Amoeboid - Thierry Karsenti
Amoeboid - Thierry Karsenti
Amoeboid - Thierry Karsenti
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
http://en.wikipedia.org/wiki/<strong>Amoeboid</strong><br />
<strong>Amoeboid</strong><br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
<strong>Amoeboid</strong><br />
Scientific classification<br />
Classes and subclasses<br />
Class Lobose pseudopods<br />
Amoebozoa<br />
Percolozoa<br />
Class Filose pseudopods<br />
Cercozoa<br />
Vampyrellids<br />
Nucleariids<br />
Class Reticulose pseudopods<br />
Foraminifera<br />
Gymnophryids<br />
Class Actinopods<br />
Radiolaria<br />
Heliozoa<br />
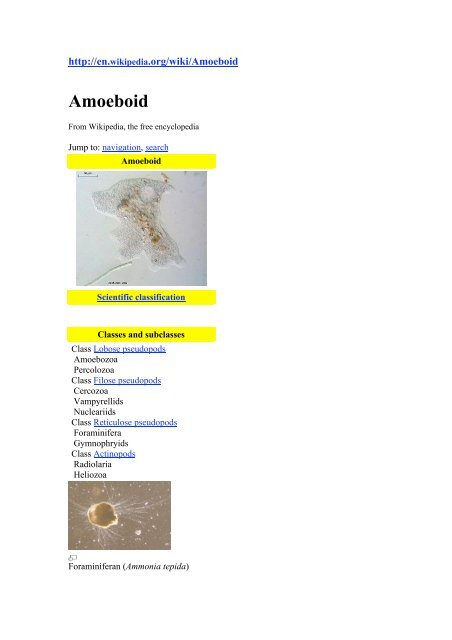
Foraminiferan (Ammonia tepida)
Heliozoan (Actinophrys sol)<br />
<strong>Amoeboid</strong>s are unicellular lifeforms that mainly consist of contractile vacuoles, a<br />
nucleus, and cytoplasm as their basic structure. They move and feed by means of<br />
temporary cytoplasmic projections, called pseudopods (false feet). They have<br />
appeared in a number of different groups. Some cells in multicellular animals may be<br />
amoeboid, for instance human white blood cells, which consume pathogens. Many<br />
protists also exist as individual amoeboid cells, or take such a form at some point in<br />
their life-cycle. The most famous such organism is Amoeba proteus; the name amoeba<br />
is variously used to describe its close relatives, other organisms similar to it, or the<br />
amoeboids in general.<br />
[edit] Morphological categories<br />
<strong>Amoeboid</strong>s may be divided into several morphological categories based on the form<br />
and structure of the pseudopods. Those where the pseudopods are supported by<br />
regular arrays of microtubules are called actinopods, and forms where they are not are<br />
called rhizopods, further divided into lobose, filose, and reticulose amoebae. There is<br />
also a strange group of giant marine amoeboids, the xenophyophores, that do not fall<br />
into any of these categories.<br />
• Lobose pseudopods are blunt, and there may be one or several on a cell,<br />
which is usually divided into a layer of clear ectoplasm surrounding more<br />
granular endoplasm. Most, including Amoeba itself, move by the body mass<br />
flowing into an anterior pseudopod. The vast majority form a monophyletic<br />
group called the Amoebozoa, which also includes most slime moulds. A<br />
second group, the Percolozoa, includes protists that can transform between<br />
amoeboid and flagellate forms.<br />
• Filose pseudopods are narrow and tapering. The vast majority of filose<br />
amoebae, including all those that produce shells, are placed within the<br />
Cercozoa together with various flagellates that tend to have amoeboid forms.<br />
The naked filose amoebae comprise two other groups, the vampyrellids and<br />
nucleariids. The latter appear to be close relatives of animals and fungi.<br />
• Reticulose pseudopods are cytoplasmic strands that branch and merge to<br />
form a net. They are found most notably among the Foraminifera, a large
group of marine protists that generally produce multi-chambered shells. There<br />
are only a few sorts of naked reticulose amoeboids, notably the gymnophryids,<br />
and their relationships are not certain.<br />
• Actinopods are divided into the radiolaria and heliozoa. The radiolaria are<br />
mostly marine protists with complex internal skeletons, including central<br />
capsules that divide the cells into granular endoplasm and frothy ectoplasm<br />
that keeps them buoyant. The heliozoa include both freshwater and marine<br />
forms that use their axopods to capture small prey, and only have simple<br />
scales or spines for skeletal elements. Both groups appear to be polyphyletic.<br />
. However, amoeboids have appeared separately in many other groups, including<br />
various different lines of algae not listed above.<br />
Δů==Subphylum Sarcodina== Sarcodina is a subphylum of the phylum<br />
Sarcomastigophora, of unicellular life forms that move by cytoplasmic flow. Some<br />
species use cytoplasmic extensions called pseudopodia for locomotion or feeding. The<br />
subphylum includes such protozoa as the common amoeba and the Foraminifera and<br />
Radiolaria. Most members of the subphylum reproduce asexually through fission,<br />
although some reproduce sexually. Sarcodina is sometimes subdůivided into two<br />
classes - Rhizopoda and Actinopoda.ÒΜκŁΔβΑhi mom.<br />
[edit] External links<br />
• The Amoebae website brings together information from published sources.<br />
• Amoebas are more than just blobs<br />
• sun animacules and amoebas<br />
• Molecular Expressions Digital Video Gallery: Pond Life - Amoeba (Protozoa)<br />
Some good, informative Amoeba videos.<br />
• Joseph Leidy's Amoeba Plates<br />
Retrieved from "http://en.wikipedia.org/wiki/<strong>Amoeboid</strong>"<br />
Categories: Protista | Cell biology | <strong>Amoeboid</strong>s | Motile cells<br />
Views<br />
• Article<br />
• Discussion<br />
• Edit this page<br />
• History<br />
Personal tools<br />
• Log in / create account<br />
Navigation
• All text is available under the terms of the GNU Free Documentation License.<br />
(See Copyrights for details.)<br />
Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc., a<br />
U.S. registered 501(c)(3) tax-deductible nonprofit charity.<br />
• Privacy policy<br />
• About Wikipedia<br />
• Disclaimers<br />
http://en.wikipedia.org/wiki/Sporozoans<br />
Apicomplexa<br />
From Wikipedia, the free encyclopedia<br />
(Redirected from Sporozoans)<br />
Jump to: navigation, search<br />
Apicomplexa<br />
Scientific classification<br />
Domain: Eukaryota<br />
Kingdom: Chromalveolata<br />
Superphylum: Alveolata<br />
Phylum: Apicomplexa<br />
Aconoidasida<br />
Classes & Subclasses<br />
• Haemosporasina<br />
• Piroplasmasina<br />
Blastocystea
Conoidasida<br />
• Coccidiasina<br />
• Gregarinasina<br />
The Apicomplexa are a large group of protists, characterized by the presence of a<br />
unique organelle called an apical complex (see also apicoplast). They are unicellular,<br />
spore-forming, and exclusively parasites of animals. Motile structures such as flagella<br />
or pseudopods are absent except in certain gamete stages. This is a diverse group<br />
including organisms such as coccidia, gregarines, piroplasms, haemogregarines, and<br />
malarias; some diseases caused by apicomplexan organisms include:<br />
• Babesiosis (Babesia)<br />
• Malaria (Plasmodium)<br />
• Cryptosporidiosis (Cryptosporidium)<br />
• Coccidian diseases including:<br />
o Cryptosporidiosis (Cryptosporidium parvum)<br />
o Cyclosporiasis (Cyclospora cayetanensis)<br />
o Toxoplasmosis (Toxoplasma gondii)<br />
Most members have a complex life-cycle, involving both asexual and sexual<br />
reproduction. Typically, a host is infected by ingesting cysts, which divide to produce<br />
sporozoites that enter its cells. Eventually, the cells burst, releasing merozoites which<br />
infect new cells. This may occur several times, until gamonts are produced, forming<br />
gametes that fuse to create new cysts. There are many variations on this basic pattern,<br />
however, and many Apicomplexa have more than one host.<br />
Generic life cycle of an apicomplexa: 1-zygote (cyst), 2-sporozoites, 3-merozoites, 4gametocytes.
Apicomplexan structure: 1-polar ring, 2-conoid, 3-micronemes, 4-rhoptries, 5nucleus,<br />
6-nucleolus, 7-mitochondria, 8-posterior ring, 9-alveoli, 10-golgi apparatus,<br />
11-micropore.<br />
The apical complex includes vesicles called rhoptries and micronemes, which open at<br />
the anterior of the cell. These secrete enzymes that allow the parasite to enter other<br />
cells. The tip is surrounded by a band of microtubules, called the polar ring, and<br />
among the Conoidasida there is also a funnel of rods called the conoid.. [1] Over the<br />
rest of the cell, except for a diminished mouth called the micropore, the membrane is<br />
supported by vesicles called alveoli, forming a semi-rigid pellicle.<br />
The presence of alveoli and other traits place the Apicomplexa among a group called<br />
the alveolates. Several related flagellates, such as Perkinsus and Colpodella have<br />
structures similar to the polar ring and were formerly included here, but most appear<br />
to be closer relatives of the dinoflagellates. They are probably similar to the common<br />
ancestor of the two groups.<br />
Another similarity is that apicomplexan cells contain a single plastid, called the<br />
apicoplast, surrounded by either 3 or four membranes. Its functions are thought to<br />
include tasks such as lipid synthesis, it appears to be necessary for survival. They are<br />
generally considered to share a common origin with the chloroplasts of<br />
dinoflagellates, although some studies suggest they are ultimately derived from green<br />
rather than red algae.<br />
The Apicomplexa comprise the bulk of what used to be called the Sporozoa, a group<br />
for parasitic protozoans without flagella, pseudopods, or cilia. Most of the<br />
Apicomplexa are motile however. The other main lines were the Ascetosporea, the<br />
Myxozoa (now known to be derived from animals), and the Microsporidia (now<br />
known to be derived from fungi). Sometimes the name Sporozoa is taken as a<br />
synonym for the Apicomplexa, or occasionally as a subset.<br />
Contents<br />
[hide]<br />
• 1 Blood borne genera<br />
• 2 Disease Genomics<br />
• 3 References<br />
• 4 External links<br />
[edit] Blood borne genera<br />
Within the Apicomplexa there are three groups of blood borne parasites. These<br />
species lie within in three suborders.<br />
• suborder Adeleorina - 8 genera<br />
• suborder Haemosporina - all genera in this suborder
• suborder Eimeriorina - 2 genera (Lankesterella and Schellackia)<br />
Blood parasites belonging to the suborder Adeleorina are collectively known as<br />
haemogregarines. Currently their sister group is thought to be the piroplasms.<br />
Suborder Adeleorina has ~400 species and has been organised into four large and 4<br />
small genera.<br />
The larger genera are:<br />
genera:<br />
• family Haemogregarinidae - taxon created by Neveu-Lemaire in 1901<br />
• Haemogregarina - taxon created by Danilewsky in 1885<br />
• Cyrilia - taxon created by Lainson in 1981<br />
genera:<br />
genera:<br />
• family Karyolysidae - taxon created by Wenyon in 1926<br />
• Karyolysus - taxon created by Labbe in 1894<br />
• family Hepatozoidae - taxon created by Wenyon in 1926<br />
• Hepatozoon - taxon created by Miller in 1908<br />
The smaller genera are :<br />
• Hemolivia - taxon created by Petit et al in 1990<br />
• Desseria - taxon created by Siddall in 1995<br />
genera:<br />
• family Dactylosomatidae<br />
• Dactylosoma<br />
• Babesiosoma
Notes:<br />
Species of the genus Desseria infect fish and lack erythrocytic merogony.<br />
The species of the genera Dactylosoma and Babesiosoma infect fish and reptiles.<br />
Leeches are the only known vectors for these species and their vertebrate hosts are<br />
aquatic.<br />
[edit] Disease Genomics<br />
As noted above, many of the apicomplexan parasites are important pathogens of<br />
human and domestic animals. In contrast to bacterial pathogens, these apicomplexan<br />
parasites are eukaryotes and share many metabolic pathways with their animal hosts.<br />
This fact makes therapeutic target development extremely difficult – a drug that<br />
harms an apicomplexan parasite is also likely to harm its human host. Currently there<br />
are no effective vaccines or treatments available for most diseases caused by these<br />
parasites. Biomedical research on these parasites is challenging because it is often<br />
difficult, if not impossible, to maintain live parasite cultures in the laboratory and to<br />
genetically manipulate these organisms. In the recent years, several of the<br />
apicomplexan species have been selected for genome sequencing. The availability of<br />
genome sequences provides a new opportunity for scientists to learn more about the<br />
evolution and biochemical capacity of these parasite. A NIH-funded database,<br />
ApiDB.org, provides public access to currently available genomic data sets.<br />
[edit] References<br />
1. ^ Duszynski1, Donald W.; Steve J. Upton and Lee Couch (2004-02-21). The<br />
Coccidia of the World (Online database). Department of Biology, University of New<br />
Mexico, and Division of Biology, Kansas State University.<br />
[edit] External links<br />
• The Taxonomicon & Systema Naturae (Website database). Taxon: Genus<br />
Cryptosporidium. Universal Taxonomic Services, Amsterdam, The<br />
Netherlands (2000).<br />
Retrieved from "http://en.wikipedia.org/wiki/Apicomplexa"<br />
Categories: Parasitic protists | Apicomplexa<br />
Views<br />
• Article<br />
• Discussion<br />
• Edit this page<br />
• History<br />
Personal tools
• Türkçe<br />
http://en.wikipedia.org/wiki/Bacterial_growth<br />
Bacterial growth<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
Growth is shown as L = log(numbers) where numbers is the number of colony<br />
forming units per ml, versus T (time.)<br />
Bacterial growth is the division of one bacterium into two idential daughter cells<br />
during a process called binary fission. Hence, local doubling of the bacterial<br />
population occurs. Both daughter cells from the division do not necessarily survive.<br />
However, if the number surviving exceeds unity on average, the bacterial population<br />
undergoes exponential growth. The measurement of an exponential bacterial growth<br />
curve in batch culture was traditionally a part of the training of all microbiologists; the<br />
basic means requires bacterial enumeration (cell counting) by direct and individual<br />
(microscopic, flow cytometry [1] ), direct and bulk (biomass), indirect and individual<br />
(colony counting), or indirect and bulk (most probable number, turbidity, nutrient<br />
uptake) methods. Models reconcile theory with the measurements [2] .<br />
In autecological studies, bacterial growth in batch culture can be modeled with four<br />
different phases: lag phase (A), exponential or log phase (B), stationary phase (C),<br />
and death phase (D).<br />
1. During lag phase, bacteria adapt themselves to growth conditions. It is the<br />
period where the individual bacteria are maturing and not yet able to divide.<br />
During the lag phase of the bacterial growth cycle, synthesis of RNA, enzymes<br />
and other molecules occurs.
2. During the exponential phase (sometimes called the log phase), the number of<br />
new bacteria appearing per unit time is proportional to the present population.<br />
This gives rise to the classic exponential growth curve, in which the logarithm<br />
of the population density rises linearly with time (see figure). The actual rate<br />
of this growth (i.e. the slope of the line in the figure) depends upon the growth<br />
conditions, which affect the frequency of cell division events and the<br />
probability of both daughter cells surviving. Exponential growth cannot<br />
continue indefinitely, however, because the medium is soon depleted of<br />
nutrients and enriched with wastes.<br />
3. During stationary phase, the growth rate slows as a result of nutrient depletion<br />
and accumulation of toxic products. This phase is reached as the bacteria<br />
begin to exhaust the resources that are available to them.<br />
4. At death phase, bacteria run out of nutrients and die.<br />
This basic batch culture growth model draws out and emphasizes aspects of bacterial<br />
growth which may differ from the growth of macrofauna. It emphasizes clonality,<br />
asexual binary division, the short development time relative to replication itself, the<br />
seemingly low death rate, the need to move from a dormant state to a reproductive<br />
state or to condition the media, and finally, the tendency of lab adapted strains to<br />
exhaust their nutrients.<br />
In reality, even in batch culture, the four phases are not well defined. The cells do not<br />
reproduce in synchrony without explicit and continual prompting (as in experiments<br />
with stalked bacteria [3] ) and their logarithmic phase growth is often not ever a<br />
constant rate, but instead a slowly decaying rate, a constant stochastic response to<br />
pressures both to reproduce and to go dormant in the face of declining nutrient<br />
concentrations and increasing waste concentrations.<br />
Batch culture is the most common laboratory growth environment in which bacterial<br />
growth is studied, but it is only one of many. It is ideally spatially unstructured and<br />
temporally structured. The bacterial culture is incubated in a closed vessel with a<br />
single batch of medium. In some experimental regimes, some of the bacterial culture<br />
is periodically removed to a fresh sterile media is added. In the extreme case, this<br />
leads to the continual renewal of the nutrients. This is a chemostat also known as<br />
continuous culture. It is ideally spatially unstructured and temporally unstructured, in<br />
an equilibrium state defined by the nutrient supply rate and the reaction of the<br />
bacteria. In comparison to batch culture, bacteria are maintained in expodential<br />
growth phase and the grow growth rate of the bacteria is known. Related devices<br />
include turbidostats and auxostats.<br />
Bacterial growth can be suppressed with bacteriostats, without necessarily<br />
killing the bacteria. In a synecological, a true-to-nature situation, where more than<br />
one bacterial species is present, the growth of microbes is more dynamic and<br />
continual.<br />
Liquid is not the only laboratory environment for bacterial growth. Spatially<br />
structured environments such as biofilms or agar surfaces present additional complex<br />
growth models.
[edit] References<br />
1. ^ Skarstad K, Steen HB, Boye E (1983). "Cell cycle parameters of slowly growing<br />
Escherichia coli B/r studied by flow cytometry". J. Bacteriol. 154 (2): 656–62. PMID<br />
6341358.<br />
2. ^ Zwietering M H, Jongenburger I, Rombouts F M, van 'T Riet K (1990). "Modeling<br />
of the Bacterial Growth Curve". Applied and Environmental Microbiology 56 (6):<br />
1875-1881.<br />
3. ^ Novick A (1955). "Growth of Bacteria". Annual Review of Microbiology 9: 97-<br />
110.<br />
[edit] External links<br />
• An examination of the exponential growth of bacterial populations<br />
• Science aid: Microbial Populations<br />
• Microbial Growth, BioMineWiki<br />
This article includes material from an article posted on 26 April 2003 on Nupedia;<br />
written by Nagina Parmar; reviewed and approved by the Biology group; editor,<br />
Gaytha Langlois; lead reviewer, Gaytha Langlois ; lead copyeditors, Ruth Ifcher. and<br />
Jan Hogle.<br />
Retrieved from "http://en.wikipedia.org/wiki/Bacterial_growth"<br />
Categories: Bacteriology | Population<br />
Views<br />
• Article<br />
• Discussion<br />
• Edit this page<br />
• History<br />
Personal tools<br />
• Log in / create account<br />
Navigation<br />
• Main Page<br />
• Contents<br />
• Featured content<br />
• Current events<br />
• Random article<br />
Interaction<br />
• About Wikipedia<br />
• Community portal
Search<br />
• Recent changes<br />
• Contact Wikipedia<br />
• Donate to Wikipedia<br />
• Help<br />
Toolbox<br />
• What links here<br />
• Related changes<br />
• Upload file<br />
• Special pages<br />
• Printable version<br />
• Permanent link<br />
• Cite this page<br />
Languages<br />
• Polski<br />
• Українська<br />
• This page was last modified on 12 March 2008, at 23:31.<br />
• All text is available under the terms of the GNU Free Documentation License.<br />
(See Copyrights for details.)<br />
Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc., a<br />
U.S. registered 501(c)(3) tax-deductible nonprofit charity.<br />
• Privacy policy<br />
• About Wikipedia<br />
• Disclaimers<br />
Bacteria<br />
http://en.wikipedia.org/wiki/Bacteria<br />
From Wikipedia, the free encyclopedia
Jump to: navigation, search<br />
For other uses, see Bacteria (disambiguation).<br />
Bacteria<br />
Fossil range: Archean or earlier -<br />
Recent<br />
Escherichia coli cells magnified<br />
25,000 times<br />
Scientific classification<br />
Domain: Bacteria<br />
Phyla<br />
Acidobacteria<br />
Actinobacteria<br />
Aquificae<br />
Bacteroidetes<br />
Chlamydiae<br />
Chlorobi<br />
Chloroflexi<br />
Chrysiogenetes<br />
Cyanobacteria<br />
Deferribacteres<br />
Deinococcus-Thermus<br />
Dictyoglomi<br />
Fibrobacteres<br />
Firmicutes<br />
Fusobacteria<br />
Gemmatimonadetes<br />
Nitrospirae<br />
Planctomycetes<br />
Proteobacteria<br />
Spirochaetes<br />
Thermodesulfobacteria<br />
Thermomicrobia<br />
Thermotogae<br />
Verrucomicrobia<br />
Bacteria (singular: bacterium) are unicellular microorganisms. Typically a few<br />
micrometres in length, bacteria have a wide range of shapes, ranging from spheres to
ods to spirals. Bacteria are ubiquitous in every habitat on Earth, growing in soil,<br />
acidic hot springs, radioactive waste, [1] seawater, and deep in the Earth's crust. There<br />
are typically 40 million bacterial cells in a gram of soil and a million bacterial cells in<br />
a millilitre of fresh water; in all, there are approximately five nonillion (5×10 30 )<br />
bacteria on Earth, [2] forming much of the world's biomass. [3] Bacteria are vital in<br />
recycling nutrients, and many important steps in nutrient cycles depend on bacteria,<br />
such as the fixation of nitrogen from the atmosphere. However, most of these bacteria<br />
have not been characterized, and only about half of the phyla of bacteria have species<br />
that can be cultured in the laboratory. [4] The study of bacteria is known as<br />
bacteriology, a branch of microbiology.<br />
There are approximately ten times as many bacterial cells as human cells in the<br />
human body, with large numbers of bacteria on the skin and in the digestive tract. [5]<br />
Although the vast majority of these bacteria are rendered harmless or beneficial by the<br />
protective effects of the immune system, a few are pathogenic bacteria and cause<br />
infectious diseases, including cholera, syphilis, anthrax, leprosy and bubonic plague.<br />
The most common fatal bacterial diseases are respiratory infections, with tuberculosis<br />
alone killing about 2 million people a year, mostly in sub-Saharan Africa. [6] In<br />
developed countries, antibiotics are used to treat bacterial infections and in various<br />
agricultural processes, so antibiotic resistance is becoming common. In industry,<br />
bacteria are important in processes such as sewage treatment, the production of cheese<br />
and yoghurt, and the manufacture of antibiotics and other chemicals. [7]<br />
Bacteria are prokaryotes. Unlike cells of animals and other eukaryotes, bacterial cells<br />
do not contain a nucleus and rarely harbour membrane-bound organelles. Although<br />
the term bacteria traditionally included all prokaryotes, the scientific classification<br />
changed after the discovery in the 1990s that prokaryotic life consists of two very<br />
different groups of organisms that evolved independently from an ancient common<br />
ancestor. These evolutionary domains are called Bacteria and Archaea. [8]<br />
Contents<br />
[hide]<br />
• 1 History of bacteriology<br />
• 2 Origin and early evolution<br />
• 3 Morphology<br />
• 4 Cellular structure<br />
o 4.1 Intracellular structures<br />
o 4.2 Extracellular structures<br />
o 4.3 Endospores<br />
• 5 Metabolism<br />
• 6 Growth and reproduction<br />
• 7 Genetics<br />
• 8 Movement<br />
• 9 Classification and identification<br />
• 10 Interactions with other organisms<br />
o 10.1 Mutualists<br />
o 10.2 Pathogens
• 11 Significance in technology and industry<br />
• 12 See also<br />
• 13 References<br />
• 14 Further reading<br />
• 15 External links<br />
History of bacteriology<br />
Further information: Microbiology<br />
Antonie van Leeuwenhoek, the first microbiologist and the first person to observe<br />
bacteria using a microscope.<br />
Bacteria were first observed by Antonie van Leeuwenhoek in 1676, using a singlelens<br />
microscope of his own design. [9] He called them "animalcules" and published his<br />
observations in a series of letters to the Royal Society. [10][11][12] The name bacterium<br />
was introduced much later, by Christian Gottfried Ehrenberg in 1828, and is derived<br />
from the Greek word βακτήριον -α , bacterion -a , meaning "small staff". [13]<br />
Louis Pasteur demonstrated in 1859 that the fermentation process is caused by the<br />
growth of microorganisms, and that this growth is not due to spontaneous generation.<br />
(Yeasts and molds, commonly associated with fermentation, are not bacteria, but<br />
rather fungi.) Along with his contemporary, Robert Koch, Pasteur was an early<br />
advocate of the germ theory of disease. [14] Robert Koch was a pioneer in medical<br />
microbiology and worked on cholera, anthrax and tuberculosis. In his research into<br />
tuberculosis, Koch finally proved the germ theory, for which he was awarded a Nobel<br />
Prize in 1905. [15] In Koch's postulates, he set out criteria to test if an organism is the<br />
cause of a disease; these postulates are still used today. [16]<br />
Though it was known in the nineteenth century that bacteria are the cause of many<br />
diseases, no effective antibacterial treatments were available. [17] In 1910, Paul Ehrlich<br />
developed the first antibiotic, by changing dyes that selectively stained Treponema<br />
pallidum—the spirochaete that causes syphilis—into compounds that selectively
killed the pathogen. [18] Ehrlich had been awarded a 1908 Nobel Prize for his work on<br />
immunology, and pioneered the use of stains to detect and identify bacteria, with his<br />
work being the basis of the Gram stain and the Ziehl-Neelsen stain. [19]<br />
A major step forward in the study of bacteria was the recognition in 1977 by Carl<br />
Woese that archaea have a separate line of evolutionary descent from bacteria. [20] This<br />
new phylogenetic taxonomy was based on the sequencing of 16S ribosomal RNA, and<br />
divided prokaryotes into two evolutionary domains, as part of the three-domain<br />
system. [21]<br />
Origin and early evolution<br />
Further information: Timeline of evolution<br />
The ancestors of modern bacteria were single-celled microorganisms that were the<br />
first forms of life to develop on earth, about 4 billion years ago. For about 3 billion<br />
years, all organisms were microscopic, and bacteria and archaea were the dominant<br />
forms of life. [22][23] Although bacterial fossils exist, such as stromatolites, their lack of<br />
distinctive morphology prevents them from being used to examine the past history of<br />
bacterial evolution, or to date the time of origin of a particular bacterial species.<br />
However, gene sequences can be used to reconstruct the bacterial phylogeny, and<br />
these studies indicate that bacteria diverged first from the archaeal/eukaryotic<br />
lineage. [24] The most recent common ancestor of bacteria and archaea was probably a<br />
hyperthermophile that lived about 2.5 billion–3.2 billion years ago. [25][26]<br />
Bacteria were also involved in the second great evolutionary divergence, that of the<br />
archaea and eukaryotes. Here, eukaryotes resulted from ancient bacteria entering into<br />
endosymbiotic associations with the ancestors of eukaryotic cells, which were<br />
themselves possibly related to the Archaea. [27][28] This involved the engulfment by<br />
proto-eukaryotic cells of alpha-proteobacterial symbionts to form either mitochondria<br />
or hydrogenosomes, which are still being found in all known Eukarya (sometimes in<br />
highly reduced form, e.g. in ancient "amitochondrial" protozoa). Later on, an<br />
independent second engulfment by some mitochondria-containing eukaryotes of<br />
cyanobacterial-like organisms led to the formation of chloroplasts in algae and plants.<br />
There are even some algal groups known that clearly originated from subsequent<br />
events of endosymbiosis by heterotrophic eukaryotic hosts engulfing a eukaryotic<br />
algae that developed into "second-generation" plastids. [29][30]<br />
Morphology
Bacteria display a large diversity of cell morphologies and arrangements<br />
Bacteria display a wide diversity of shapes and sizes, called morphologies. Bacterial<br />
cells are about 10 times smaller than eukaryotic cells and are typically 0.5–<br />
5.0 micrometres in length. However, a few species–for example Thiomargarita<br />
namibiensis and Epulopiscium fishelsoni–are up to half a millimetre long and are<br />
visible to the unaided eye. [31] Among the smallest bacteria are members of the genus<br />
Mycoplasma, which measure only 0.3 micrometres, as small as the largest viruses. [32]<br />
Most bacterial species are either spherical, called cocci (sing. coccus, from Greek<br />
kókkos, grain, seed) or rod-shaped, called bacilli (sing. bacillus, from Latin baculus,<br />
stick). Some rod-shaped bacteria, called vibrio, are slightly curved or comma-shaped;<br />
others, can be spiral-shaped, called spirilla, or tightly coiled, called spirochaetes. A<br />
small number of species even have tetrahedral or cuboidal shapes. [33] This wide<br />
variety of shapes is determined by the bacterial cell wall and cytoskeleton, and is<br />
important because it can influence the ability of bacteria to acquire nutrients, attach to<br />
surfaces, swim through liquids and escape predators. [34][35]<br />
Many bacterial species exist simply as single cells, others associate in characteristic<br />
patterns: Neisseria form diploids (pairs), Streptococcus form chains, and<br />
Staphylococcus group together in "bunch of grapes" clusters. Bacteria can also be<br />
elongated to form filaments, for example the Actinobacteria. Filamentous bacteria are<br />
often surrounded by a sheath that contains many individual cells; certain types, such<br />
as species of the genus Nocardia, even form complex, branched filaments, similar in<br />
appearance to fungal mycelia. [36]
The range of sizes shown by prokaryotes, relative to those of other organisms and<br />
biomolecules<br />
Bacteria often attach to surfaces and form dense aggregations called biofilms or<br />
bacterial mats. These films can range from a few micrometers in thickness to up to<br />
half a meter in depth, and may contain multiple species of bacteria, protists and<br />
archaea. Bacteria living in biofilms display a complex arrangement of cells and<br />
extracellular components, forming secondary structures such as microcolonies,<br />
through which there are networks of channels to enable better diffusion of<br />
nutrients. [37][38] In natural environments, such as soil or the surfaces of plants, the<br />
majority of bacteria are bound to surfaces in biofilms. [39] Biofilms are also important<br />
for chronic bacterial infections and infections of implanted medical devices, as<br />
bacteria protected within these structures are much harder to kill than individual<br />
bacteria. [40]<br />
Even more complex morphological changes are sometimes possible. For example,<br />
when starved of amino acids, Myxobacteria detect surrounding cells in a process<br />
known as quorum sensing, migrate towards each other, and aggregate to form fruiting<br />
bodies up to 500 micrometres long and containing approximately 100,000 bacterial<br />
cells. [41] In these fruiting bodies, the bacteria perform separate tasks; this type of<br />
cooperation is a simple type of multicellular organisation. For example, about one in<br />
10 cells migrate to the top of these fruiting bodies and differentiate into a specialised<br />
dormant state called myxospores, which are more resistant to desiccation and other<br />
adverse environmental conditions than are ordinary cells. [42]<br />
Cellular structure<br />
Further information: Bacterial cell structure
Diagram of the cellular structure of a typical bacterial cell<br />
Intracellular structures<br />
The bacterial cell is surrounded by a lipid membrane, or cell membrane, which<br />
encompasses the contents of the cell and acts as a barrier to hold nutrients, proteins<br />
and other essential components of the cytoplasm within the cell. As they are<br />
prokaryotes, bacteria do not tend to have membrane-bound organelles in their<br />
cytoplasm and thus contain few intracellular structures. They consequently lack a<br />
nucleus, mitochondria, chloroplasts and the other organelles present in eukaryotic<br />
cells, such as the Golgi apparatus and endoplasmic reticulum. [43] However, recent<br />
research is identifying increasing amounts of structural complexity in bacteria, such as<br />
the discovery of the prokaryotic cytoskeleton. [44][45]<br />
Many important biochemical reactions, such as energy generation, occur due to<br />
concentration gradients across membranes, creating a potential difference analogous<br />
to a battery. The absence of internal membranes in bacteria means these reactions,<br />
such as electron transport, occur across the cell membrane, between the cytoplasm<br />
and the periplasmic space. [46] Additionally, while some transporter proteins consume<br />
chemical energy, others harness concentration gradients to import nutrients across the<br />
cell membrane or to expel undesired molecules from the cytoplasm.<br />
Bacteria do not have a membrane-bound nucleus, and their genetic material is<br />
typically a single circular chromosome located in the cytoplasm in an irregularly<br />
shaped body called the nucleoid. [47] The nucleoid contains the chromosome with<br />
associated proteins and RNA. Like all living organisms, bacteria contain ribosomes<br />
for the production of proteins, but the structure of the bacterial ribosome is different<br />
from those of eukaryotes and Archaea. [48] The order Planctomycetes are an exception<br />
to the general absence of internal membranes in bacteria, because they have a<br />
membrane around their nucleoid and contain other membrane-bound cellular<br />
structures. [49]<br />
Some bacteria produce intracellular nutrient storage granules, such as glycogen, [50]<br />
polyphosphate, [51] sulfur [52] or polyhydroxyalkanoates. [53] These granules enable<br />
bacteria to store compounds for later use. Certain bacterial species, such as the
photosynthetic Cyanobacteria, produce internal gas vesicles, which they use to<br />
regulate their buoyancy - allowing them to move up or down into water layers with<br />
different light intensities and nutrient levels. [54]<br />
Extracellular structures<br />
Further information: Cell envelope<br />
Around the outside of the cell membrane is the bacterial cell wall. Bacterial cell walls<br />
are made of peptidoglycan (called murein in older sources), which is made from<br />
polysaccharide chains cross-linked by unusual peptides containing D-amino acids. [55]<br />
Bacterial cell walls are different from the cell walls of plants and fungi, which are<br />
made of cellulose and chitin, respectively. [56] The cell wall of bacteria is also distinct<br />
from that of Archaea, which do not contain peptidoglycan. The cell wall is essential to<br />
the survival of many bacteria, and the antibiotic penicillin is able to kill bacteria by<br />
inhibiting a step in the synthesis of peptidoglycan. [56]<br />
There are broadly speaking two different types of cell wall in bacteria, called Grampositive<br />
and Gram-negative. The names originate from the reaction of cells to the<br />
Gram stain, a test long-employed for the classification of bacterial species. [57]<br />
Gram-positive bacteria possess a thick cell wall containing many layers of<br />
peptidoglycan and teichoic acids. In contrast, Gram-negative bacteria have a relatively<br />
thin cell wall consisting of a few layers of peptidoglycan surrounded by a second lipid<br />
membrane containing lipopolysaccharides and lipoproteins. Most bacteria have the<br />
Gram-negative cell wall, and only the Firmicutes and Actinobacteria (previously<br />
known as the low G+C and high G+C Gram-positive bacteria, respectively) have the<br />
alternative Gram-positive arrangement. [58] These differences in structure can produce<br />
differences in antibiotic susceptibility; for instance, vancomycin can kill only Grampositive<br />
bacteria and is ineffective against Gram-negative pathogens, such as<br />
Haemophilus influenzae or Pseudomonas aeruginosa. [59]<br />
In many bacteria an S-layer of rigidly arrayed protein molecules covers the outside of<br />
the cell. [60] This layer provides chemical and physical protection for the cell surface<br />
and can act as a macromolecular diffusion barrier. S-layers have diverse but mostly<br />
poorly understood functions, but are known to act as virulence factors in<br />
Campylobacter and contain surface enzymes in Bacillus stearothermophilus. [61]<br />
Helicobacter pylori electron micrograph, showing multiple flagella on the cell surface
Flagella are rigid protein structures, about 20 nanometres in diameter and up to<br />
20 micrometres in length, that are used for motility. Flagella are driven by the energy<br />
released by the transfer of ions down an electrochemical gradient across the cell<br />
membrane. [62]<br />
Fimbriae are fine filaments of protein, just 2–10 nanometres in diameter and up to<br />
several micrometers in length. They are distributed over the surface of the cell, and<br />
resemble fine hairs when seen under the electron microscope. Fimbriae are believed<br />
to be involved in attachment to solid surfaces or to other cells and are essential for the<br />
virulence of some bacterial pathogens. [63] Pili (sing. pilus) are cellular appendages,<br />
slightly larger than fimbriae, that can transfer genetic material between bacterial cells<br />
in a process called conjugation (see bacterial genetics, below). [64]<br />
Capsules or slime layers are produced by many bacteria to surround their cells, and<br />
vary in structural complexity: ranging from a disorganised slime layer of extracellular<br />
polymer, to a highly structured capsule or glycocalyx. These structures can<br />
protect cells from engulfment by eukaryotic cells, such as macrophages. [65] They can<br />
also act as antigens and be involved in cell recognition, as well as aiding attachment<br />
to surfaces and the formation of biofilms. [66]<br />
The assembly of these extracellular structures is dependent on bacterial secretion<br />
systems. These transfer proteins from the cytoplasm into the periplasm or into the<br />
environment around the cell. Many types of secretion systems are known and these<br />
structures are often essential for the virulence of pathogens, so are intensively<br />
studied. [67]<br />
Endospores<br />
Further information: Endospores<br />
Bacillus anthracis (stained purple) growing in cerebrospinal fluid<br />
Certain genera of Gram-positive bacteria, such as Bacillus, Clostridium,<br />
Sporohalobacter, Anaerobacter and Heliobacterium, can form highly resistant,<br />
dormant structures called endospores. [68] In almost all cases, one endospore is formed<br />
and this is not a reproductive process, although Anaerobacter can make up to seven<br />
endospores in a single cell. [69] Endospores have a central core of cytoplasm containing<br />
DNA and ribosomes surrounded by a cortex layer and protected by an impermeable<br />
and rigid coat.
Endospores show no detectable metabolism and can survive extreme physical and<br />
chemical stresses, such as high levels of UV light, gamma radiation, detergents,<br />
disinfectants, heat, pressure and desiccation. [70] In this dormant state, these organisms<br />
may remain viable for millions of years, [71][72] and endospores even allow bacteria to<br />
survive exposure to the vacuum and radiation in space. [73] Endospore-forming bacteria<br />
can also cause disease: for example, anthrax can be contracted by the inhalation of<br />
Bacillus anthracis endospores, and contamination of deep puncture wounds with<br />
Clostridium tetani endospores causes tetanus. [74]<br />
Metabolism<br />
Further information: Microbial metabolism<br />
Filaments of photosynthetic cyanobacteria<br />
In contrast to higher organisms, bacteria exhibit an extremely wide variety of<br />
metabolic types. [75] The distribution of metabolic traits within a group of bacteria has<br />
traditionally been used to define their taxonomy, but these traits often do not<br />
correspond with modern genetic classifications. [76] Bacterial metabolism is classified<br />
on the basis of three major criteria: the kind of energy used for growth, the source of<br />
carbon, and the electron donors used for growth. An additional criterion of respiratory<br />
microorganisms are the electron acceptors used for aerobic or anaerobic<br />
respiration. [77]<br />
Carbon metabolism in bacteria is either heterotrophic, where organic carbon<br />
compounds are used as carbon sources, or autotrophic, meaning that cellular carbon is<br />
obtained by fixing carbon dioxide. Typical autotrophic bacteria are phototrophic<br />
cyanobacteria, green sulfur-bacteria and some purple bacteria, but also many<br />
chemolithotrophic species, such as nitrifying or sulfur-oxidising bacteria. [78] Energy<br />
metabolism of bacteria is either based on phototrophy, the use of light through<br />
photosynthesis, or on chemotrophy, the use of chemical substances for energy, which<br />
are mostly oxidised at the expense of oxygen or alternative electron acceptors<br />
(aerobic/anaerobic respiration).<br />
Finally, bacteria are further divided into lithotrophs that use inorganic electron donors<br />
and organotrophs that use organic compounds as electron donors. Chemotrophic<br />
organisms use the respective electron donors for energy conservation (by<br />
aerobic/anaerobic respiration or fermentation) and biosynthetic reactions (e.g. carbon<br />
dioxide fixation), whereas phototrophic organisms use them only for biosynthetic<br />
purposes. Respiratory organisms use chemical compounds as a source of energy by<br />
taking electrons from the reduced substrate and transferring them to a terminal
electron acceptor in a redox reaction. This reaction releases energy that can be used to<br />
synthesise ATP and drive metabolism. In aerobic organisms, oxygen is used as the<br />
electron acceptor. In anaerobic organisms other inorganic compounds, such as nitrate,<br />
sulfate or carbon dioxide are used as electron acceptors. This leads to the ecologically<br />
important processes of denitrification, sulfate reduction and acetogenesis,<br />
respectively.<br />
Another way of life of chemotrophs in the absence of possible electron acceptors is<br />
fermentation, where the electrons taken from the reduced substrates are transferred to<br />
oxidised intermediates to generate reduced fermentation products (e.g. lactate,<br />
ethanol, hydrogen, butyric acid). Fermentation is possible, because the energy content<br />
of the substrates is higher than that of the products, which allows the organisms to<br />
synthesise ATP and drive their metabolism. [79][80]<br />
These processes are also important in biological responses to pollution; for example,<br />
sulfate-reducing bacteria are largely responsible for the production of the highly toxic<br />
forms of mercury (methyl- and dimethylmercury) in the environment. [81] Nonrespiratory<br />
anaerobes use fermentation to generate energy and reducing power,<br />
secreting metabolic by-products (such as ethanol in brewing) as waste. Facultative<br />
anaerobes can switch between fermentation and different terminal electron acceptors<br />
depending on the environmental conditions in which they find themselves.<br />
Lithotrophic bacteria can use inorganic compounds as a source of energy. Common<br />
inorganic electron donors are hydrogen, carbon monoxide, ammonia (leading to<br />
nitrification), ferrous iron and other reduced metal ions, and several reduced sulfur<br />
compounds. Unusually, the gas methane can be used by methanotrophic bacteria as<br />
both a source of electrons and a substrate for carbon anabolism. [82] In both aerobic<br />
phototrophy and chemolithotrophy, oxygen is used as a terminal electron acceptor,<br />
while under anaerobic conditions inorganic compounds are used instead. Most<br />
lithotrophic organisms are autotrophic, whereas organotrophic organisms are<br />
heterotrophic.<br />
In addition to fixing carbon dioxide in photosynthesis, some bacteria also fix nitrogen<br />
gas (nitrogen fixation) using the enzyme nitrogenase. This environmentally important<br />
trait can be found in bacteria of nearly all the metabolic types listed above, but is not<br />
universal. [83]<br />
Growth and reproduction<br />
Further information: Bacterial growth<br />
Unlike multicellular organisms, increases in the size of bacteria (cell growth) and<br />
their reproduction by cell division are tightly linked in unicellular organisms. Bacteria<br />
grow to a fixed size and then reproduce through binary fission, a form of asexual<br />
reproduction. [84] Under optimal conditions, bacteria can grow and divide extremely<br />
rapidly, and bacterial populations can double as quickly as every 9.8 minutes. [85] In<br />
cell division, two identical clone daughter cells are produced. Some bacteria, while<br />
still reproducing asexually, form more complex reproductive structures that help<br />
disperse the newly-formed daughter cells. Examples include fruiting body formation
y Myxobacteria and arial hyphae formation by Streptomyces, or budding. Budding<br />
involves a cell forming a protrusion that breaks away and produces a daughter cell.<br />
A growing colony of Escherichia coli cells [86]<br />
In the laboratory, bacteria are usually grown using solid or liquid media. Solid growth<br />
media such as agar plates are used to isolate pure cultures of a bacterial strain.<br />
However, liquid growth media are used when measurement of growth or large<br />
volumes of cells are required. Growth in stirred liquid media occurs as an even cell<br />
suspension, making the cultures easy to divide and transfer, although isolating single<br />
bacteria from liquid media is difficult. The use of selective media (media with specific<br />
nutrients added or deficient, or with antibiotics added) can help identify specific<br />
organisms. [87]<br />
Most laboratory techniques for growing bacteria use high levels of nutrients to<br />
produce large amounts of cells cheaply and quickly. However, in natural<br />
environments nutrients are limited, meaning that bacteria cannot continue to<br />
reproduce indefinitely. This nutrient limitation has led the evolution of different<br />
growth strategies (see r/K selection theory). Some organisms can grow extremely<br />
rapidly when nutrients become available, such as the formation of algal (and<br />
cyanobacterial) blooms that often occur in lakes during the summer. [88] Other<br />
organisms have adaptations to harsh environments, such as the production of multiple<br />
antibiotics by Streptomyces that inhibit the growth of competing microorganisms. [89]<br />
In nature, many organisms live in communities (e.g. biofilms) which may allow for<br />
increased supply of nutrients and protection from environmental stresses. [39] These<br />
relationships can be essential for growth of a particular organism or group of<br />
organisms (syntrophy). [90]<br />
Bacterial growth follows three phases. When a population of bacteria first enter a<br />
high-nutrient environment that allows growth, the cells need to adapt to their new<br />
environment. The first phase of growth is the lag phase, a period of slow growth when<br />
the cells are adapting to the high-nutrient environment and preparing for fast growth.<br />
The lag phase has high biosynthesis rates, as proteins necessary for rapid growth are<br />
produced. [91] The second phase of growth is the logarithmic phase (log phase), also
known as the exponential phase. The log phase is marked by rapid exponential<br />
growth. The rate at which cells grow during this phase is known as the growth rate<br />
(k), and the time it takes the cells to double is known as the generation time (g).<br />
During log phase, nutrients are metabolised at maximum speed until one of the<br />
nutrients is depleted and starts limiting growth. The final phase of growth is the<br />
stationary phase and is caused by depleted nutrients. The cells reduce their metabolic<br />
activity and consume non-essential cellular proteins. The stationary phase is a<br />
transition from rapid growth to a stress response state and there is increased<br />
expression of genes involved in DNA repair, antioxidant metabolism and nutrient<br />
transport. [92]<br />
Genetics<br />
Further information: Plasmid, Genome<br />
Most bacteria have a single circular chromosome that can range in size from only<br />
160,000 base pairs in the endosymbiotic bacteria Candidatus Carsonella ruddii, [93] to<br />
12,200,000 base pairs in the soil-dwelling bacteria Sorangium cellulosum. [94]<br />
Spirochaetes of the genus Borrelia are a notable exception to this arrangement, with<br />
bacteria such as Borrelia burgdorferi, the cause of Lyme disease, containing a single<br />
linear chromosome. [95] The genes in bacterial genomes are usually a single continuous<br />
stretch of DNA and although several different types of introns do exist in bacteria,<br />
these are much more rare than in eukaryotes. [96]<br />
Bacteria may also contain plasmids, which are small extra-chromosomal DNAs that<br />
may contain genes for antibiotic resistance or virulence factors. Another type of<br />
bacterial DNA are integrated viruses (bacteriophages). Many types of bacteriophage<br />
exist, some simply infect and lyse their host bacteria, while others insert into the<br />
bacterial chromosome. A bacteriophage can contain genes that contribute to its host's<br />
phenotype: for example, in the evolution of Escherichia coli O157:H7 and<br />
Clostridium botulinum, the toxin genes in an integrated phage converted a harmless<br />
ancestral bacteria into a lethal pathogen. [97]<br />
Bacteria, as asexual organisms, inherit identical copies of their parent's genes (i.e.,<br />
they are clonal). However, all bacteria can evolve by selection on changes to their<br />
genetic material DNA caused by genetic recombination or mutations. Mutations come<br />
from errors made during the replication of DNA or from exposure to mutagens.<br />
Mutation rates vary widely among different species of bacteria and even among<br />
different clones of a single species of bacteria. [98] Genetic changes in bacterial<br />
genomes come from either random mutation during replication or "stress-directed<br />
mutation", where genes involved in a particular growth-limiting process have an<br />
increased mutation rate. [99]<br />
Some bacteria also transfer genetic material between cells. This can occur in three<br />
main ways. Firstly, bacteria can take up exogenous DNA from their environment, in a<br />
process called transformation. Genes can also be transferred by the process of<br />
transduction, when the integration of a bacteriophage introduces foreign DNA into the<br />
chromosome. The third method of gene transfer is bacterial conjugation, where DNA<br />
is transferred through direct cell contact. This gene acquisition from other bacteria or
the environment is called horizontal gene transfer and may be common under natural<br />
conditions. [100] Gene transfer is particularly important in antibiotic resistance as it<br />
allows the rapid transfer of resistance genes between different pathogens. [101]<br />
Movement<br />
Further information: Chemotaxis, Flagella, Pilus<br />
The different arrangements of bacterial flagella: A-Monotrichous; B-Lophotrichous;<br />
C-Amphitrichous and D-Peritrichous<br />
Motile bacteria can move using flagella, bacterial gliding, twitching motility or<br />
changes of buoyancy. [102] In twitching motility, bacterial use their type IV pili as a<br />
grappling hook, repeatedly extending it, anchoring it and then retracting it with<br />
remarkable force (>80 pN). [103]<br />
Bacterial species differ in the number and arrangement of flagella on their surface;<br />
some have a single flagellum (monotrichous), a flagellum at each end<br />
(amphitrichous), clusters of flagella at the poles of the cell (lophotrichous), while<br />
others have flagella distributed over the entire surface of the cell (peritrichous). The<br />
bacterial flagella is the best-understood motility structure in any organism and is made<br />
of about 20 proteins, with approximately another 30 proteins required for its<br />
regulation and assembly. [102] The flagellum is a rotating structure driven by a motor at<br />
the base that uses the electrochemical gradient across the membrane for power. This<br />
motor drives the motion of the filament, which acts as a propeller. Many bacteria<br />
(such as E. coli) have two distinct modes of movement: forward movement<br />
(swimming) and tumbling. The tumbling allows them to reorient and makes their<br />
movement a three-dimensional random walk. [104] (See external links below for link to<br />
videos.) The flagella of a unique group of bacteria, the spirochaetes, are found<br />
between two membranes in the periplasmic space. They have a distinctive helical<br />
body that twists about as it moves. [102]<br />
Motile bacteria are attracted or repelled by certain stimuli in behaviors called taxes:<br />
these include chemotaxis, phototaxis and magnetotaxis. [105][106] In one peculiar group,
the myxobacteria, individual bacteria move together to form waves of cells that then<br />
differentiate to form fruiting bodies containing spores. [107] The myxobacteria move<br />
only when on solid surfaces, unlike E. coli which is motile in liquid or solid media.<br />
Several Listeria and Shigella species move inside host cells by usurping the<br />
cytoskeleton, which is normally used to move organelles inside the cell. By promoting<br />
actin polymerization at one pole of their cells, they can form a kind of tail that pushes<br />
them through the host cell's cytoplasm. [108]<br />
Classification and identification<br />
Streptococcus mutans visualized with a Gram stain<br />
Further information: Scientific classification, Systematics and Clinical<br />
pathology<br />
Classification seeks to describe the diversity of bacterial species by naming and<br />
grouping organisms based on similarities. Bacteria can be classified on the basis of<br />
cell structure, cellular metabolism or on differences in cell components such as DNA,<br />
fatty acids, pigments, antigens and quinones. [87] While these schemes allowed the<br />
identification and classification of bacterial strains, it was unclear whether these<br />
differences represented variation between distinct species or between strains of the<br />
same species. This uncertainty was due to the lack of distinctive structures in most<br />
bacteria, as well as lateral gene transfer between unrelated species. [109] Due to lateral<br />
gene transfer, some closely related bacteria can have very different morphologies and<br />
metabolisms. To overcome this uncertainty, modern bacterial classification<br />
emphasizes molecular systematics, using genetic techniques such as guanine cytosine<br />
ratio determination, genome-genome hybridization, as well as sequencing genes that<br />
have not undergone extensive lateral gene transfer, such as the rRNA gene. [110]<br />
Classification of bacteria is determined by publication in the International Journal of<br />
Systematic Bacteriology, [111] and Bergey's Manual of Systematic Bacteriology. [112]<br />
The term "bacteria" was traditionally applied to all microscopic, single-celled<br />
prokaryotes. However, molecular systematics showed prokaryotic life to consist of<br />
two separate domains, originally called Eubacteria and Archaebacteria, but now<br />
called Bacteria and Archaea that evolved independently from an ancient common<br />
ancestor. [113] The archaea and eukaryotes are more closely-related to each other than<br />
either is to the bacteria. These two domains, along with Eukarya, are the basis of the<br />
three-domain system, which is currently the most widely used classification system in<br />
microbiolology. [114] However, due to the relatively recent introduction of molecular<br />
systematics and a rapid increase in the number of genome sequences that are
available, bacterial classification remains a changing and expanding field. [4][115] For<br />
example, a few biologists argue that the Archaea and Eukaryotes evolved from Grampositive<br />
bacteria. [116]<br />
Identification of bacteria in the laboratory is particularly relevant in medicine, where<br />
the correct treatment is determined by the bacterial species causing an infection.<br />
Consequently, the need to identify human pathogens was a major impetus for the<br />
development of techniques to identify bacteria.<br />
Phylogenetic tree showing the diversity of bacteria, compared to other organisms. [117]<br />
Eukaryotes are colored red, archaea green and bacteria blue.<br />
The Gram stain, developed in 1884 by Hans Christian Gram, characterises bacteria<br />
based on the structural characteristics of their cell walls. [57] The thick layers of<br />
peptidoglycan in the "Gram-positive" cell wall stain purple, while the thin "Gramnegative"<br />
cell wall appears pink. By combining morphology and Gram-staining, most<br />
bacteria can be classified as belonging to one of four groups (Gram-positive cocci,<br />
Gram-positive bacilli, Gram-negative cocci and Gram-negative bacilli). Some<br />
organisms are best identified by stains other than the Gram stain, particularly<br />
mycobacteria or Nocardia, which show acid-fastness on Ziehl–Neelsen or similar<br />
stains. [118] Other organisms may need to be identified by their growth in special<br />
media, or by other techniques, such as serology.<br />
Culture techniques are designed to promote the growth and identify particular<br />
bacteria, while restricting the growth of the other bacteria in the sample. Often these<br />
techniques are designed for specific specimens; for example, a sputum sample will be<br />
treated to identify organisms that cause pneumonia, while stool specimens are<br />
cultured on selective media to identify organisms that cause diarrhoea, while<br />
preventing growth of non-pathogenic bacteria. Specimens that are normally sterile,<br />
such as blood, urine or spinal fluid, are cultured under conditions designed to grow all<br />
possible organisms. [119][87] Once a pathogenic organism has been isolated, it can be<br />
further characterised by its morphology, growth patterns such as (aerobic or anaerobic<br />
growth, patterns of hemolysis) and staining.
As with bacterial classification, identification of bacteria is increasingly using<br />
molecular methods. Diagnostics using such DNA-based tools, such as polymerase<br />
chain reaction, are increasingly popular due to their specificity and speed, compared<br />
to culture-based methods. [120] These methods also allow the detection and<br />
identification of "viable but nonculturable" cells that are metabolically active but nondividing.<br />
[121] However, even using these improved methods, the total number of<br />
bacterial species is not known and cannot even be estimated with any certainty.<br />
Attempts to quantify bacterial diversity have ranged from 10 7 to 10 9 total species, but<br />
even these diverse estimates may be out by many orders of magnitude. [122][123]<br />
Interactions with other organisms<br />
Despite their apparent simplicity, bacteria can form complex associations with other<br />
organisms. These symbiotic associations can be divided into parasitism, mutualism<br />
and commensalism. Due to their small size, commensal bacteria are ubiquitous and<br />
grow on animals and plants exactly as they will grow on any other surface. However,<br />
their growth can be increased by warmth and sweat, and large populations of these<br />
organisms in humans are the cause of body odor.<br />
Mutualists<br />
Certain bacteria form close spatial associations that are essential for their survival.<br />
One such mutualistic association, called interspecies hydrogen transfer, occurs<br />
between clusters of anaerobic bacteria that consume organic acids such as butyric acid<br />
or propionic acid and produce hydrogen, and methanogenic Archaea that consume<br />
hydrogen. [124] The bacteria in this association are unable to consume the organic acids<br />
as this reaction produces hydrogen that accumulates in their surroundings. Only the<br />
intimate association with the hydrogen-consuming Archaea keeps the hydrogen<br />
concentration low enough to allow the bacteria to grow.<br />
In soil, microorganisms which reside in the rhizosphere (a zone that includes the root<br />
surface and the soil that adheres to the root after gentle shaking) carry out nitrogen<br />
fixation, converting nitrogen gas to nitrogenous compounds. [125] This serves to<br />
provide an easily absorbable form of nitrogen for many plants, which cannot fix<br />
nitrogen themselves. Many other bacteria are found as symbionts in humans and other<br />
organisms. For example, the presence of over 1,000 bacterial species in the normal<br />
human gut flora of the intestines can contribute to gut immunity, synthesise vitamins<br />
such as folic acid, vitamin K and biotin, convert milk protein to lactic acid (see<br />
Lactobacillus), as well as fermenting complex undigestible carbohydrates. [126][127][128]<br />
The presence of this gut flora also inhibits the growth of potentially pathogenic<br />
bacteria (usually through competitive exclusion) and these beneficial bacteria are<br />
consequently sold as probiotic dietary supplements. [129]<br />
Pathogens<br />
Main article: Pathogenic bacteria
Color-enhanced scanning electron micrograph showing Salmonella typhimurium (red)<br />
invading cultured human cells<br />
If bacteria form a parasitic association with other organisms, they are classed as<br />
pathogens. Pathogenic bacteria are a major cause of human death and disease and<br />
cause infections such as tetanus, typhoid fever, diphtheria, syphilis, cholera,<br />
foodborne illness, leprosy and tuberculosis. A pathogenic cause for a known medical<br />
disease may only be discovered many years after, as was the case with Helicobacter<br />
pylori and peptic ulcer disease. Bacterial diseases are also important in agriculture,<br />
with bacteria causing leaf spot, fire blight and wilts in plants, as well as Johne's<br />
disease, mastitis, salmonella and anthrax in farm animals.<br />
Each species of pathogen has a characteristic spectrum of interactions with its human<br />
hosts. Some organisms, such as Staphylococcus or Streptococcus, can cause skin<br />
infections, pneumonia, meningitis and even overwhelming sepsis, a systemic<br />
inflammatory response producing shock, massive vasodilation and death. [130] Yet<br />
these organisms are also part of the normal human flora and usually exist on the skin<br />
or in the nose without causing any disease at all. Other organisms invariably cause<br />
disease in humans, such as the Rickettsia, which are obligate intracellular parasites<br />
able to grow and reproduce only within the cells of other organisms. One species of<br />
Rickettsia causes typhus, while another causes Rocky Mountain spotted fever.<br />
Chlamydia, another phylum of obligate intracellular parasites, contains species that<br />
can cause pneumonia, or urinary tract infection and may be involved in coronary heart<br />
disease. [131] Finally, some species such as Pseudomonas aeruginosa, Burkholderia<br />
cenocepacia, and Mycobacterium avium are opportunistic pathogens and cause<br />
disease mainly in people suffering from immunosuppression or cystic fibrosis. [132][133]<br />
Bacterial infections may be treated with antibiotics, which are classified as<br />
bacteriocidal if they kill bacteria, or bacteriostatic if they just prevent bacterial<br />
growth. There are many types of antibiotics and each class inhibits a process that is<br />
different in the pathogen from that found in the host. An example of how antibiotics<br />
produce selective toxicity are chloramphenicol and puromycin, which inhibit the<br />
bacterial ribosome, but not the structurally different eukaryotic ribosome. [134]<br />
Antibiotics are used both in treating human disease and in intensive farming to<br />
promote animal growth, where they may be contributing to the rapid development of<br />
antibiotic resistance in bacterial populations. [135] Infections can be prevented by<br />
antiseptic measures such as sterilizating the skin prior to piercing it with the needle of<br />
a syringe, and by proper care of indwelling catheters. Surgical and dental instruments
are also sterilized to prevent contamination and infection by bacteria. Disinfectants<br />
such as bleach are used to kill bacteria or other pathogens on surfaces to prevent<br />
contamination and further reduce the risk of infection.<br />
Significance in technology and industry<br />
Further information: Economic importance of bacteria<br />
Bacteria, often Lactobacillus in combination with yeasts and molds, have been used<br />
for thousands of years in the preparation of fermented foods such as cheese, pickles,<br />
soy sauce, sauerkraut, vinegar, wine and yoghurt. [136][137]<br />
The ability of bacteria to degrade a variety of organic compounds is remarkable and<br />
has been used in waste processing and bioremediation. Bacteria capable of digesting<br />
the hydrocarbons in petroleum are often used to clean up oil spills. [138] Fertilizer was<br />
added to some of the beaches in Prince William Sound in an attempt to promote the<br />
growth of these naturally occurring bacteria after the infamous 1989 Exxon Valdez oil<br />
spill. These efforts were effective on beaches that were not too thickly covered in oil.<br />
Bacteria are also used for the bioremediation of industrial toxic wastes. [139] In the<br />
chemical industry, bacteria are most important in the production of enantiomerically<br />
pure chemicals for use as pharmaceuticals or agrichemicals. [140]<br />
Bacteria can also be used in the place of pesticides in the biological pest control. This<br />
commonly involves Bacillus thuringiensis (also called BT), a Gram-positive, soil<br />
dwelling bacterium. Subspecies of this bacteria are used as a Lepidopteran-specific<br />
insecticides under trade names such as Dipel and Thuricide. [141] Because of their<br />
specificity, these pesticides are regarded as environmentally friendly, with little or no<br />
effect on humans, wildlife, pollinators and most other beneficial insects. [142][143]<br />
Because of their ability to quickly grow and the relative ease with which they can be<br />
manipulated, bacteria are the workhorses for the fields of molecular biology, genetics<br />
and biochemistry. By making mutations in bacterial DNA and examining the resulting<br />
phenotypes, scientists can determine the function of genes, enzymes and metabolic<br />
pathways in bacteria, then apply this knowledge to more complex organisms. [144] This<br />
aim of understanding the biochemistry of a cell reaches its most complex expression<br />
in the synthesis of huge amounts of enzyme kinetic and gene expression data into<br />
mathematical models of entire organisms. This is achievable in some well-studied<br />
bacteria, with models of Escherichia coli metabolism now being produced and<br />
tested. [145][146] This understanding of bacterial metabolism and genetics allows the use<br />
of biotechnology to bioengineer bacteria for the production of therapeutic proteins,<br />
such as insulin, growth factors, or antibodies. [147][148]<br />
See also<br />
• Human flora<br />
• Bioaerosol<br />
• Biotechnology<br />
• Contamination control<br />
• Denitrification
• Desulforudis audaxviator<br />
• Extremophiles<br />
• Transgenic bacteria<br />
References<br />
1. ^ Fredrickson J, Zachara J, Balkwill D, et al (2004). "Geomicrobiology of high-level<br />
nuclear waste-contaminated vadose sediments at the Hanford site, Washington state".<br />
Appl Environ Microbiol 70 (7): 4230–41. PMID 15240306.<br />
2. ^ Whitman W, Coleman D, Wiebe W (1998). "Prokaryotes: the unseen majority".<br />
Proc Natl Acad Sci U S A 95 (12): 6578–83. PMID 9618454.<br />
3. ^ Whitman W, Coleman D, Wiebe W (1998). "Prokaryotes: the unseen majority".<br />
Proc Natl Acad Sci U S A 95 (12): 6578–83. PMID 9618454.<br />
4. ^ a b Rappé MS, Giovannoni SJ (2003). "The uncultured microbial majority". Annu.<br />
Rev. Microbiol. 57: 369-94. doi:10.1146/annurev.micro.57.030502.090759. PMID<br />
14527284.<br />
5. ^ Sears CL (2005). "A dynamic partnership: celebrating our gut flora". Anaerobe 11<br />
(5): 247-51. doi:10.1016/j.anaerobe.2005.05.001. PMID 16701579.<br />
6. ^ 2002 WHO mortality data. Retrieved on 2007-01-20.<br />
7. ^ Ishige T, Honda K, Shimizu S (2005). "Whole organism biocatalysis". Curr Opin<br />
Chem Biol 9 (2): 174–80. PMID 15811802.<br />
8. ^ Woese C, Kandler O, Wheelis M (1990). "Towards a natural system of organisms:<br />
proposal for the domains Archaea, Bacteria, and Eucarya". Proc Natl Acad Sci U S A<br />
87 (12): 4576–9. PMID 2112744.<br />
9. ^ Porter JR (1976). "Antony van Leeuwenhoek: Tercentenary of his discovery of<br />
bacteria". Bacteriological reviews 40 (2): 260-9. PMID 786250. Retrieved on 2007-<br />
08-19.<br />
10. ^ van Leeuwenhoek A (1684). "An abstract of a letter from Mr. Anthony<br />
Leevvenhoek at Delft, dated Sep. 17, 1683, Containing Some Microscopical<br />
Observations, about Animals in the Scurf of the Teeth, the Substance Call'd Worms in<br />
the Nose, the Cuticula Consisting of Scales". Philosophical Transactions (1683–<br />
1775) 14: 568-74. Retrieved on 2007-08-19.<br />
11. ^ van Leeuwenhoek A (1700). "Part of a Letter from Mr Antony van Leeuwenhoek,<br />
concerning the Worms in Sheeps Livers, Gnats, and Animalcula in the Excrements of<br />
Frogs". Philosophical Transactions (1683–1775) 22: 509–18. Retrieved on 2007-08-<br />
19.<br />
12. ^ van Leeuwenhoek A (1702). "Part of a Letter from Mr Antony van Leeuwenhoek,<br />
F. R. S. concerning Green Weeds Growing in Water, and Some Animalcula Found<br />
about Them". Philosophical Transactions (1683–1775) 23: 1304–11. Retrieved on<br />
2007-08-19.<br />
13. ^ Etymology of the word "bacteria". Online Etymology dictionary. Retrieved on<br />
2006-11-23.<br />
14. ^ Pasteur's Papers on the Germ Theory. LSU Law Center's Medical and Public<br />
Health Law Site, Historic Public Health Articles. Retrieved on 2006-11-23.<br />
15. ^ The Nobel Prize in Physiology or Medicine 1905. Nobelprize.org. Retrieved on<br />
2006-11-22.<br />
16. ^ O'Brien S, Goedert J (1996). "HIV causes AIDS: Koch's postulates fulfilled". Curr<br />
Opin Immunol 8 (5): 613–18. PMID 8902385.<br />
17. ^ Thurston A (2000). "Of blood, inflammation and gunshot wounds: the history of<br />
the control of sepsis". Aust N Z J Surg 70 (12): 855-61. PMID 11167573.<br />
18. ^ Schwartz R (2004). "Paul Ehrlich's magic bullets". N Engl J Med 350 (11): 1079–<br />
80. PMID 15014180.<br />
19. ^ Biography of Paul Ehrlich. Nobelprize.org. Retrieved on 2006-11-26.
20. ^ Woese C, Fox G (1977). "Phylogenetic structure of the prokaryotic domain: the<br />
primary kingdoms". Proc Natl Acad Sci U S A 74 (11): 5088–90. PMID 270744.<br />
21. ^ Woese C, Kandler O, Wheelis M (1990). "Towards a natural system of organisms:<br />
proposal for the domains Archaea, Bacteria, and Eucarya". Proc Natl Acad Sci U S A<br />
87 (12): 4576–79. PMID 2112744.<br />
22. ^ Schopf J (1994). "Disparate rates, differing fates: tempo and mode of evolution<br />
changed from the Precambrian to the Phanerozoic". Proc Natl Acad Sci U S A 91<br />
(15): 6735–42. PMID 8041691.<br />
23. ^ DeLong E, Pace N (2001). "Environmental diversity of bacteria and archaea". Syst<br />
Biol 50 (4): 470–78. PMID 12116647.<br />
24. ^ Brown JR, Doolittle WF (1997). "Archaea and the prokaryote-to-eukaryote<br />
transition". Microbiol. Mol. Biol. Rev. 61 (4): 456-502. PMID 9409149.<br />
25. ^ Di Giulio M (2003). "The universal ancestor and the ancestor of bacteria were<br />
hyperthermophiles". J Mol Evol 57 (6): 721–30. PMID 14745541.<br />
26. ^ Battistuzzi F, Feijao A, Hedges S. "A genomic timescale of prokaryote evolution:<br />
insights into the origin of methanogenesis, phototrophy, and the colonization of<br />
land.". BMC Evol Biol 4: 44. PMID 15535883.<br />
27. ^ Poole A, Penny D (2007). "Evaluating hypotheses for the origin of eukaryotes".<br />
Bioessays 29 (1): 74–84. PMID 17187354.<br />
28. ^ Dyall S, Brown M, Johnson P (2004). "Ancient invasions: from endosymbionts to<br />
organelles". Science 304 (5668): 253–7. PMID 15073369.<br />
29. ^ Lang B, Gray M, Burger G. "Mitochondrial genome evolution and the origin of<br />
eukaryotes". Annu Rev Genet 33: 351-97. PMID 10690412.<br />
30. ^ McFadden G (1999). "Endosymbiosis and evolution of the plant cell". Curr Opin<br />
Plant Biol 2 (6): 513-9. PMID 10607659.<br />
31. ^ Schulz H, Jorgensen B. "Big bacteria". Annu Rev Microbiol 55: 105–37. PMID<br />
11544351.<br />
32. ^ Robertson J, Gomersall M, Gill P. (1975). "Mycoplasma hominis: growth,<br />
reproduction, and isolation of small viable cells". J Bacteriol. 124 (2): 1007–18.<br />
PMID 1102522.<br />
33. ^ Fritz I, Strömpl C, Abraham W (2004). "Phylogenetic relationships of the genera<br />
Stella, Labrys and Angulomicrobium within the 'Alphaproteobacteria' and description<br />
of Angulomicrobium amanitiforme sp. nov". Int J Syst Evol Microbiol 54 (Pt 3): 651-<br />
7. PMID 15143003.<br />
34. ^ Cabeen M, Jacobs-Wagner C (2005). "Bacterial cell shape". Nat Rev Microbiol 3<br />
(8): 601–10. PMID 16012516.<br />
35. ^ Young K (2006). "The selective value of bacterial shape". Microbiol Mol Biol Rev<br />
70 (3): 660–703. PMID 16959965.<br />
36. ^ Douwes K, Schmalzbauer E, Linde H, Reisberger E, Fleischer K, Lehn N,<br />
Landthaler M, Vogt T (2003). "Branched filaments no fungus, ovoid bodies no<br />
bacteria: Two unusual cases of mycetoma". J Am Acad Dermatol 49 (2 Suppl Case<br />
Reports): S170–3. PMID 12894113.<br />
37. ^ Donlan R (2002). "Biofilms: microbial life on surfaces". Emerg Infect Dis 8 (9):<br />
881–90. PMID 12194761.<br />
38. ^ Branda S, Vik S, Friedman L, Kolter R (2005). "Biofilms: the matrix revisited".<br />
Trends Microbiol 13 (1): 20–26. PMID 15639628.<br />
39. ^ a b Davey M, O'toole G (2000). "Microbial biofilms: from ecology to molecular<br />
genetics". Microbiol Mol Biol Rev 64 (4): 847–67. PMID 11104821.<br />
40. ^ Donlan RM, Costerton JW (2002). "Biofilms: survival mechanisms of clinically<br />
relevant microorganisms". Clin Microbiol Rev 15 (2): 167–93. PMID 11932229.<br />
41. ^ Shimkets L. "Intercellular signaling during fruiting-body development of<br />
Myxococcus xanthus.". Annu Rev Microbiol 53: 525–49. PMID 10547700.<br />
42. ^ Kaiser D. "Signaling in myxobacteria". Annu Rev Microbiol 58: 75–98. PMID<br />
15487930.
43. ^ Berg JM, Tymoczko JL Stryer L (2002). Molecular Cell Biology, 5th ed., WH<br />
Freeman. ISBN 0-7167-4955-6.<br />
44. ^ Gitai Z (2005). "The new bacterial cell biology: moving parts and subcellular<br />
architecture". Cell 120 (5): 577–86. PMID 15766522.<br />
45. ^ Shih YL, Rothfield L (2006). "The bacterial cytoskeleton". Microbiol. Mol. Biol.<br />
Rev. 70 (3): 729–54. PMID 16959967.<br />
46. ^ Harold F (1972). "Conservation and transformation of energy by bacterial<br />
membranes". Bacteriol Rev 36 (2): 172–230. PMID 4261111.<br />
47. ^ Thanbichler M, Wang S, Shapiro L (2005). "The bacterial nucleoid: a highly<br />
organized and dynamic structure". J Cell Biochem 96 (3): 506–21. PMID<br />
15988757.<br />
48. ^ Poehlsgaard J, Douthwaite S (2005). "The bacterial ribosome as a target for<br />
antibiotics". Nat Rev Microbiol 3 (11): 870–81. PMID 16261170.<br />
49. ^ Fuerst J (2005). "Intracellular compartmentation in planctomycetes". Annu Rev<br />
Microbiol 59: 299–328. PMID 15910279.<br />
50. ^ Yeo M, Chater K (2005). "The interplay of glycogen metabolism and<br />
differentiation provides an insight into the developmental biology of Streptomyces<br />
coelicolor". Microbiology 151 (Pt 3): 855–61. PMID 15758231.<br />
51. ^ Shiba T, Tsutsumi K, Ishige K, Noguchi T (2000). "Inorganic polyphosphate and<br />
polyphosphate kinase: their novel biological functions and applications".<br />
Biochemistry (Mosc) 65 (3): 315–23. PMID 10739474.<br />
52. ^ Brune DC. (1995). "Isolation and characterization of sulfur globule proteins from<br />
Chromatium vinosum and Thiocapsa roseopersicina". Arch Microbiol 163 (6): 391–<br />
99. PMID 7575095.<br />
53. ^ Kadouri D, Jurkevitch E, Okon Y, Castro-Sowinski S. (2005). "Ecological and<br />
agricultural significance of bacterial polyhydroxyalkanoates". Crit Rev Microbiol 31<br />
(2): 55–67. PMID 15986831.<br />
54. ^ Walsby A (1994). "Gas vesicles". Microbiol Rev 58 (1): 94–144. PMID 8177173.<br />
55. ^ van Heijenoort J (2001). "Formation of the glycan chains in the synthesis of<br />
bacterial peptidoglycan". Glycobiology 11 (3): 25R–36R. PMID 11320055.<br />
56. ^ a b Koch A (2003). "Bacterial wall as target for attack: past, present, and future<br />
research". Clin Microbiol Rev 16 (4): 673–87. PMID 14557293.<br />
57. ^ a b Gram, HC (1884). "Über die isolierte Färbung der Schizomyceten in Schnitt- und<br />
Trockenpräparaten". Fortschr. Med. 2: 185–189.<br />
58. ^ Hugenholtz P (2002). "Exploring prokaryotic diversity in the genomic era".<br />
Genome Biol 3 (2): REVIEWS0003. PMID 11864374.<br />
59. ^ Walsh F, Amyes S (2004). "Microbiology and drug resistance mechanisms of fully<br />
resistant pathogens.". Curr Opin Microbiol 7 (5): 439-44. PMID 15451497.<br />
60. ^ Engelhardt H, Peters J (1998). "Structural research on surface layers: a focus on<br />
stability, surface layer homology domains, and surface layer-cell wall interactions". J<br />
Struct Biol 124 (2–3): 276–302. PMID 10049812.<br />
61. ^ Beveridge T, Pouwels P, Sára M, Kotiranta A, Lounatmaa K, Kari K, Kerosuo E,<br />
Haapasalo M, Egelseer E, Schocher I, Sleytr U, Morelli L, Callegari M, Nomellini J,<br />
Bingle W, Smit J, Leibovitz E, Lemaire M, Miras I, Salamitou S, Béguin P, Ohayon<br />
H, Gounon P, Matuschek M, Koval S (1997). "Functions of S-layers". FEMS<br />
Microbiol Rev 20 (1–2): 99–149. PMID 9276929.<br />
62. ^ Kojima S, Blair D. "The bacterial flagellar motor: structure and function of a<br />
complex molecular machine". Int Rev Cytol 233: 93–134. PMID 15037363.<br />
63. ^ Beachey E (1981). "Bacterial adherence: adhesin-receptor interactions mediating<br />
the attachment of bacteria to mucosal surface". J Infect Dis 143 (3): 325–45. PMID<br />
7014727.<br />
64. ^ Silverman P (1997). "Towards a structural biology of bacterial conjugation". Mol<br />
Microbiol 23 (3): 423–9. PMID 9044277.<br />
65. ^ Stokes R, Norris-Jones R, Brooks D, Beveridge T, Doxsee D, Thorson L (2004).<br />
"The glycan-rich outer layer of the cell wall of Mycobacterium tuberculosis acts as an
antiphagocytic capsule limiting the association of the bacterium with macrophages".<br />
Infect Immun 72 (10): 5676–86. PMID 15385466.<br />
66. ^ Daffé M, Etienne G (1999). "The capsule of Mycobacterium tuberculosis and its<br />
implications for pathogenicity". Tuber Lung Dis 79 (3): 153–69. PMID 10656114.<br />
67. ^ Finlay B, Falkow S (1997). "Common themes in microbial pathogenicity revisited".<br />
Microbiol Mol Biol Rev 61 (2): 136–69. PMID 9184008.<br />
68. ^ Nicholson W, Munakata N, Horneck G, Melosh H, Setlow P (2000). "Resistance of<br />
Bacillus endospores to extreme terrestrial and extraterrestrial environments".<br />
Microbiol Mol Biol Rev 64 (3): 548–72. PMID 10974126.<br />
69. ^ Siunov A, Nikitin D, Suzina N, Dmitriev V, Kuzmin N, Duda V. "Phylogenetic<br />
status of Anaerobacter polyendosporus, an anaerobic, polysporogenic bacterium". Int<br />
J Syst Bacteriol 49 Pt 3: 1119–24. PMID 10425769.<br />
70. ^ Nicholson W, Fajardo-Cavazos P, Rebeil R, Slieman T, Riesenman P, Law J, Xue<br />
Y (2002). "Bacterial endospores and their significance in stress resistance". Antonie<br />
Van Leeuwenhoek 81 (1–4): 27–32. PMID 12448702.<br />
71. ^ Vreeland R, Rosenzweig W, Powers D (2000). "Isolation of a 250 million-year-old<br />
halotolerant bacterium from a primary salt crystal". Nature 407 (6806): 897–900.<br />
PMID 11057666.<br />
72. ^ Cano R, Borucki M (1995). "Revival and identification of bacterial spores in 25- to<br />
40-million-year-old Dominican amber". Science 268 (5213): 1060–4. PMID<br />
7538699.<br />
73. ^ Nicholson W, Schuerger A, Setlow P (2005). "The solar UV environment and<br />
bacterial spore UV resistance: considerations for Earth-to-Mars transport by natural<br />
processes and human spaceflight". Mutat Res 571 (1–2): 249–64. PMID 15748651.<br />
74. ^ Hatheway C (1990). "Toxigenic clostridia". Clin Microbiol Rev 3 (1): 66–98. PMID<br />
2404569.<br />
75. ^ Nealson K (1999). "Post-Viking microbiology: new approaches, new data, new<br />
insights". Orig Life Evol Biosph 29 (1): 73–93. PMID 11536899.<br />
76. ^ Xu J (2006). "Microbial ecology in the age of genomics and metagenomics:<br />
concepts, tools, and recent advances". Mol Ecol 15 (7): 1713–31. PMID 16689892.<br />
77. ^ Zillig W (1991). "Comparative biochemistry of Archaea and Bacteria". Curr Opin<br />
Genet Dev 1 (4): 544-51. PMID 1822288.<br />
78. ^ Hellingwerf K, Crielaard W, Hoff W, Matthijs H, Mur L, van Rotterdam B (1994).<br />
"Photobiology of bacteria". Antonie Van Leeuwenhoek 65 (4): 331–47. PMID<br />
7832590.<br />
79. ^ Zumft W (1997). "Cell biology and molecular basis of denitrification". Microbiol<br />
Mol Biol Rev 61 (4): 533–616. PMID 9409151.<br />
80. ^ Drake H, Daniel S, Küsel K, Matthies C, Kuhner C, Braus-Stromeyer S (1997).<br />
"Acetogenic bacteria: what are the in situ consequences of their diverse metabolic<br />
versatilities?". Biofactors 6 (1): 13–24. PMID 9233536.<br />
81. ^ Morel, FMM; Kraepiel AML, Amyot M (1998). "The chemical cycle and<br />
bioaccumulation of mercury". Annual Review of Ecological Systems 29: 543—566.<br />
82. ^ Dalton H (2005). "The Leeuwenhoek Lecture 2000 the natural and unnatural<br />
history of methane-oxidizing bacteria". Philos Trans R Soc Lond B Biol Sci 360<br />
(1458): 1207–22. PMID 16147517.<br />
83. ^ Zehr J, Jenkins B, Short S, Steward G (2003). "Nitrogenase gene diversity and<br />
microbial community structure: a cross-system comparison". Environ Microbiol 5<br />
(7): 539–54. PMID 12823187.<br />
84. ^ Koch A (2002). "Control of the bacterial cell cycle by cytoplasmic growth". Crit<br />
Rev Microbiol 28 (1): 61–77. PMID 12003041.<br />
85. ^ Eagon R. "Pseudomonas natriegens, a marine bacterium with a generation time of<br />
less than 10 minutes". J Bacteriol 83: 736–7. PMID 13888946.<br />
86. ^ Stewart EJ, Madden R, Paul G, Taddei F (2005). "Aging and death in an organism<br />
that reproduces by morphologically symmetric division". PLoS Biol. 3 (2): e45.<br />
PMID 15685293.
87. ^ a b c Thomson R, Bertram H (2001). "Laboratory diagnosis of central nervous<br />
system infections". Infect Dis Clin North Am 15 (4): 1047–71. PMID 11780267.<br />
88. ^ Paerl H, Fulton R, Moisander P, Dyble J. "Harmful freshwater algal blooms, with<br />
an emphasis on cyanobacteria". ScientificWorldJournal 1: 76–113. PMID<br />
12805693.<br />
89. ^ Challis G, Hopwood D. "Synergy and contingency as driving forces for the<br />
evolution of multiple secondary metabolite production by Streptomyces species".<br />
Proc Natl Acad Sci U S A 100 Suppl 2: 14555–61. PMID 12970466.<br />
90. ^ Kooijman S, Auger P, Poggiale J, Kooi B (2003). "Quantitative steps in<br />
symbiogenesis and the evolution of homeostasis". Biol Rev Camb Philos Soc 78 (3):<br />
435–63. PMID 14558592.<br />
91. ^ Prats C, López D, Giró A, Ferrer J, Valls J (2006). "Individual-based modelling of<br />
bacterial cultures to study the microscopic causes of the lag phase". J Theor Biol 241<br />
(4): 939–53. PMID 16524598.<br />
92. ^ Hecker M, Völker U. "General stress response of Bacillus subtilis and other<br />
bacteria". Adv Microb Physiol 44: 35–91. PMID 11407115.<br />
93. ^ Nakabachi A, Yamashita A, Toh H, Ishikawa H, Dunbar H, Moran N, Hattori M<br />
(2006). "The 160-kilobase genome of the bacterial endosymbiont Carsonella".<br />
Science 314 (5797): 267. PMID 17038615.<br />
94. ^ Pradella S, Hans A, Spröer C, Reichenbach H, Gerth K, Beyer S (2002).<br />
"Characterisation, genome size and genetic manipulation of the myxobacterium<br />
Sorangium cellulosum So ce56". Arch Microbiol 178 (6): 484-92. PMID 12420170.<br />
95. ^ Hinnebusch J, Tilly K (1993). "Linear plasmids and chromosomes in bacteria". Mol<br />
Microbiol 10 (5): 917-22. PMID 7934868.<br />
96. ^ Belfort M, Reaban ME, Coetzee T, Dalgaard JZ (1995). "Prokaryotic introns and<br />
inteins: a panoply of form and function". J. Bacteriol. 177 (14): 3897–903. PMID<br />
7608058.<br />
97. ^ Brüssow H, Canchaya C, Hardt W (2004). "Phages and the evolution of bacterial<br />
pathogens: from genomic rearrangements to lysogenic conversion". Microbiol Mol<br />
Biol Rev 68 (3): 560–602. PMID 15353570.<br />
98. ^ Denamur E, Matic I (2006). "Evolution of mutation rates in bacteria". Mol<br />
Microbiol 60 (4): 820–7. PMID 16677295.<br />
99. ^ Wright B (2004). "Stress-directed adaptive mutations and evolution". Mol<br />
Microbiol 52 (3): 643–50. PMID 15101972.<br />
100. ^ Davison J (1999). "Genetic exchange between bacteria in the<br />
environment". Plasmid 42 (2): 73–91. PMID 10489325.<br />
101. ^ Hastings P, Rosenberg S, Slack A (2004). "Antibiotic-induced lateral<br />
transfer of antibiotic resistance". Trends Microbiol 12 (9): 401–4. PMID 15337159.<br />
102. ^ a b c Bardy S, Ng S, Jarrell K (2003). "Prokaryotic motility structures".<br />
Microbiology 149 (Pt 2): 295–304. PMID 12624192.<br />
103. ^ Merz A, So M, Sheetz M (2000). "Pilus retraction powers bacterial<br />
twitching motility". Nature 407 (6800): 98–102. PMID 10993081.<br />
104. ^ Wu M, Roberts J, Kim S, Koch D, DeLisa M (2006). "Collective bacterial<br />
dynamics revealed using a three-dimensional population-scale defocused particle<br />
tracking technique". Appl Environ Microbiol 72 (7): 4987–94. PMID 16820497.<br />
105. ^ Lux R, Shi W (2004). "Chemotaxis-guided movements in bacteria". Crit<br />
Rev Oral Biol Med 15 (4): 207-20. PMID 15284186.<br />
106. ^ Frankel R, Bazylinski D, Johnson M, Taylor B (1997). "Magneto-aerotaxis<br />
in marine coccoid bacteria". Biophys J 73 (2): 994–1000. PMID 9251816.<br />
107. ^ Kaiser D. "Signaling in myxobacteria". Annu Rev Microbiol 58: 75–98.<br />
PMID 15487930.<br />
108. ^ Goldberg MB (2001). "Actin-based motility of intracellular microbial<br />
pathogens". Microbiol Mol Biol Rev 65 (4): 595–626. PMID 11729265.
109. ^ Boucher Y, Douady CJ, Papke RT, Walsh DA, Boudreau ME, Nesbo CL,<br />
Case RJ, Doolittle WF (2003). "Lateral gene transfer and the origins of prokaryotic<br />
groups.". Annu Rev Genet 37: 283–328. PMID 14616063.<br />
110. ^ Olsen G, Woese C, Overbeek R (1994). "The winds of (evolutionary)<br />
change: breathing new life into microbiology". J Bacteriol 176 (1): 1–6. PMID<br />
8282683.<br />
111. ^ IJSEM - Home<br />
112. ^ Bergey's Manual Trust<br />
113. ^ Woese C, Kandler O, Wheelis M (1990). "Towards a natural system of<br />
organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proc Natl<br />
Acad Sci U S A 87 (12): 4576–9. PMID 2112744.<br />
114. ^ Gupta R (2000). "The natural evolutionary relationships among<br />
prokaryotes.". Crit Rev Microbiol 26 (2): 111-31. PMID 10890353.<br />
115. ^ Doolittle RF (2005). "Evolutionary aspects of whole-genome biology".<br />
Curr Opin Struct Biol 15 (3): 248–253. PMID 11837318.<br />
116. ^ Cavalier-Smith T (2002). "The neomuran origin of archaebacteria, the<br />
negibacterial root of the universal tree and bacterial megaclassification.". Int J Syst<br />
Evol Microbiol 52 (Pt 1): 7–76. PMID 11837318.<br />
117. ^ Ciccarelli FD, Doerks T, von Mering C, Creevey CJ, Snel B, Bork P<br />
(2006). "Toward automatic reconstruction of a highly resolved tree of life". Science<br />
311 (5765): 1283-7. PMID 16513982.<br />
118. ^ Woods G, Walker D (1996). "Detection of infection or infectious agents by<br />
use of cytologic and histologic stains". Clin Microbiol Rev 9 (3): 382–404. PMID<br />
8809467.<br />
119. ^ Weinstein M (1994). "Clinical importance of blood cultures". Clin Lab<br />
Med 14 (1): 9–16. PMID 8181237.<br />
120. ^ Louie M, Louie L, Simor AE (2000). "The role of DNA amplification<br />
technology in the diagnosis of infectious diseases". CMAJ 163 (3): 301–309. PMID<br />
10951731.<br />
121. ^ Oliver J. "The viable but nonculturable state in bacteria". J Microbiol 43<br />
Spec No: 93–100. PMID 15765062.<br />
122. ^ Curtis T, Sloan W, Scannell J (2002). "Estimating prokaryotic diversity and<br />
its limits". Proc Natl Acad Sci U S A 99 (16): 10494-9. PMID 12097644.<br />
123. ^ Schloss P, Handelsman J (2004). "Status of the microbial census".<br />
Microbiol Mol Biol Rev 68 (4): 686-91. PMID 15590780.<br />
124. ^ Stams A, de Bok F, Plugge C, van Eekert M, Dolfing J, Schraa G (2006).<br />
"Exocellular electron transfer in anaerobic microbial communities". Environ<br />
Microbiol 8 (3): 371–82. PMID 16478444.<br />
125. ^ Barea J, Pozo M, Azcón R, Azcón-Aguilar C (2005). "Microbial cooperation<br />
in the rhizosphere". J Exp Bot 56 (417): 1761–78. PMID 15911555.<br />
126. ^ O'Hara A, Shanahan F (2006). "The gut flora as a forgotten organ". EMBO<br />
Rep 7 (7): 688–93. PMID 16819463.<br />
127. ^ Zoetendal E, Vaughan E, de Vos W (2006). "A microbial world within us".<br />
Mol Microbiol 59 (6): 1639–50. PMID 16553872.<br />
128. ^ Gorbach S (1990). "Lactic acid bacteria and human health". Ann Med 22<br />
(1): 37–41. PMID 2109988.<br />
129. ^ Salminen S, Gueimonde M, Isolauri E (2005). "Probiotics that modify<br />
disease risk". J Nutr 135 (5): 1294–8. PMID 15867327.<br />
130. ^ Fish D. "Optimal antimicrobial therapy for sepsis". Am J Health Syst<br />
Pharm 59 Suppl 1: S13–9. PMID 11885408.<br />
131. ^ Belland R, Ouellette S, Gieffers J, Byrne G (2004). "Chlamydia<br />
pneumoniae and atherosclerosis". Cell Microbiol 6 (2): 117–27. PMID 14706098.<br />
132. ^ Heise E. "Diseases associated with immunosuppression". Environ Health<br />
Perspect 43: 9–19. PMID 7037390.
133. ^ Saiman, L. "Microbiology of early CF lung disease". Paediatr Respir<br />
Rev.volume=5 Suppl A: S367–369. PMID 14980298<br />
134. ^ Yonath A, Bashan A (2004). "Ribosomal crystallography: initiation,<br />
peptide bond formation, and amino acid polymerization are hampered by antibiotics".<br />
Annu Rev Microbiol 58: 233–51. PMID 15487937.<br />
135. ^ Khachatourians G (1998). "Agricultural use of antibiotics and the evolution<br />
and transfer of antibiotic-resistant bacteria". CMAJ 159 (9): 1129–36. PMID<br />
9835883.<br />
136. ^ Johnson M, Lucey J (2006). "Major technological advances and trends in<br />
cheese". J Dairy Sci 89 (4): 1174–8. PMID 16537950.<br />
137. ^ Hagedorn S, Kaphammer B (1994). "Microbial biocatalysis in the<br />
generation of flavor and fragrance chemicals". Annu. Rev. Microbiol. 48: 773-800.<br />
doi:10.1146/annurev.mi.48.100194.004013. PMID 7826026.<br />
138. ^ Cohen Y (2002). "Bioremediation of oil by marine microbial mats". Int<br />
Microbiol 5 (4): 189–93. PMID 12497184.<br />
139. ^ Neves LC, Miyamura TT, Moraes DA, Penna TC, Converti A (2006).<br />
"Biofiltration methods for the removal of phenolic residues". Appl. Biochem.<br />
Biotechnol. 129-132: 130-52. PMID 16915636.<br />
140. ^ Liese A, Filho M (1999). "Production of fine chemicals using biocatalysis".<br />
Curr Opin Biotechnol 10 (6): 595–603. PMID 10600695.<br />
141. ^ Aronson AI, Shai Y (2001). "Why Bacillus thuringiensis insecticidal toxins<br />
are so effective: unique features of their mode of action". FEMS Microbiol. Lett. 195<br />
(1): 1-8. PMID 11166987.<br />
142. ^ Bozsik A (2006). "Susceptibility of adult Coccinella septempunctata<br />
(Coleoptera: Coccinellidae) to insecticides with different modes of action". Pest<br />
Manag Sci 62 (7): 651–4. PMID 16649191.<br />
143. ^ Chattopadhyay A, Bhatnagar N, Bhatnagar R (2004). "Bacterial insecticidal<br />
toxins". Crit Rev Microbiol 30 (1): 33–54. PMID 15116762.<br />
144. ^ Serres M, Gopal S, Nahum L, Liang P, Gaasterland T, Riley M (2001). "A<br />
functional update of the Escherichia coli K-12 genome". Genome Biol 2 (9):<br />
RESEARCH0035. PMID 11574054.<br />
145. ^ Almaas E, Kovács B, Vicsek T, Oltvai Z, Barabási A (2004). "Global<br />
organization of metabolic fluxes in the bacterium Escherichia coli". Nature 427<br />
(6977): 839–43. PMID 14985762.<br />
146. ^ Reed JL, Vo TD, Schilling CH, Palsson BO (2003). "An expanded<br />
genome-scale model of Escherichia coli K-12 (iJR904 GSM/GPR)". Genome Biol. 4<br />
(9): R54. doi:10.1186/gb-2003-4-9-r54. PMID 12952533.<br />
147. ^ Walsh G (2005). "Therapeutic insulins and their large-scale manufacture".<br />
Appl Microbiol Biotechnol 67 (2): 151–9. PMID 15580495.<br />
148. ^ Graumann K, Premstaller A (2006). "Manufacturing of recombinant<br />
therapeutic proteins in microbial systems". Biotechnol J 1 (2): 164–86. PMID<br />
16892246.<br />
Further reading<br />
• Alcamo IE (2001). Fundamentals of microbiology. Boston: Jones and Bartlett.<br />
ISBN 0-7637-1067-9.<br />
• Atlas RM (1995). Principles of microbiology. St. Louis: Mosby. ISBN 0-<br />
8016-7790-4.<br />
• Martinko JM, Madigan MT (2005). Brock Biology of Microorganisms, 11th<br />
ed., Englewood Cliffs, N.J: Prentice Hall. ISBN 0-13-144329-1.<br />
• Holt JC, Bergey DH (1994). Bergey's manual of determinative bacteriology,<br />
9th ed., Baltimore: Williams & Wilkins. ISBN 0-683-00603-7.
• Hugenholtz P, Goebel BM, Pace NR (1998). "Impact of culture-independent<br />
studies on the emerging phylogenetic view of bacterial diversity". J Bacteriol<br />
180 (18): 4765–74. PMID 9733676.<br />
• Funke BR, Tortora GJ, Case CL (2004). Microbiology: an introduction, 8th<br />
ed,, San Francisco: Benjamin Cummings. ISBN 0-8053-7614-3.<br />
External links<br />
Find more about Bacteria on<br />
Wikipedia's sister projects:<br />
Dictionary definitions<br />
Textbooks<br />
Quotations<br />
Source texts<br />
Images and media<br />
News stories<br />
Learning resources<br />
• Bacterial Nomenclature Up-To-Date from DSMZ<br />
• The largest bacteria<br />
• Tree of Life: Eubacteria<br />
• Videos of bacteria swimming and tumbling, use of optical tweezers and other<br />
videos.<br />
• Planet of the Bacteria by Stephen Jay Gould<br />
• On-line text book on bacteriology<br />
• Animated guide to bacterial cell structure.<br />
Chemotaxis<br />
http://en.wikipedia.org/wiki/Chemotaxis<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search
Chemotaxis, a kind of taxis, is the phenomenon in which bodily cells, bacteria, and<br />
other single-cell or multicellular organisms direct their movements according to<br />
certain chemicals in their environment. This is important for bacteria to find food (for<br />
example, glucose) by swimming towards the highest concentration of food molecules,<br />
or to flee from poisons (for example, phenol). In multicellular organisms, chemotaxis<br />
is critical to early (e.g. movement of sperm towards the egg during fertilization) and<br />
subsequent phases of development (e.g. migration of neurons or lymphocytes) as well<br />
as in normal function. In addition, it has been recognized that mechanisms that allow<br />
chemotaxis in animals can be subverted during cancer metastasis.<br />
Chemotaxis is called positive if movement is in the direction of a higher concentration<br />
of the chemical in question, and negative if the direction is opposite.<br />
Contents<br />
[hide]<br />
• 1 History of chemotaxis research<br />
• 2 Phylogeny and chemotactic signalling<br />
• 3 Bacterial chemotaxis<br />
o 3.1 Behavior<br />
o 3.2 Signal transduction<br />
3.2.1 Flagellum regulation<br />
3.2.2 Receptor regulation<br />
• 4 Eukaryotic chemotaxis<br />
o 4.1 Motility<br />
4.1.1 Chemotaxis related migratory responses<br />
o 4.2 Receptors<br />
4.2.1 Chemotactic selection<br />
o 4.3 Chemotactic ligands<br />
4.3.1 Chemotactic range fitting (CRF)<br />
• 5 Clinical significance<br />
• 6 In the mirror of publications<br />
• 7 Measurement of chemotaxis<br />
• 8 References<br />
• 9 External links<br />
[edit] History of chemotaxis research
Milestones of chemotaxis research<br />
Although migration of cells was detected from the early days of the development of<br />
microscopy (Leeuwenhoek), erudite description of chemotaxis was first made by T.W.<br />
Engelmann (1881) and W.F. Pfeffer (1884) in bacteria and H.S. Jennings (1906) in<br />
ciliates. The Nobel Prize Laureate E. Metchnikoff also contributed to the study of the<br />
field with investigations of the process as an initial step of phagocytosis. The<br />
significance of chemotaxis in biology and clinical pathology was widely accepted in<br />
the 1930s. The most fundamental definitions belonging to the phenomenon were also<br />
drafted by this time. The most important aspects in quality control of chemotaxis<br />
assays were described by H. Harris in the 1950s. In the 1960s and 1970s, the<br />
revolution of modern cell biology and biochemistry provided a series of novel<br />
techniques which became available to investigate the migratory responder cells and<br />
subcellular fractions responsible for chemotactic activity. The pioneering works of J.<br />
Adler represented a significant turning point in understanding the whole process of<br />
intracellular signal transduction of bacteria. [1]<br />
On November 3, 2006, Dr. Dennis Bray of University of Cambridge was awarded the<br />
Microsoft European Science Award for his work on chemotaxis on E. coli. [2][3]<br />
[edit] Phylogeny and chemotactic signalling<br />
Chemotaxis is one of the most basic cell physiological responses. Development of<br />
receptor systems for the detection of harmful and favorable substances in the<br />
environment was most essential to unicellular organisms from the very early stages of<br />
phylogeny. Comprehensive analysis of chemotactic activity of the eukaryotic<br />
protozoon Tetrahymena pyriformis and consensus sequences of appearance of amino<br />
acids in the primordial soup suggest that there was a good correlation between the<br />
chemotactic character of these relative simple organic molecules and their<br />
development on the Earth. In this way the earliest molecules are suggested to be
highly chemoattractant (e.g. Gly, Glu, Pro), while latter ones are thought to be<br />
strongly chemorepellent (e.g. Tyr, Trp, Phe) amino acids. [4]<br />
[edit] Bacterial chemotaxis<br />
Some bacteria, such as E. coli, have several flagella per cell (4–10 typically). These<br />
can rotate in two ways :<br />
1. Counter-clockwise rotation aligns the flagella into a single rotating bundle,<br />
causing the bacterium to swim in a straight line.<br />
2. Clockwise rotation breaks the flagella bundle apart such that each flagellum<br />
points in a different direction, causing the bacterium to tumble in place.<br />
The directions of rotation are given for an observer outside the cell looking down the<br />
flagella toward the cell.<br />
[edit] Behavior<br />
The overall movement of a bacterium is the result of alternating tumble and swim<br />
phases. If one watches a bacterium swimming in a uniform environment, its<br />
movement will look like a random walk with relatively straight swims interrupted by<br />
random tumbles that reorient the bacterium. Bacteria such as E. coli are unable to<br />
choose the direction in which they swim, and are unable to swim in a straight line for<br />
more than a few seconds due to rotational diffusion. In other words, bacteria "forget"<br />
the direction in which they are going. Given these limitations, it is remarkable that<br />
bacteria can direct their motion to find favorable locations with high concentrations of<br />
attractants (usually food) and avoid repellents (usually poisons).<br />
In the presence of a chemical gradient bacteria will chemotax, or direct their overall<br />
motion based on the gradient. If the bacterium senses that it is moving in the correct<br />
direction (toward attractant/away from repellent), it will keep swimming in a straight<br />
line for a longer time before tumbling. If it is moving in the wrong direction, it will<br />
tumble sooner and try a new direction at random. In other words, bacteria like E. coli<br />
use temporal sensing to decide whether life is getting better or worse. In this way, it
finds the location with the highest concentration of attractant (usually the source)<br />
quite well. Even under very high concentrations, it can still distinguish very small<br />
differences in concentration. Fleeing from a repellent works with the same efficiency.<br />
It seems remarkable that this purposeful random walk is a result of simply choosing<br />
between two methods of random movement; namely tumbling and straight swimming.<br />
In fact, chemotactic responses such as forgetting direction and choosing movements<br />
resemble the decision-making abilities of higher lifeforms with brains that process<br />
sensory data.<br />
The helical nature of the individual flagellar filament is critical for this movement to<br />
occur. As such, the protein that makes up the flagellar filament, flagellin, is quite<br />
similar among all flagellated bacteria. Vertebrates seem to have taken advantage of<br />
this fact by possessing an immune receptor (TLR5) designed to recognize this<br />
conserved protein.<br />
As in many instances in biology, there are bacteria that do not follow this rule. Many<br />
bacteria, such as Vibrio, are monoflagellated and have a single flagellum at one pole<br />
of the cell. Their method of chemotaxis is different. Others possess a single flagellum<br />
that is kept inside the cell wall. These bacteria move by spinning the whole cell,<br />
which is shaped like a corkscrew. [5]<br />
[edit] Signal transduction<br />
Chemical gradients are sensed through multiple transmembrane receptors, called<br />
methyl accepting chemotaxis proteins (MCPs), which vary in the molecules that they<br />
detect. These receptors may bind attractants or repellents directly or indirectly through<br />
interaction with proteins of periplasmatic space. The signals from these receptors are<br />
transmitted across the plasma membrane into the cytosol, where Che proteins are<br />
activated. The Che proteins alter the tumbling frequency, and alter the receptors.<br />
[edit] Flagellum regulation<br />
The proteins CheW and CheA bind to the receptor. The activation of the receptor by<br />
an external stimulus causes autophosphorylation in the histidine kinase, CheA, at a
single highly conserved histidine residue. CheA in turn transfers phosphoryl groups to<br />
conserved aspartate residues in the response regulators CheB and CheY [ note: CheA<br />
is a histidine kinase and it does not actively transfer the phosphoryl group. The<br />
response regulator CheB takes the phosphoryl group from CheA]. This mechanism of<br />
signal transduction is called a 'Two Component System' and is a common form of<br />
signal transduction in bacteria. CheY induces tumbling by interacting with the<br />
flagellar switch protein FliM, inducing a change from counter-clockwise to clockwise<br />
rotation of the flagellum. Change in the rotation state of a single flagellum can disrupt<br />
the entire flagella bundle and cause a tumble.<br />
[edit] Receptor regulation<br />
CheB, when activated by CheA, acts as a methylesterase, removing methyl groups<br />
from glutamate residues on the cytosolic side of the receptor. It works antagonistically<br />
with CheR, a methyltransferase, which adds methyl residues to the same glutamate<br />
residues. The more methyl residues are attached to the receptor, the more sensitive the<br />
receptor. As the signal from the receptor induces demethylation of the receptor in a<br />
feedback loop, the system is continuously adjusted to environmental chemical levels,<br />
remaining sensitive for small changes even under extreme chemical concentrations.<br />
This regulation allows the bacterium to 'remember' chemical concentrations from the<br />
recent past and compare them to those it is currently experiencing, thus 'know'<br />
whether it is traveling up or down a gradient. However, the methylation system alone<br />
cannot account for the wide range of sensitivity that bacteria have to chemical<br />
gradients. Additional regulatory mechanisms such as receptor clustering and receptorreceptor<br />
interactions also modulate the signalling pathway.<br />
http://en.wikipedia.org/wiki/Coccidia<br />
Coccidia<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
Coccidia<br />
Coccidia oocysts<br />
Scientific classification<br />
Kingdom: Protista
Phylum: Apicomplexa<br />
Class: Conoidasida<br />
Subclass: Coccidiasina<br />
Order: Eucoccidiorida<br />
Suborder, Family, Genera & Species<br />
Adeleorina<br />
• Adeleidae<br />
• Dactylosomatidae<br />
• Haemogregarinidae<br />
• Hepatozoidae<br />
o Hepatozoon<br />
• Karyolysidae<br />
• Klossiellidae<br />
• Legerellidae<br />
Eimeriorina<br />
• Aggregatidae<br />
o Aggregata<br />
o Merocystis<br />
o Selysina<br />
• Calyptosporiidae<br />
o Calyptospora<br />
• Cryptosporidiidae<br />
o Cryptosporidium<br />
• Eimeriidae<br />
o Atoxoplasma<br />
o Barrouxia<br />
o Caryospora<br />
o Caryotropha<br />
o Cyclospora<br />
o Diaspora<br />
o Dorisa<br />
o Dorisiella<br />
o Eimeria<br />
o Grasseella<br />
o Isospora<br />
o Mantonella<br />
o Ovivora<br />
o Pfeifferinella<br />
o Pseudoklossia<br />
o Tyzzeria<br />
o Wenyonella<br />
• Elleipsisomatidae<br />
o Elleipsisoma<br />
• Lankesterellidae
o Lankesterella<br />
o Schellackia<br />
• Sarcocystidae<br />
o Sarcocystinae<br />
Frenkelia<br />
Sarcocystis<br />
o Toxoplasmatinae<br />
Besnoitia<br />
Hammondia<br />
Neospora<br />
Toxoplasma<br />
• Selenococcidiidae<br />
o Selenococcidium<br />
• Spirocystidae<br />
o Spirocystis<br />
Coccidia are microscopic, spore-forming, single-celled parasites belonging to the<br />
apicomplexan class Conoidasida. [1] Coccidian parasites infect the intestinal tracts of<br />
animals [2] , and are the largest group of apicomplexan protozoa.<br />
Coccidia are obligate, intracellular parasites, which means that they must live and<br />
reproduce within an animal cell.<br />
Contents<br />
[hide]<br />
• 1 Coccidiosis<br />
o 1.1 Coccidia in dogs<br />
• 2 Genera and species that cause coccidiosis<br />
• 3 References<br />
• 4 See also<br />
• 5 External links<br />
[edit] Coccidiosis<br />
Coccidiosis is the disease caused by coccidian infection. Coccidiosis is a parasitic<br />
disease of the intestinal tract of animals, caused by coccidian protozoa. The disease<br />
spreads from one animal to another by contact with infected feces, or ingestion of<br />
infected tissue. Diarrhea, which may become bloody in severe cases, is the primary<br />
symptom. Most animals infected with coccidia are asymptomatic; however, young or<br />
immuno-compromised animals may suffer severe symptoms, including death.<br />
While coccidian organisms can infect a wide variety of animals, including humans,<br />
birds, and livestock, they are usually species-specific. One well-known exception is<br />
toxoplasmosis, caused by Toxoplasma gondii.
[edit] Coccidia in dogs<br />
People often first encounter coccidia when they acquire a young puppy who is<br />
infected. The infectious organisms are canine-specific and are not contagious to<br />
humans (compare to zoonotic diseases).<br />
Young puppies are frequently infected with coccidia and often develop active<br />
Coccidiosis -- even puppies obtained from diligent professional breeders. Infected<br />
puppies almost always have received the parasite from their mother's feces. Typically,<br />
healthy adult animals shedding the parasite's oocysts in their feces will be<br />
asymptomatic because of their developed immune systems. However, undeveloped<br />
immune systems make puppies more susceptible. Further, stressors such as new<br />
owners, travel, weather changes, and unsanitary conditions are believed to activate<br />
infections in susceptible animals.<br />
Symptoms in young dogs are universal: at some point around 2-3 months of age, an<br />
infected dog develops persistently loose stools. This diarrhea proceeds to stool<br />
containing liquid, thick mucus, and light colored fecal matter. As the infection<br />
progresses, spots of blood may become apparent in the stool, and sudden bowel<br />
movements may surprise both dog and owner alike. Coccidia infection is so common<br />
that any pup under 4 months old with these symptoms can almost surely be assumed<br />
to have coccidiosis.<br />
Fortunately, the treatment is inexpensive, extremely effective, and routine. A<br />
veterinarian can easily diagnose the disease through low-powered microscopic<br />
examination of an affected dog's feces, which usually will be replete with oocysts.<br />
One of many easily administered and inexpensive drugs will be prescribed, and, in the<br />
course of just a few days, an infection will be eliminated or perhaps reduced to such a<br />
level that the dog's immune system can make its own progress against the infection.<br />
Even when an infection has progressed sufficiently that blood is present in feces,<br />
permanent damage to the gastrointestinal system is rare, and the dog will most likely<br />
make a complete recovery without long-lasting negative effects.<br />
If one dog of a litter has coccidiosis, then most certainly all dogs at a breeder's<br />
kennels have active coccidia infections. Breeders should be notified if a newlyacquired<br />
pup is discovered to be infected with coccidia. Breeders can take steps to<br />
eradicate the organism from their kennels, including applying medications in bulk to<br />
an entire facility.<br />
[edit] Genera and species that cause coccidiosis<br />
• Genus Isospora is the most common cause of intestinal coccidiosis in dogs<br />
and cats and is usually what is meant by coccidiosis. Species of Isospora are<br />
species specific, meaning they only infects one type of species. Species that<br />
infect dogs include I. canis, I. ohioensis, I. burrowsi, and I. neorivolta. Species<br />
that infect cats include I. felis and I. rivolta. The most common symptom is<br />
diarrhea. Sulfonamides are the most common treatment. [3]<br />
• Genus Cryptosporidium contains two species known to cause<br />
cryptosporidiosis, C. parvum and C. muris. Cattle are most commonly affected
y Cryptosporidium, and their feces are often assumed to be a source of<br />
infection for other mammals including humans. Recent genetic analyses of<br />
Cryptosporidium in humans have identified Cryptosporidium hominis as a<br />
new species specific for humans. Infection occurs most commonly in<br />
individuals that are immunocompromised, e.g. dogs with canine distemper,<br />
cats with feline leukemia virus infection, and humans with AIDS. Very young<br />
puppies and kittens can also become infected with Cryptosporidium, but the<br />
infection is usually eliminated without treatment. [3]<br />
• Genus Hammondia is transmitted by ingestion of cysts found in the tissue of<br />
grazing animals and rodents. Dogs and cats are the definitive hosts, with the<br />
species H. heydorni infecting dogs and the species H. hammondi and H.<br />
pardalis infecting cats. Hammondia usually does not cause disease. [3]<br />
• Genus Besnoitia infect cats that ingest cysts found in the tissue of rodents and<br />
opossum, but usually does not cause disease. [3]<br />
• Genus Sarcocystis infect carnivores that ingest cysts from various intermediate<br />
hosts. It is possible for Sarcocystis to cause disease in dogs and cats. [3]<br />
• Genus Toxoplasma has one important species, Toxoplasma gondii. Cats are<br />
the definitive host, but all mammals and some fish, reptiles, and amphibians<br />
can be intermediate hosts. Therefore, only cat feces will hold infective<br />
oocysts, but infection through ingestion of cysts can occur with the tissue of<br />
any intermediate host. Toxoplasmosis occurs in humans usually as low-grade<br />
fever or muscle pain for a few days. A normal immune system will suppress<br />
the infection but the tissue cysts will persist in that animal or human for years<br />
or the rest of its life. In immunocompromised individuals, those dormant cysts<br />
can be reactivated and cause many lesions in the brain, heart, lungs, eyes, etc.<br />
Without a competent immune system, the animal or human will most likely<br />
die from the infection. For pregnant women, the fetus is at risk if the pregnant<br />
woman becomes infected for the first time during pregnancy. If the woman<br />
had been infected during childhood or adolescence, she will have an immunity<br />
that will protect her developing fetus during pregnancy. The most important<br />
misconception about the transmission of toxoplasmosis comes from statements<br />
like 'ingestion of raw or undercooked meat, or cat feces.' Kitchen hygiene is<br />
much more important because people do tend to taste marinades or sauces<br />
before being cooked, or chop meat then vegetables without properly cleaning<br />
the knife and cutting board. Many physicians mistakenly put panic in their<br />
pregnant clients and advise them to get rid of their cat without really warning<br />
them of the likely sources of infection. Adult cats are very unlikely to shed<br />
infective oocysts. Symptoms in cats include fever, weight loss, diarrhea,<br />
vomiting, uveitis, and central nervous system signs. Disease in dogs includes a<br />
rapidly progressive form seen in dogs also infected with distemper, and a<br />
neurological form causing paralysis, tremors, and seizures. Dogs and cats are<br />
usually treated with clindamycin. [3]<br />
• Genus Neospora has one important species, Neospora caninum, that affects<br />
dogs in a manner similar to toxoplasmosis. Neosporosis is difficult to treat. [3]<br />
• Genus Hepatozoon contains one species that causes hepatozoonosis in dogs<br />
and cats, Hepatozoon canis. Animals become infected by ingesting an infected<br />
Rhipicephalus sanguineus, also known as the brown dog tick. Symptoms<br />
include fever, weight loss, and pain of the spine and limbs.
The most common medications used to treat coccidian infections are in the<br />
sulphonamide family. Although unusual, sulphonamides can damage the tear glands<br />
in some dogs, causing keratoconjunctivitis sicca, or "dry eye", which may have a lifelong<br />
impact. Some veterinarians recommend measuring tear production prior to<br />
sulphonamide administration, and at various intervals after administration. Other<br />
veterinarians will simply avoid using sulphonamides, instead choosing another<br />
product effective against coccidia.<br />
Left untreated, the infection may clear of its own accord, or in some cases may<br />
continue to ravage an animal and cause permanent damage or, occasionally, death.<br />
[edit] References<br />
1. ^ The Taxonomicon & Systema Naturae (Website database). Taxon: Genus<br />
Cryptosporidium. Universal Taxonomic Services, Amsterdam, The<br />
Netherlands (2000).<br />
2. ^ Biodiversity explorer: Apicomplexa (apicomplexans, sporozoans). Iziko<br />
Museums of Cape Town.<br />
3. ^ a b c d e f g Ettinger, Stephen J.; Feldman, Edward C. (1995). Textbook of<br />
Veterinary Internal Medicine, 4th ed., W.B. Saunders Company. ISBN 0-<br />
7216-6795-3.<br />
[edit] See also<br />
• cryptosporidiosis<br />
• Zoalene is a fodder additive for poultry, used to prevent infections from<br />
coccidia.<br />
[edit] External links<br />
• Mar Vista Animal Medical Center.<br />
• The Coccidia of the World, Donald W. Duszynski, Steve J. Upton, Lee Couch,<br />
Feb. 21, 2004.<br />
• Life Cycle EIMERIA, Andreas Weck-Heimann, 1996-2005<br />
• FarmingUK, Information about Coccidiosis<br />
• Lillehoj, Hyun S. (October 1996). "Two Strategies for Protecting Poultry<br />
From Coccidia". Agricultural Research magazine (October 1996). United<br />
States Department of Agriculture: Agrigultural Research Service. Describes<br />
using live-parasite vaccine versus a monoclonal antibody to block the<br />
sporozoite from invading a host's cell.<br />
Retrieved from "http://en.wikipedia.org/wiki/Coccidia"<br />
Categories: Apicomplexa | Dog diseases | Cat diseases | Animal diseases | Veterinary<br />
protozoology
http://en.wikipedia.org/wiki/Deer_Island_Waste_Water_Treatment_Plant<br />
http://ludb.clui.org/ex/i/MA3134/<br />
Deer Island Waste Water Treatment<br />
Plant<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
The Deer Island Waste Water Treatment Plant (also known as Deer Island Sewage<br />
Treatment Plant) run and operated by The Massachusetts Water Resources Authority<br />
is located on Deer Island, one of the Boston Harbor Islands in Boston Harbor.<br />
[1] [2] [3]<br />
It is the second largest sewage treatment plant in the United States.<br />
It is a key part of the program to protect Boston Harbor against pollution from sewer<br />
systems in eastern Massachusetts.<br />
The plant removes human, household, business and industrial pollutants from<br />
wastewater that originates in homes and businesses in forty three greater Boston<br />
communities. It complies with all federal and state environmental standards and<br />
subject to the discharge permit issued for it by EPA and DEP. Its treated wastewater<br />
can safely be released into the marine environment.<br />
It has an array of 150 foot tall egg-like sludge digesters and these are major harbor<br />
landmarks. [4][5]<br />
[edit] Notes<br />
1. ^ Deer Island Sewage Treatment Plant. The Center for Land Use<br />
Interpretation.<br />
2. ^ Jardine Water Purification Plant article<br />
3. ^ 1867 "The First Tunnel Under the Lake". Chicago Public Library.<br />
4. ^ Islands You Can Visit - Deer Island. Boston Harbor Islands Partnership.<br />
Retrieved on August 21, 2006.<br />
5. ^ Deer Island Factsheet. Boston Harbor Islands Partnership. Retrieved on<br />
August 21, 2006.<br />
[edit] Bibliography<br />
• Baldwin, Sandy, "Boston Harbor Pipe Dreams Come True!": USGS Visits the<br />
Deer Island Sewage Treatment Plant and a Cleaner Harbor, USGS Sound<br />
Waves, April 2006.
[edit] External links<br />
• MWRA article on The Deer Island Sewage Treatment Plant<br />
• A History of the sewer system in Boston<br />
Retrieved from<br />
"http://en.wikipedia.org/wiki/Deer_Island_Waste_Water_Treatment_Plant"<br />
Categories: Buildings and structures in Boston, Massachusetts | Sewage treatment<br />
plants<br />
Views<br />
• Article<br />
• Discussion<br />
• Edit this page<br />
• History<br />
Personal tools<br />
• Log in / create account<br />
Navigation<br />
• Main Page<br />
• Contents<br />
• Featured content<br />
• Current events<br />
• Random article<br />
Interaction<br />
Search<br />
• About Wikipedia<br />
• Community portal<br />
• Recent changes<br />
• Contact Wikipedia<br />
• Donate to Wikipedia<br />
• Help<br />
Toolbox<br />
• What links here<br />
• Related changes<br />
• Upload file
• Special pages<br />
• Printable version<br />
• Permanent link<br />
• Cite this page<br />
• This page was last modified on 19 October 2007, at 15:11.<br />
• All text is available under the terms of the GNU Free Documentation License.<br />
(See Copyrights for details.)<br />
Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc., a<br />
U.S. registered 501(c)(3) tax-deductible nonprofit charity.<br />
• Privacy policy<br />
• About Wikipedia<br />
• Disclaimers<br />
http://en.wikipedia.org/wiki/Flagellate<br />
Flagellate<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
"Flagellata" from Ernst Haeckel's Artforms of Nature, 1904
Parasitic excavate (Giardia lamblia)<br />
Green alga (Chlamydomonas)<br />
Flagellates are cells with one or more whip-like organelles called flagella. Some cells<br />
in animals may be flagellate, for instance the spermatozoa of most phyla. Higher<br />
plants and fungi do not produce flagellate cells, but the closely related green algae and<br />
chytrids do. Many protists take the form of single-celled flagellates.<br />
[edit] Form and behavior<br />
Flagellates r protozoans (animal-like protists). Eukaryotic flagella are supported by<br />
microtubules in a characteristic arrangement, with nine fused pairs surrounding two<br />
central singlets. These arise from a basal body or kinetosome, with microtubule roots<br />
that are an important part of the cell's brain. In some, for instance, they support a<br />
cytostome or mouth, where food is ingested. The flagella often support hairs, called<br />
mastigonemes, or contain rods. Their ultrastructure plays an important role in<br />
classifying eukaryotes.<br />
In protists and microscopic animals, flagella are generally used for propulsion. They<br />
may also be used to create a current that brings in food. In most things, one or more<br />
flagella are located at or near the anterior of the cell eg Euglena. Often there is one<br />
directed forwards and one trailing behind. Among animals, fungi, and Choanozoa,<br />
which make up a group called the opisthokonts, there is a single posterior flagellum.<br />
They are from the phylum Mastigophora. They can cause diseases and they can make<br />
their own food. For example, Trypanosome which causes the African sleeping<br />
sickness.<br />
[edit] Groups of flagellates<br />
Originally the flagellated protozoa were treated as a single class of phylum, the<br />
Mastigophora. This was divided into the Phytomastigina or phytoflagellates, which<br />
have chloroplasts or are closely related to such forms, and the Zoomastigina or<br />
zooflagellates, which do not. Most phytoflagellates were given a separate<br />
classification by botanists, treating them in several divisions of algae.<br />
This scheme has generally been abandoned or is retained only for convenience.<br />
However, the relationships among the flagellates are still mostly unknown, and their
higher classification is confused. Some argue that the Linnaean ranks are not<br />
appropriate for such a diverse set of organisms.<br />
Phytoflagellates are found in most groups of algae. Both the green algae and<br />
heterokonts include a variety of flagellates in addition to non-motile and multicellular<br />
forms. The dinoflagellates, cryptomonads, haptophytes, and euglenids are almost<br />
entirely single-celled flagellates.<br />
Many of the other flagellates make up what are called the excavate taxa. These<br />
include the euglenids and a number of important parasites, such as trypanosomes and<br />
Giardia. The excavates generally show similarities in the structure of their flagella<br />
and typically have a cytostome. However, they may be a paraphyletic group, and in<br />
particular may have been ancestral to most or all other eukaryotes. Aprils fools day<br />
yaeaahhhhh<br />
Other notable groups including flagellates are the Cercozoa, alveolates (including<br />
dinoflagellates), ebriids, and Apusozoa.<br />
[edit] External links<br />
• MeSH Flagellata<br />
Retrieved from "http://en.wikipedia.org/wiki/Flagellate"<br />
http://en.wikipedia.org/wiki/Fungus#With_plants<br />
accessed 03/04/08<br />
Fungus<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
For the fictional character, see Fungus the Bogeyman. For the music genre, see<br />
Fungi (music).<br />
Fungi<br />
Fossil range: Early Silurian - Recent
Clockwise from top left: Amanita muscaria, a<br />
basidiomycete; Sarcoscypha coccinea, an<br />
ascomycete; black bread mold, a zygomycete; a<br />
chytrid; a Penicillium conidiophore.<br />
Scientific classification<br />
Domain: Eukarya<br />
(unranked) Opisthokonta<br />
Kingdom: Fungi<br />
(L., 1753) R.T. Moore, 1980 [1]<br />
Subkingdoms/Phyla<br />
Chytridiomycota<br />
Blastocladiomycota<br />
Neocallimastigomycota<br />
Glomeromycota<br />
Zygomycota<br />
Dikarya (inc. Deuteromycota)<br />
Ascomycota<br />
Basidiomycota<br />
A fungus (pronounced / f ŋgәs/) is any eukaryotic organism that is a member of the<br />
kingdom Fungi (pronounced / f nd a /). [2] The fungi are heterotrophic organisms<br />
characterized by a chitinous cell wall, and in the majority of species, filamentous<br />
growth as multicellular hyphae forming a mycelium; some fungal species also grow<br />
as single cells. Sexual and asexual reproduction is commonly via spores, often<br />
produced on specialized structures or in fruiting bodies. Some fungal species have lost<br />
the ability to form specialized reproductive structures, and propagate solely by<br />
vegetative growth. Yeasts, molds, and mushrooms are examples of fungi. The fungi<br />
are a monophyletic group that is phylogenetically clearly distinct from the<br />
morphologically similar slime molds (myxomycetes) and water molds (oomycetes).<br />
The fungi are more closely related to animals than plants, yet the discipline of biology<br />
devoted to the study of fungi, known as mycology, often falls under a branch of<br />
botany.
Occurring worldwide, most fungi are largely invisible to the naked eye, living for the<br />
most part in soil, dead matter and as symbionts of plants, animals, or other fungi.<br />
They perform an essential role in all ecosystems in decomposing matter and are<br />
indispensable in nutrient cycling and exchange. Some fungi become noticeable when<br />
fruiting, either as mushrooms or molds. Many fungal species have long been used as a<br />
direct source of food, such as mushrooms and truffles and in fermentation of various<br />
food products, such as wine, beer, and soy sauce. More recently, fungi are being used<br />
as sources for antibiotics and various enzymes, such as cellulases, pectinases, and<br />
proteases, important for industrial use or as active ingredients of detergents. Many<br />
fungi produce bioactive compounds, such as alkaloids and polyketides that are toxic<br />
to animals including humans and are, therefore, called mycotoxins. Some fungi are<br />
used recreationally or in traditional ceremonies as a source of psychotropic<br />
compounds. Several species of the fungi are significant pathogens of humans and<br />
other animals, and losses due to diseases of crops (e.g., rice blast disease) or food<br />
spoilage caused by fungi can have a large impact on human food supply and local<br />
economies.<br />
Contents<br />
[hide]<br />
• 1 Etymology and definition<br />
• 2 Diversity<br />
• 3 Importance for human use<br />
o 3.1 Cultured foods<br />
o 3.2 Other human uses<br />
o 3.3 Mycotoxins<br />
o 3.4 Edible and poisonous fungi<br />
o 3.5 Fungi in the biological control of pests<br />
• 4 Ecology<br />
o 4.1 Symbiosis<br />
4.1.1 With plants<br />
4.1.2 With insects<br />
4.1.3 As pathogens and parasites<br />
o 4.2 Nutrition and possible autotrophy<br />
• 5 Morphology<br />
o 5.1 Microscopic structures<br />
o 5.2 Macroscopic structures<br />
o 5.3 Morphological and physiological features for substrate penetration<br />
• 6 Reproduction<br />
o 6.1 Asexual reproduction<br />
o 6.2 Sexual reproduction<br />
o 6.3 Spore dispersal<br />
o 6.4 Other sexual processes<br />
• 7 Phylogeny and classification<br />
o 7.1 Physiological and morphological traits<br />
o 7.2 Evolutionary history<br />
7.2.1 Cladogram<br />
o 7.3 The taxonomic groups of fungi
o 7.4 Phylogenetic relationships with other fungus-like organisms<br />
• 8 See also<br />
• 9 Notes and references<br />
• 10 Further reading<br />
• 11 External links<br />
Etymology and definition<br />
The English word fungus is directly adopted from the Latin fungus, meaning<br />
"mushroom", used in Horace and Pliny. [3] This in turn is derived from the Greek word<br />
sphongos/σφογγος ("sponge"), referring to the macroscopic structures and<br />
morphology of some mushrooms and molds and also used in other languages (e.g., the<br />
German Schwamm ("sponge") or Schwammerl for some types of mushroom).<br />
Diversity<br />
Fungi have a worldwide distribution, and grow in a wide range of habitats, including<br />
deserts. Most fungi grow in terrestrial environments, but several species occur only in<br />
aquatic habitats. Fungi along with bacteria are the primary decomposers of organic<br />
matter in most if not all terrestrial ecosystems worldwide. Based on observations of<br />
the ratio of the number of fungal species to the number of plant species in some<br />
environments, the fungal kingdom has been estimated to contain about 1.5 million<br />
species. [4] Around 70,000 fungal species have been formally described by<br />
taxonomists, but the true dimension of fungal diversity is still unknown. [5] Most fungi<br />
grow as thread-like filaments called hyphae, which form a mycelium, while others<br />
grow as single cells. [6][7] Until recently many fungal species were described based<br />
mainly on morphological characteristics, such as the size and shape of spores or<br />
fruiting structures, and biological species concepts; the application of molecular tools,<br />
such as DNA sequencing, to study fungal diversity has greatly enhanced the<br />
resolution and added robustness to estimates of diversity within various taxonomic<br />
groups. [8]<br />
Importance for human use<br />
Sacharomyces cerevisiae cells in DIC microscopy.<br />
Human use of fungi for food preparation or preservation and other purposes is<br />
extensive and has a long history: yeasts are required for fermentation of beer, wine [9]
and bread, some other fungal species are used in the production of soy sauce and<br />
tempeh. Mushroom farming and mushroom gathering are large industries in many<br />
countries. Many fungi are producers of antibiotics, including β-lactam antibiotics such<br />
as penicillin and cephalosporin. [10] Widespread use of these antibiotics for the<br />
treatment of bacterial diseases, such as tuberculosis, syphilis, leprosy, and many<br />
others began in the early 20th century and continues to play a major part in antibacterial<br />
chemotherapy. The study of the historical uses and sociological impact of<br />
fungi is known as ethnomycology.<br />
Cultured foods<br />
Baker's yeast or Saccharomyces cerevisiae, a single-cell fungus, is used in the baking<br />
of bread and other wheat-based products, such as pizza and dumplings. [11] Several<br />
yeast species of the genus Saccharomyces are also used in the production of alcoholic<br />
beverages through fermentation. [12] Mycelial fungi, such as the shoyu koji mold<br />
(Aspergillus oryzae), are used in the brewing of Shoyu (soy sauce) and preparation of<br />
tempeh. [13] Quorn is a high-protein product made from the mold, Fusarium<br />
venenatum, and is used in vegetarian cooking.<br />
Other human uses<br />
Fungi are also used extensively to produce industrial chemicals like lactic acid,<br />
antibiotics and even to make stonewashed jeans. [14] Several fungal species are<br />
ingested for their psychedelic properties, both recreationally and religiously (see main<br />
article, Psilocybin mushrooms).<br />
Mycotoxins<br />
Main article: Mycotoxins<br />
Many fungi produce compounds with biological activity. Several of these compounds<br />
are toxic and are therefore called mycotoxins, referring to their fungal origin and toxic<br />
activity. Of particular relevance to humans are those mycotoxins that are produced by<br />
moulds causing food spoilage and poisonous mushrooms (see below). Particularly<br />
infamous are the aflatoxins, which are insidious liver toxins and highly carcinogenic<br />
metabolites produced by Aspergillus species often growing in or on grains and nuts<br />
consumed by humans, and the lethal amatoxins produced by mushrooms of the genus<br />
Amanita. Other notable mycotoxins include ochratoxins, patulin, ergot alkaloids, and<br />
trichothecenes and fumonisins, all of which have significant impact on human food<br />
supplies or animal livestock. [15]<br />
Mycotoxins belong to the group of secondary metabolites (or natural products).<br />
Originally, this group of compounds had been thought to be mere byproducts of<br />
primary metabolism, hence the name "secondary" metabolites. However, recent<br />
research has shown the existence of biochemical pathways solely for the purpose of<br />
producing mycotoxins and other natural products in fungi. [16] Mycotoxins provide a<br />
number of fitness benefits to the fungi that produce them in terms of physiological<br />
adaptation, competition with other microbes and fungi, and protection from<br />
fungivory. [17][18] These fitness benefits and the existence of dedicated biosynthetic
pathways for mycotoxin production suggest that the mycotoxins are important for<br />
fungal persistence and survival.<br />
Edible and poisonous fungi<br />
Asian mushrooms, clockwise from left, enokitake, buna-shimeji, bunapi-shimeji, king<br />
oyster mushroom and shiitake.<br />
Black Périgord Truffle (Tuber melanosporum), cut in half.<br />
Stilton cheese veined with Penicillium roqueforti.<br />
Some of the best known types of fungi are the edible and the poisonous mushrooms.<br />
Many species are commercially raised, but others must be harvested from the wild.<br />
Agaricus bisporus, sold as button mushrooms when small or Portobello mushrooms<br />
when larger, are the most commonly eaten species, used in salads, soups, and many<br />
other dishes. Many Asian fungi are commercially grown and have gained in<br />
popularity in the West. They are often available fresh in grocery stores and markets,<br />
including straw mushrooms (Volvariella volvacea), oyster mushrooms (Pleurotus<br />
ostreatus), shiitakes (Lentinula edodes), and enokitake (Flammulina spp.).<br />
There are many more mushroom species that are harvested from the wild for personal<br />
consumption or commercial sale. Milk mushrooms, morels, chanterelles, truffles,<br />
black trumpets, and porcini mushrooms (Boletus edulis) (also known as king boletes)<br />
all demand a high price on the market. They are often used in gourmet dishes.
For certain types of cheeses, it is also a common practice to inoculate milk curds with<br />
fungal spores to foment the growth of specific species of mold that impart a unique<br />
flavor and texture to the cheese. This accounts for the blue colour in cheeses such as<br />
Stilton or Roquefort which is created using Penicillium roqueforti spores. [19] Molds<br />
used in cheese production are usually non-toxic and are thus safe for human<br />
consumption; however, mycotoxins (e.g., aflatoxins, roquefortine C, patulin, or<br />
others) may accumulate due to fungal spoilage during cheese ripening or storage. [20]<br />
Many mushroom species are toxic to humans, with toxicities ranging from slight<br />
digestive problems or allergic reactions as well as hallucinations to severe organ<br />
failures and death. Some of the most deadly mushrooms belong to the genera Inocybe,<br />
Cortinarius, and most infamously, Amanita, which includes the destroying angel (A.<br />
virosa) and the death cap (A. phalloides), the most common cause of deadly<br />
mushroom poisoning. [21] The false morel (Gyromitra esculenta) is considered a<br />
delicacy by some when cooked yet can be deadly when raw. Tricholoma equestre is<br />
one which was considered edible for centuries yet recently responsible for a series of<br />
serious poisonings in France.<br />
Fly agaric mushrooms (A. muscaria) also cause occasional poisonings, mostly as a<br />
result of ingestion for use as a recreational drug for its hallucinogenic properties.<br />
Historically Fly agaric was used by Celtic Druids in Northern Europe and the Koryak<br />
people of north-eastern Siberia for religious or shamanic purposes. [22] It is difficult to<br />
identify a safe mushroom without proper training and knowledge, thus it is often<br />
advised to assume that a mushroom in the wild is poisonous and not to consume it.<br />
Fungi in the biological control of pests<br />
In agricultural settings, fungi that actively compete for nutrients and space with, and<br />
eventually prevail over, pathogenic microorganisms, such as bacteria or other fungi,<br />
via the competitive exclusion principle, [23] or are parasites of these pathogens, may be<br />
beneficial agents for human use. For example, some fungi may be used to suppress<br />
growth or eliminate harmful plant pathogens, such as insects, mites, weeds,<br />
nematodes and other fungi that cause diseases of important crop plants. [24] This has<br />
generated strong interest in the use and practical application of these fungi for the<br />
biological control of these agricultural pests. Entomopathogenic fungi can be used as<br />
biopesticides, as they actively kill insects. [25] Examples of fungi that have been used<br />
as bioinsecticides are Beauveria bassiana, Metarhizium anisopliae, Hirsutella spp,<br />
Paecilomyces fumosoroseus, and Verticillium lecanii. [26] [27] Endophytic fungi of<br />
grasses of the genus Neotyphodium, such as N. coenophialum produce alkaloids that<br />
are toxic to a range of invertebrate and vertebrate herbivores. These alkaloids protect<br />
the infected grass plants from herbivory, but some endophyte alkaloids can cause<br />
poisoning of grazing animals, such as cattle and sheep. [28] Infection of grass cultivars<br />
of turf or forage grasses with isolates of the grass endophytes that produce only<br />
specific alkaloids to improve grass hardiness and resistance to herbivores such as<br />
insects, while being non-toxic to livestock, is being used in grass breeding<br />
programs. [29]<br />
Ecology
Polypores growing on a tree in Borneo<br />
Although often inconspicuous, fungi occur in every environment on Earth and play<br />
very important roles in most ecosystems. Along with bacteria, fungi are the major<br />
decomposers in most terrestrial (and some aquatic) ecosystems, and therefore play a<br />
critical role in biogeochemical cycles and in many food webs. As decomposers, they<br />
play an indispensable role in nutrient cycling, especially as saprotrophs and<br />
symbionts, degrading organic matter to inorganic molecules, which can then re-enter<br />
anabolic metabolic pathways in plants or other organisms. [30][31]<br />
Symbiosis<br />
Many fungi have important symbiotic relationships with organisms from most if not<br />
all Kingdoms. [32][33][34] These interactions can be mutualistic or antagonistic in nature,<br />
or in case of commensal fungi are of no apparent benefit or detriment to the host.<br />
[35][36][37]<br />
With plants<br />
Mycorrhizal symbiosis between plants and fungi is one of the most well-known plantfungus<br />
associations and is of significant importance for plant growth and persistence<br />
in many ecosystems; over 90% of all plant species engage in some kind of<br />
mycorrhizal relationship with fungi and are dependent upon this relationship for<br />
survival. [38][39][40] The mycorrhizal symbiosis is ancient, dating to at least 400 million<br />
years ago. [41] It often increases the plant's uptake of inorganic compounds, such as<br />
nitrate and phosphate from soils having low concentrations of these key plant<br />
nutrients. [30] In some mycorrhizal associations, the fungal partners may mediate plantto-plant<br />
transfer of carbohydrates and other nutrients. Such mycorrhizal communities<br />
are called "common mycorrhizal networks". [42]<br />
Lichens are formed by a symbiotic relationship between algae or cyanobacteria<br />
(referred to in lichens as "photobionts") and fungi (mostly various species of<br />
ascomycetes and a few basidiomycetes), in which individual photobiont cells are<br />
embedded in a tissue formed by the fungus. [43] As in mycorrhizas, the photobiont<br />
provides sugars and other carbohydrates, while the fungus provides minerals and<br />
water. The functions of both symbiotic organisms are so closely intertwined that they<br />
function almost as a single organism.
With insects<br />
Many insects also engage in mutualistic relationships with various types of fungi.<br />
Several groups of ants cultivate fungi in the order Agaricales as their primary food<br />
source, while ambrosia beetles cultivate various species of fungi in the bark of trees<br />
that they infest. [44] Termites on the African Savannah are also known to cultivate<br />
fungi. [45]<br />
As pathogens and parasites<br />
However, many fungi are parasites on plants, animals (including humans), and other<br />
fungi. Serious fungal pathogens of many cultivated plants causing extensive damage<br />
and losses to agriculture and forestry include the rice blast fungus Magnaporthe<br />
oryzae, [46] tree pathogens such as Ophiostoma ulmi and Ophiostoma novo-ulmi<br />
causing Dutch elm disease, [47] and Cryphonectria parasitica responsible for chestnut<br />
blight, [48] and plant-pathogenic fungi in the genera Fusarium, Ustilago, Alternaria,<br />
and Cochliobolus; [36] fungi with the potential to cause serious human diseases,<br />
especially in persons with immuno-deficiencies, are in the genera Aspergillus,<br />
Candida, Cryptoccocus, [49][37][50] Histoplasma, [51] and Pneumocystis. [52] Several<br />
pathogenic fungi are also responsible for relatively minor human diseases, such as<br />
athlete’s foot and ringworm. Some fungi are predators of nematodes, which they<br />
capture using an array of specialized structures, such as constricting rings or adhesive<br />
nets. [53]<br />
Nutrition and possible autotrophy<br />
Growth of fungi as hyphae on or in solid substrates or single cells in aquatic<br />
environments is adapted to efficient extraction of nutrients from these environments,<br />
because these growth forms have high surface area to volume ratios. These<br />
adaptations in morphology are complemented by hydrolytic enzymes secreted into the<br />
environment for digestion of large organic molecules, such as polysaccharides,<br />
proteins, lipids, and other organic substrates into smaller molecules. [54][55][56] These<br />
molecules are then absorbed as nutrients into the fungal cells.<br />
Traditionally, the fungi are considered heterotrophs, organisms that rely solely on<br />
carbon fixed by other organisms for metabolism. Fungi have evolved a remarkable<br />
metabolic versatility that allows many of them to use a large variety of organic<br />
substrates for growth, including simple compounds as nitrate, ammonia, acetate, or<br />
ethanol. [57] [58] Recent research raises the possibility that some fungi utilize the<br />
pigment melanin to extract energy from ionizing radiation, such as gamma radiation<br />
for "radiotrophic" growth. [59] It has been proposed that this process might bear some<br />
similarity to photosynthesis in plants, [59] but detailed biochemical data supporting the<br />
existence of this hypothetical pathway are presently lacking.<br />
Morphology<br />
Microscopic structures
Mold covering a decaying peach over a period of six days. The frames were taken<br />
approximately 12 hours apart.<br />
Though fungi are part of the opisthokont clade, all phyla except for the chytrids have<br />
lost their posterior flagella. [60] Fungi are unusual among the eukaryotes in having a<br />
cell wall that, besides glucans (e.g., β-1,3-glucan) and other typical components,<br />
contains the biopolymer chitin. [61]<br />
Many fungi grow as thread-like filamentous microscopic structures called hyphae,<br />
and an assemblage of intertwined and interconnected hyphae is called a mycelium. [6]<br />
Hyphae can be septate, i.e., divided into hyphal compartments separated by a septum,<br />
each compartment containing one or more nuclei or can be coenocytic, i.e., lacking<br />
hyphal compartmentalization. However, septa have pores, such as the doliporus in the<br />
basidiomycetes that allow cytoplasm, organelles, and sometimes nuclei to pass<br />
through. [6] Coenocytic hyphae are essentially multinucleate supercells. [62] In some<br />
cases, fungi have developed specialized structures for nutrient uptake from living<br />
hosts; examples include haustoria in plant-parasitic fungi of nearly all divisions, and<br />
arbuscules of several mycorrhizal fungi, [63] which penetrate into the host cells for<br />
nutrient uptake by the fungus.<br />
Macroscopic structures<br />
Fungal mycelia can become visible macroscopically, for example, as concentric rings<br />
on various surfaces, such as damp walls, and on other substrates, such as spoilt food<br />
(see figure), and are commonly and generically called mould (American spelling,<br />
mold); fungal mycelia grown on solid agar media in laboratory petri dishes are usually<br />
referred to as colonies, with many species exhibiting characteristic macroscopic<br />
growth morphologies and colours, due to spores or pigmentation.<br />
Specialized fungal structures important in sexual reproduction are the apothecia,<br />
perithecia, and cleistothecia in the ascomycetes, and the fruiting bodies of the<br />
basidiomycetes, and a few ascomycetes. These reproductive structures can sometimes<br />
grow very large, and are well known as mushrooms.<br />
Morphological and physiological features for substrate penetration<br />
Fungal hyphae are specifically adapted to growth on solid surfaces and within<br />
substrates, and can exert astoundingly large penetrative mechanical forces. The plant<br />
pathogen, Magnaporthe grisea, forms a structure called an appressorium specifically
designed for penetration of plant tissues, and the pressure generated by the<br />
appressorium, which is directed against the plant epidermis can exceed 8 MPa (80<br />
bars). [64] The generation of these mechanical pressures is the result of an interplay<br />
between physiological processes to increase intracellular turgor by production of<br />
osmolytes such as glycerol, and the morphology of the appressorium. [65]<br />
Reproduction<br />
Fungi on a fence post near Orosí, Costa Rica.<br />
Reproduction of fungi is complex, reflecting the heterogeneity in lifestyles and<br />
genetic make up within this group of organisms. [6] Many fungi reproduce both<br />
sexually or asexually, depending on conditions in the environment. These conditions<br />
trigger genetically determined developmental programs leading to the expression of<br />
specialized structures for sexual or asexual reproduction. These structures aid both<br />
reproduction and efficient dissemination of spores or spore-containing propagules.<br />
Asexual reproduction<br />
Asexual reproduction via vegetative spores or through mycelial fragmentation is<br />
common in many fungal species and allows more rapid dispersal than sexual<br />
reproduction. In the case of the "Fungi imperfecti" or Deuteromycota, which lack a<br />
sexual cycle, it is the only means of propagation. Asexual spores, upon germination,<br />
may found a population that is clonal to the population from which the spore<br />
originated, and thus colonize new environments.<br />
Sexual reproduction<br />
Sexual reproduction with meiosis exists in all fungal phyla, except the<br />
Deuteromycota. It differs in many aspects from sexual reproduction in animals or<br />
plants. Many differences also exist between fungal groups and have been used to<br />
discriminate fungal clades and species based on morphological differences in sexual<br />
structures and reproductive strategies. Experimental crosses between fungal isolates<br />
can also be used to identify species based on biological species concepts. The major<br />
fungal clades have initially been delineated based on the morphology of their sexual<br />
structures and spores; for example, the spore-containing structures, asci and basidia,<br />
can be used in the identification of ascomycetes and basidiomycetes, respectively.<br />
Many fungal species have elaborate vegetative incompatibility systems that allow<br />
mating only between individuals of opposite mating type, while others can mate and
sexually reproduce with any other individual or itself. Species of the former mating<br />
system are called heterothallic, and of the latter homothallic. [66]<br />
Most fungi have both a haploid and diploid stage in their life cycles. In all sexually<br />
reproducing fungi, compatible individuals combine by cell fusion of vegetative<br />
hyphae by anastomosis, required for the initiation of the sexual cycle. Ascomycetes<br />
and basidiomycetes go through a dikaryotic stage, in which the nuclei inherited from<br />
the two parents do not fuse immediately after cell fusion, but remain separate in the<br />
hyphal cells (see heterokaryosis).<br />
In ascomycetes, dikaryotic hyphae of the hymenium form a characteristic hook at the<br />
hyphal septum. During cell division formation of the hook ensures proper distribution<br />
of the newly divided nuclei into the apical and basal hyphal compartments. An ascus<br />
(plural asci) is then formed, in which karyogamy (nuclear fusion) occurs. These asci<br />
are embedded in an ascocarp, or fruiting body, of the fungus. Karyogamy in the asci is<br />
followed immediately by meiosis and the production of ascospores. The ascospores<br />
are disseminated and germinate and may form a new haploid mycelium. [67]<br />
Sexual reproduction in basidiomycetes is similar to that of the ascomycetes.<br />
Compatible haploid hyphae fuse to produce a dikaryotic mycelium. However, the<br />
dikaryotic phase is more extensive in the basidiomycetes, in many cases also present<br />
in the vegetatively growing mycelium. A specialized anatomical structure, called a<br />
clamp connection, is formed at each hyphal septum. As with the structurally similar<br />
hook in the ascomycetes, formation of the clamp connection in the basidiomycetes is<br />
required for controlled transfer of nuclei during cell division, to maintain the<br />
dikaryotic stage with two genetically different nuclei in each hyphal compartment. [67]<br />
A basidiocarp is formed in which club-like structures known as basidia generate<br />
haploid basidiospores after karyogamy and meiosis. [68] The most commonly known<br />
basidiocarps are mushrooms, but they may also take many other forms (see<br />
Morphology section).<br />
In zygomycetes, haploid hyphae of two individuals fuse, forming a zygote, which<br />
develops into a zygospore. When the zygospore germinates, it quickly undergoes<br />
meiosis, generating new haploid hyphae, which in turn may form asexual<br />
sporangiospores. These sporangiospores are means of rapid dispersal of the fungus<br />
and germinate into new genetically identical haploid fungal colonies, able to mate and<br />
undergo another sexual cycle followed by the generation of new zygospores, thus<br />
completing the lifecycle.<br />
Spore dispersal<br />
Both asexual and sexual spores or sporangiospores of many fungal species are<br />
actively dispersed by forcible ejection from their reproductive structures. This<br />
ejection ensures exit of the spores from the reproductive structures as well as<br />
travelling through the air over long distances. Many fungi thereby possess specialized<br />
mechanical and physiological mechanisms as well as spore-surface structures, such as<br />
hydrophobins, for spore ejection. These mechanisms include, for example, forcible<br />
discharge of ascospores enabled by the structure of the ascus and accumulation of<br />
osmolytes in the fluids of the ascus that lead to explosive discharge of the ascospores<br />
into the air. [69] The forcible discharge of single spores termed ballistospores involves
formation of a small drop of water (Buller's drop), which upon contact with the spore<br />
leads to its projectile release with an initial acceleration of more than 10,000 g. [70]<br />
Other fungi rely on alternative mechanisms for spore release, such as external<br />
mechanical forces, exemplified by puffballs. Attracting insects, such as flies, to<br />
fruiting structures, by virtue of their having lively colours and a putrid odour, for<br />
dispersal of fungal spores is yet another strategy, most prominently used by the<br />
stinkhorns.<br />
Other sexual processes<br />
Besides regular sexual reproduction with meiosis, some fungal species may exchange<br />
genetic material via parasexual processes, initiated by anastomosis between hyphae<br />
and plasmogamy of fungal cells. The frequency and relative importance of parasexual<br />
events is unclear and may be lower than other sexual processes. However, it is known<br />
to play a role in intraspecific hybridization [71] and is also likely required for<br />
hybridization between fungal species, which has been associated with major events in<br />
fungal evolution. [72]<br />
Phylogeny and classification<br />
The mushroom Oudemansiella nocturnum eats wood<br />
For a long time taxonomists considered fungi to be members of the Plant Kingdom.<br />
This early classification was based mainly on similarities in lifestyle: both fungi and<br />
plant are mainly sessile, have similarities in general morphology and growth habitat<br />
(like plants, fungi often grow in soil, in the case of mushrooms forming conspicuous<br />
fruiting bodies, which sometimes bear resemblance to plants such as mosses).<br />
Moreover, both groups possess a cell wall, which is absent in the Animal Kingdom.<br />
However, the fungi are now considered a separate kingdom, distinct from both plants<br />
and animals, from which they appear to have diverged approximately one billion<br />
years ago. [73] Many studies have identified several distinct morphological,<br />
biochemical, and genetic features in the Fungi, clearly delineating this group from the<br />
other kingdoms. For these reasons, the fungi are placed in their own kingdom.<br />
Physiological and morphological traits<br />
Similar to animals and unlike most plants, fungi lack the capacity to synthesize<br />
organic carbon by chlorophyll-based photosynthesis; whereas plants store the reduced<br />
carbon as starch, fungi, like animals and some bacteria, use glycogen [74] for storage<br />
of carbohydrates. A major component of the cell wall in many fungal species is the<br />
nitrogen-containing carbohydrate, chitin, [75] also present in some animals, such as the<br />
insects and crustaceans, while the plant cell wall consists chiefly of the carbohydrate<br />
cellulose. The defining and unique characteristics of fungal cells include growth as
hyphae, which are microscopic filaments of between 2-10 microns in diameter and up<br />
to several centimetres in length, and which combined form the fungal mycelium.<br />
Some fungi, such as yeasts, grow as single ovoid cells, similar to unicellular algae and<br />
the protists.<br />
Unlike many plants, most fungi lack an efficient vascular system, such as xylem or<br />
phloem for long-distance transport of water and nutrients; as an example for<br />
convergent evolution, some fungi, such as Armillaria, form rhizomorphs or mycelial<br />
cords, [76] resembling and functionally related to, but morphologically distinct from,<br />
plant roots.<br />
Some characteristics shared between plants and fungi include the presence of<br />
vacuoles in the cell, [77] and a similar pathway in the biosynthesis of terpenes using<br />
mevalonic acid and pyrophosphate as biochemical precursors; plants however use an<br />
additional terpene biosynthesis pathway in the chloroplasts that is apparently absent in<br />
fungi. [78] Ancestral traits shared among members of the fungi include chitinous cell<br />
walls and heterotrophy by absorption. [67] A further characteristic of the fungi that is<br />
absent from other eukaryotes, and shared only with some bacteria, is the biosynthesis<br />
of the amino acid, L-lysine, via the α-aminoadipate pathway. [79]<br />
Similar to plants, fungi produce a plethora of secondary metabolites functioning as<br />
defensive compounds or for niche adaptation; however, biochemical pathways for the<br />
synthesis of similar or even identical compounds often differ markedly between fungi<br />
and plants. [80][81]<br />
Evolutionary history<br />
Even though traditionally included in many botany curricula and textbooks, fungi are<br />
now thought to be more closely related to animals than to plants, and are placed with<br />
the animals in the monophyletic group of opisthokonts. [67] For much of the Paleozoic<br />
Era, the fungi appear to have been aquatic, and consisted of organisms similar to the<br />
extant Chytrids in having flagellum-bearing spores. [82] The first land fungi probably<br />
appeared in the Silurian, right after the first land plants appeared, even though their<br />
fossils are fragmentary. For some time after the Permian-Triassic extinction event, a<br />
fungal spike, detected as an extraordinary abundance of fungal spores in sediments<br />
formed shortly after this event, indicates that they were the dominant life form during<br />
this period—nearly 100% of the fossil record available from this period. [83]<br />
Analyses using molecular phylogenetics support a monophyletic origin of the<br />
Fungi. [8] The taxonomy of the Fungi is in a state of constant flux, especially due to<br />
recent research based on DNA comparisons. These current phylogenetic analyses<br />
often overturn classifications based on older and sometimes less discriminative<br />
methods based on morphological features and biological species concepts obtained<br />
from experimental matings. [84][85]<br />
There is no unique generally accepted system at the higher taxonomic levels and there<br />
are constant name changes at every level, from species upwards. However, efforts<br />
among fungal researchers are now underway to establish and encourage usage of a<br />
unified and more consistent nomenclature. [8] Fungal species can also have multiple<br />
scientific names depending on its life cycle and mode (sexual or asexual) of
eproduction. Web sites such as Index Fungorum and ITIS define preferred up-to-date<br />
names (with cross-references to older synonyms), but do not always agree with each<br />
other.<br />
Cladogram<br />
Unikonta<br />
Opisthokonta<br />
The taxonomic groups of fungi<br />
Amoebozoa<br />
Fungi<br />
Animalia<br />
Choanozoa<br />
Dikarya<br />
Chytridiomycota<br />
Blastocladiomycota<br />
Neocallimastigomycota<br />
Zygomycota<br />
Glomeromycota<br />
Ascomycota<br />
Basidiomycota<br />
The major divisions (phyla) of fungi have been classified based mainly on their sexual<br />
reproductive structures. Currently, seven fungal divisions are proposed: [8]<br />
Arbuscular mycorrhiza seen under microscope. Flax root cortical cells containing<br />
paired arbuscules.
Conidiophores of molds of the genus Aspergillus, an ascomycete, seen under<br />
microscope.<br />
• The Chytridiomycota are commonly known as chytrids. These fungi are<br />
ubiquitous with a worldwide distribution; chytrids produce zoospores that are<br />
capable of active movement through aqueous phases with a single flagellum.<br />
Consequently, some taxonomists had earlier classified them as protists on the<br />
basis of the flagellum. Molecular phylogenies, inferred from the rRNA-operon<br />
sequences representing the 18S, 28S, and 5.8S ribosomal subunits, suggest<br />
that the Chytrids are a basal fungal group divergent from the other fungal<br />
divisions, consisting of four major clades with some evidence for paraphyly or<br />
possibly polyphyly. [82]<br />
• The Blastocladiomycota were previously considered a taxonomic clade within<br />
the Chytridiomycota. Recent molecular data and ultrastructural characteristics,<br />
however, place the Blastocladiomycota as a sister clade to the Zygomycota,<br />
Glomeromycota, and Dikarya (Ascomycota and Basiomycota). The<br />
blastocladiomycetes are fungi that are saprotrophs and parasites of all<br />
eukaryotic groups and undergo sporic meiosis unlike their close relatives, the<br />
chytrids, which mostly exhibit zygotic meiosis. [82]<br />
• The Neocallimastigomycota were earlier placed in the phylum<br />
Chytridomycota. Members of this small phylum are anaerobic organisms,<br />
living in the digestive system of larger herbivorous mammals and possibly in<br />
other terrestrial and aquatic environments. They lack mitochondria but contain<br />
hydrogenosomes of mitochondrial origin. As the related chrytrids,<br />
neocallimastigomycetes form zoospores that are posteriorly uniflagellate or<br />
polyflagellate. [8]<br />
• The Zygomycota contain the taxa, Zygomycetes and Trichomycetes, and<br />
reproduce sexually with meiospores called zygospores and asexually with<br />
sporangiospores. Black bread mold (Rhizopus stolonifer) is a common species<br />
that belongs to this group; another is Pilobolus, which is capable of ejecting<br />
spores several meters through the air. Medically relevant genera include<br />
Mucor, Rhizomucor, and Rhizopus. Molecular phylogenetic investigation has<br />
shown the Zygomycota to be a polyphyletic phylum with evidence of<br />
paraphyly within this taxonomic group. [86]<br />
• Members of the Glomeromycota are fungi forming arbuscular mycorrhizae<br />
with higher plants. Only one species has been observed forming zygospores;<br />
all other species solely reproduce asexually. The symbiotic association<br />
between the Glomeromycota and plants is ancient, with evidence dating to 400<br />
million years ago. [41]
Diagram of an apothecium (the typical cup-like reproductive structure of<br />
Ascomycetes) showing sterile tissues as well as developing and mature asci.<br />
• The Ascomycota, commonly known as sac fungi or ascomycetes, constitute<br />
the largest taxonomic group within the Eumycota. These fungi form meiotic<br />
spores called ascospores, which are enclosed in a special sac-like structure<br />
called an ascus. This division includes morels, a few mushrooms and truffles,<br />
single-celled yeasts (e.g., of the genera Saccharomyces, Kluyveromyces,<br />
Pichia, and Candida), and many filamentous fungi living as saprotrophs,<br />
parasites, and mutualistic symbionts. Prominent and important genera of<br />
filamentous ascomycetes include Aspergillus, Penicillium, Fusarium, and<br />
Claviceps. Many ascomycetes species have only been observed undergoing<br />
asexual reproduction (called anamorphic species), but molecular data has often<br />
been able to identify their closest teleomorphs in the Ascomycota. Because the<br />
products of meiosis are retained within the sac-like ascus, several ascomyctes<br />
have been used for elucidating principles of genetics and heredity (e.g.<br />
Neurospora crassa).<br />
• Members of the Basidiomycota, commonly known as the club fungi or<br />
basidiomycetes, produce meiospores called basidiospores on club-like stalks<br />
called basidia. Most common mushrooms belong to this group, as well as rust<br />
(fungus) and smut fungi, which are major pathogens of grains. Other<br />
important Basidiomyces include the maize pathogen,Ustilago maydis, human<br />
commensal species of the genus Malassezia, and the opportunistic human<br />
pathogen, Cryptococcus neoformans.<br />
Phylogenetic relationships with other fungus-like organisms<br />
Because of some similarities in morphology and lifestyle, the slime molds<br />
(myxomycetes) and water molds (oomycetes) were formerly classified in the kingdom<br />
Fungi. Unlike true fungi, however, the cell walls of these organisms contain cellulose<br />
and lack chitin. Slime molds are unikonts like fungi, but are grouped in the<br />
Amoebozoa. Water molds are diploid bikonts, grouped in the Chromalveolate<br />
kingdom. Neither water molds nor slime molds are closely related to the true fungi,<br />
and, therefore, taxonomists no longer group them in the kingdom Fungi. Nonetheless,<br />
studies of the oomycetes and myxomycetes are still often included in mycology<br />
textbooks and primary research literature.<br />
It has been suggested that the nucleariids, currently grouped in the Choanozoa, may<br />
be a sister group to the oomycete clade, and as such could be included in an expanded<br />
fungal kingdom. [87]
See also<br />
• Bioaerosol<br />
• Carnivorous fungus<br />
• Fusicoccin<br />
• List of fungal orders<br />
• MycoBank<br />
• Mycotoxin<br />
• Plant pathology<br />
• Wood-decay fungus<br />
• Quorn<br />
• Pathogenic fungi<br />
Notes and references<br />
1. ^ (1980) "Taxonomic proposals for the classification of marine yeasts and other<br />
yeast-like fungi including the smuts". Bot. Mar. 23: 371.<br />
2. ^ These are the pronunciations listed first in most dictionaries. See, for example, the<br />
Merriam-Webster Online entry Alternative pronunciations for fungi include<br />
/ f ŋga /, / f nd i/, and / f ŋgi/. Funguses (/ f ŋgәsәz/) is an alternative<br />
plural form.<br />
3. ^ Simpson, D.P. (1979). Cassell's Latin Dictionary, 5, London: Cassell Ltd., 883.<br />
ISBN 0-304-52257-0.<br />
4. ^ Hawksworth DL (2006). "The fungal dimension of biodiversity: magnitude,<br />
significance, and conservation". Mycol. Res. 95: 641–655.<br />
5. ^ Mueller GM, Schmit JP (2006). "Fungal biodiversity: what do we know? What can<br />
we predict?". Biodivers Conserv 16: 1–5.<br />
6. ^ a b c d Alexopoulos CJ, Mims CW, Blackwell M (1996). Introductory Mycology.<br />
John Wiley and Sons. ISBN 0471522295.<br />
7. ^ Meredith Blackwell; Rytas Vilgalys, and John W. Taylor (2005-02-14). Eumycota:<br />
mushrooms, sac fungi, yeast, molds, rusts, smuts, etc. (English). Retrieved on 2007-<br />
04-06.<br />
8. ^ a b c d e Hibbett, D.S., et al. (2007). "A higher level phylogenetic classification of the<br />
Fungi". Mycol. Res. 111 (5): 509-547. doi:doi:10.1016/j.mycres.2007.03.004.<br />
9. ^ Strains of wine yeast<br />
10. ^ Demain AL. (1991). "Production of beta-lactam antibiotics and its regulation.".<br />
Proc Natl Sci Counc Repub China B. 15: 251-265. PMID 1815263.<br />
11. ^ Kulp, Karel (2000). Handbook of Cereal Science and Technology. CRC Press.<br />
ISBN 0824782941.<br />
12. ^ Piskur J, Rozpedowska E, Polakova S, Merico A, Compagno C. (2006). "How did<br />
Saccharomyces evolve to become a good brewer?". Trends Genet. 22: 183-186.<br />
PMID 16499989.<br />
13. ^ Kitamoto N, Yoshino S, Ohmiya K, Tsukagoshi N. (1999). "Sequence analysis,<br />
overexpression, and antisense inhibition of a beta-xylosidase gene, xylA, from<br />
Aspergillus oryzae KBN616.". Appl. Env. Microbiol. 65: 20-24. PMID 9872754.<br />
14. ^ Trichoderma spp., including T. harzianum, T. viride, T. koningii, T. hamatum and<br />
other spp. Deuteromycetes, Moniliales (asexual classification system). Biological<br />
Control: A Guide to Natural Enemies in North America. Retrieved on 2007-07-10.<br />
15. ^ van Egmond HP, Schothorst RC, Jonker MA (2007). "Regulations relating to<br />
mycotoxins in food: perspectives in a global and European context". Anal Bioanal<br />
Chem. 389: 147-157. PMID 17508207.
16. ^ Keller NP, Turner G, Bennett JW (2005). "Fungal secondary metabolism - from<br />
biochemistry to genomics". Nat Rev Microbiol. 3: 937-497. PMID 16322742.<br />
17. ^ Demain AL, Fang A (2000). "The natural functions of secondary metabolites". Adv<br />
Biochem Eng Biotechnol. 69: 1-39. PMID 11036689.<br />
18. ^ Rohlfs M, Albert M, Keller NP, Kempken F (2007). "Secondary chemicals protect<br />
mould from fungivory". Biol Lett. 3: 523-525. PMID 17686752.<br />
19. ^ Questions & Answers - Mold on Cheese whatscookingamerica.net. Retrieved 2007-<br />
04-06.<br />
20. ^ Erdogan A, Gurses M, Sert S. (2004). "Isolation of moulds capable of producing<br />
mycotoxins from blue mouldy Tulum cheeses produced in Turkey.". Int J Food<br />
Microbiol. 85: 83-85. PMID 12810273.<br />
21. ^ On the Trail of the Death Cap Mushroom Richard Harris, www.npr.org, 2007-02-<br />
08. Retrieved 2007-04-06.<br />
22. ^ Mythology and Folklore of Fly Agaric Paul Kendall, Trees for Life. Retrieved<br />
2007-04-06.<br />
23. ^ López-Gómez J, Molina-Meyer M (2006). "The competitive exclusion principle<br />
versus biodiversity through competitive segregation and further adaptation to spatial<br />
heterogeneities". Theor Popul Biol. 69: 94-109. PMID 16223517.<br />
24. ^ Setting the Stage To Screen Biocontrol Fungi Hank Becker, July 1998. Retrieved<br />
2007-04-06.<br />
25. ^ WHEY-BASED FUNGAL MICROFACTORY TECHNOLOGY FOR<br />
ENHANCED BIOLOGICAL PEST MANAGEMENT USING FUNGI Todd. S.<br />
Keiller, Technology Transfer, University of Vermont. Retrieved 2007-04-06.<br />
26. ^ Deshpande MV. (1999). "Mycopesticide production by fermentation: potential and<br />
challenges.". Crit Rev Microbiol. 25: 229-243. PMID 10524330.<br />
27. ^ Thomas MB, Read AF. (2007). "Can fungal biopesticides control malaria?". Nat<br />
Rev Microbiol. 5: 377-383. PMID 17426726.<br />
28. ^ Bush LP, Wilkinson HH, Schardl CL. (1997). "Bioprotective Alkaloids of Grass-<br />
Fungal Endophyte Symbioses". Plant Physiol. 114: 1-7. PMID 12223685.<br />
29. ^ Bouton JH, Latch GCM, Hill NS, Hoveland CS, McCannc MA, Watson RH, Parish<br />
JA, Hawkins LL, Thompson FN (2002). "Use of nonergot alkaloid-producing<br />
endophytes for alleviating tall fescue toxicosis in sheep.". Agron. J. 94: 567-574.<br />
http://agron.scijournals.org/cgi/content/full/94/3/567.<br />
30. ^ a b Lindahl BD, Ihrmark K, Boberg J, Trumbore SE, Högberg P, Stenlid J, Finlay<br />
RD (2007). "Spatial separation of litter decomposition and mycorrhizal nitrogen<br />
uptake in a boreal forest". New Phytol. 173: 611-620. PMID 17244056.<br />
31. ^ Barea JM, Pozo MJ, Azcón R, Azcón-Aguilar C (2005). "Microbial co-operation in<br />
the rhizosphere". J. Exp. Bot. 56: 1761-1778. PMID 15911555.<br />
32. ^ Aanen DK. (2006). "As you reap, so shall you sow: coupling of harvesting and<br />
inoculating stabilizes the mutualism between termites and fungi.". Biol Lett. 2: 209-<br />
212. PMID 17148364.<br />
33. ^ Nikoh N, Fukatsu T. (2000). "Interkingdom host jumping underground:<br />
phylogenetic analysis of entomoparasitic fungi of the genus Cordyceps.". Mol Biol<br />
Evol. 17: 2629-2638. PMID 10742053.<br />
34. ^ Perotto S, Bonfante P. (1997). "Bacterial associations with mycorrhizal fungi: close<br />
and distant friends in the rhizosphere.". Trends Microbiol. 5: 496-501. PMID<br />
9447662.<br />
35. ^ Arnold AE, Mejía LC, Kyllo D, Rojas EI, Maynard Z, Robbins N, Herre EA.<br />
(2003). "Fungal endophytes limit pathogen damage in a tropical tree.". Proc. Natl.<br />
Acad. Sci. USA 100: 15649-15654. PMID 14671327.<br />
36. ^ a b Paszkowski U. (2006). "Mutualism and parasitism: the yin and yang of plant<br />
symbioses.". Curr Opin Plant Biol. 9: 364-370. PMID 16713732.<br />
37. ^ a b Hube B. (2004). "From commensal to pathogen: stage- and tissue-specific gene<br />
expression of Candida albicans.". Curr Opin Microbiol. 7: 336-341. PMID<br />
15288621.
38. ^ Volk, Tom. Tom Volk's Fungi FAQ. Retrieved on 2006-09-21.<br />
39. ^ Wong, George. Symbiosis: Mycorrhizae and Lichens. Retrieved on 2006-09-21.<br />
40. ^ Knowledge of nitrogen transfer between plants and beneficial fungi expands<br />
southwestfarmpress.com. 2005-06-10 Retrieved 2007-04-06.<br />
41. ^ a b Remy W, Taylor TN, Hass H, Kerp H (1994). "4-hundred million year old<br />
vesicular-arbuscular mycorrhizae.". Proc. Natl. Acad. Sci 91: 11841-11843. PMID<br />
11607500.<br />
42. ^ Selosse MA, Richard F, He X, Simard SW (2006). "Mycorrhizal networks: des<br />
liaisons dangereuses?". Trends Ecol Evol. 21: 621-628. PMID 16843567.<br />
43. ^ Brodo, Irwin M.; Sylvia Duran Sharnoff (2001). Lichens of North America. Yale<br />
University Press. ISBN 0300082495.<br />
44. ^ Fungi and Insect Symbiosis www.botany.hawaii.edu. Retrieved 2007-04-06.<br />
45. ^ Pascal Jouquet, Virginie Tavernier, Luc Abbadie and Michel Lepage. Nests of<br />
subterranean fungus-growing termites (Isoptera, Macrotermitinae) as nutrient<br />
patches for grasses in savannah ecosystems. African Journal of Ecology. 2005. Vol<br />
43, 191–196<br />
46. ^ Talbot NJ (2003). "On the trail of a cereal killer: Exploring the biology of<br />
Magnaporthe grisea.". Annu Rev Microbiol. 57: 177-202. PMID 14527276.<br />
47. ^ Paoletti M, Buck KW, Brasier CM. (2006). "Selective acquisition of novel mating<br />
type and vegetative incompatibility genes via interspecies gene transfer in the<br />
globally invading eukaryote Ophiostoma novo-ulmi.". Mol Ecol. 15: 249-262. PMID<br />
16367844.<br />
48. ^ Gryzenhout M, Wingfield BD, Wingfield MJ. (2006). "New taxonomic concepts<br />
for the important forest pathogen Cryphonectria parasitica and related fungi.". FEMS<br />
Microbiol Lett. 258: 161-172. PMID 16640568.<br />
49. ^ Nielsen K, Heitman J. (2007). "Sex and virulence of human pathogenic fungi.". Adv<br />
Genet. 57: 143-173. PMID 17352904.<br />
50. ^ Brakhage AA (2005). "Systemic fungal infections caused by Aspergillus species:<br />
epidemiology, infection process and virulence determinants.". Curr. Drug Targets 6:<br />
875-886. PMID 16375671.<br />
51. ^ Kauffman CA. (2007). "Histoplasmosis: a clinical and laboratory update". Clin<br />
Microbiol Rev. 20: 115-132. PMID 17223625.<br />
52. ^ Cushion MT, Smulian AG, Slaven BE, Sesterhenn T, Arnold J, Staben C, Porollo<br />
A, Adamczak R, Meller J. (2007). "Transcriptome of Pneumocystis carinii during<br />
Fulminate Infection: Carbohydrate Metabolism and the Concept of a Compatible<br />
Parasite.". PLoS ONE 2: e423. PMID 17487271.<br />
53. ^ ILLUSTRATIONS for Predatory Fungi, wood Decay and the Carbon Cycle<br />
www.uoguelph.ca. Retrieved 2007-04-06.<br />
54. ^ Pereira JL, Noronha EF, Miller RN, Franco OL. (2007). "Novel insights in the use<br />
of hydrolytic enzymes secreted by fungi with biotechnological potential.". Lett Appl<br />
Microbiol. 44: 573-581. PMID 17576216.<br />
55. ^ Schaller M, Borelli C, Korting HC, Hube B. (2007). "Hydrolytic enzymes as<br />
virulence factors of Candida albicans.". Mycoses 48: 365-377. PMID 16262871.<br />
56. ^ Farrar JF (1985). "Carbohydrate metabolism in biotrophic plant pathogens.".<br />
Microbiol Sci. 2: 314-317. PMID 3939987.<br />
57. ^ Marzluf GA (1981). "Regulation of nitrogen metabolism and gene expression in<br />
fungi". Microbiol Rev. 45: 437-461. PMID 6117784.<br />
58. ^ Heynes MJ (1994). "Regulatory circuits of the amdS gene of Aspergillus nidulans".<br />
Antonie Van Leeuwenhoek. 65: 179-782. PMID 7847883.<br />
59. ^ a b Dadachova E, Bryan RA, Huang X, Moadel T, Schweitzer AD, Aisen P,<br />
Nosanchuk JD, Casadevall A. (2007). "Ionizing radiation changes the electronic<br />
properties of melanin and enhances the growth of melanized fungi". PLoS ONE 2:<br />
e457. PMID 17520016.
60. ^ The Protistan Origins of Animals and Fungi Emma T. Steenkamp, Jane Wright and<br />
Sandra L. Baldauf. Molecular Biology and Evolution 2006 23(1):93-106;<br />
doi:10.1093/molbev/msj011. Retrieved 2007-04-06.<br />
61. ^ Stevens DA, Ichinomiya M, Koshi Y, Horiuchi H. (2006). "Escape of Candida<br />
from caspofungin inhibition at concentrations above the MIC (paradoxical effect)<br />
accomplished by increased cell wall chitin; evidence for beta-1,6-glucan synthesis<br />
inhibition by caspofungin.". Antimicrob Agents Chemother. 50: 3160-3161.. PMID<br />
16940118.<br />
62. ^ Chang, Shu-ting; Philip G. Miles (2004). Mushrooms: Cultivation, Nutritional<br />
Value, Medicinal Effect and Environmental Impact. CRC Press. ISBN 0849310431.<br />
63. ^ “Fungal Biology” at The University of Sydney Retrieved on 26 June 2007<br />
64. ^ Howard RJ, Ferrari MA, Roach DH, Money NP (1991). "Penetration of hard<br />
substrates by a fungus employing enormous turgor pressures". Proc Natl Acad Sci U<br />
S A. 88: 11281-11284. PMID 1837147.<br />
65. ^ Wang ZY, Jenkinson JM, Holcombe LJ, Soanes DM, Veneault-Fourrey C,<br />
Bhambra GK, Talbot NJ (2005). "The molecular biology of appressorium turgor<br />
generation by the rice blast fungus Magnaporthe grisea". Biochem Soc Trans. 33:<br />
384-388. PMID 15787612.<br />
66. ^ Metzenberg RL, Glass NL. (1990). "Mating type and mating strategies in<br />
Neurospora.". Bioessays 12: 53-59. PMID 2140508.<br />
67. ^ a b c d P. Sitte, H. Ziegler, F. Ehrendorfer (1991). Strasburger Lehrbuch der Botanik<br />
(Textbook of Botany), 33 ed, Urban & Fischer. ISBN 3437204475.<br />
68. ^ Reproduction of fungi MicrobiologyBytes, 2007-01-18. Retrieved 2007-04-06.<br />
69. ^ Trail F. (2007). "Fungal cannons: explosive spore discharge in the Ascomycota".<br />
FEMS Microbiol Lett. 276: 12-18. PMID 17784861.<br />
70. ^ Pringle A, Patek SN, Fischer M, Stolze J, Money NP. (2005). "The captured launch<br />
of a ballistospore". Mycologia 97: 866-871. PMID 16457355.<br />
71. ^ Furlaneto MC, Pizzirani-Kleiner AA. (1992). "Intraspecific hybridisation of<br />
Trichoderma pseudokoningii by anastomosis and by protoplast fusion.". FEMS<br />
Microbiol Lett. 69: 191-195. PMID 1537549.<br />
72. ^ Schardl CL, Craven KD. (2003). "Interspecific hybridization in plant-associated<br />
fungi and oomycetes: a review.". Mol. Ecol. 12: 2861-2873. PMID 14629368.<br />
73. ^ Bruns T. (2006). "Evolutionary biology: a kingdom revised.". Nature 443: 758-761.<br />
PMID 17051197.<br />
74. ^ Lomako J, Lomako WM, Whelan WJ. (2004). "Glycogenin: the primer for<br />
mammalian and yeast glycogen synthesis". Biochim Biophys Acta. 1673: 45-55.<br />
PMID 15238248.<br />
75. ^ Bowman SM, Free SJ. (2006). "The structure and synthesis of the fungal cell wall".<br />
Bioessays 28: 799-808. PMID 16927300.<br />
76. ^ Mihail JD, Bruhn JN. (2005). "Foraging behaviour of Armillaria rhizomorph<br />
systems". Mycol. Res. 109: 1195-1207. PMID 16279413.<br />
77. ^ Shoji JY, Arioka M, Kitamoto K (2006). "Possible involvement of pleiomorphic<br />
vacuolar networks in nutrient recycling in filamentous fungi". Autophagy. 2: 226-227.<br />
PMID 16874107.<br />
78. ^ Wu S, Schalk M, Clark A, Miles RB, Coates R, Chappell J. (2007). "Redirection of<br />
cytosolic or plastidic isoprenoid precursors elevates terpene production in plants".<br />
Nat Biotechnol. 24: 1441-7. PMID 17057703.<br />
79. ^ Xu H, Andi B, Qian J, West AH, Cook PF (2006). "The alpha-aminoadipate<br />
pathway for lysine biosynthesis in fungi". Cell Biochem Biophys. 46: 43-64. PMID<br />
16943623.<br />
80. ^ Tudzynski B. (2005). "Gibberellin biosynthesis in fungi: genes, enzymes,<br />
evolution, and impact on biotechnology". Appl Microbiol Biotechnol. 66: 597-611.<br />
PMID 15578178.
81. ^ Siewers V, Smedsgaard J, Tudzynski P. (2004). "The P450 monooxygenase<br />
BcABA1 is essential for abscisic acid biosynthesis in Botrytis cinerea.". Appl<br />
Environ. Microbiol. 70: 3868-3876. PMID 15240257.<br />
82. ^ a b c James TY et al (2006). "Reconstructing the early evolution of Fungi using a<br />
six-gene phylogeny.". Nature 443: 818-822. PMID 17051209.<br />
83. ^ Eshet, Y. et al. (1995) Fungal event and palynological record of ecological crisis<br />
and recovery across the Permian-Triassic boundary. Geology, 23, 967-970.<br />
84. ^ See Palaeos: Fungi for an introduction to fungal taxonomy, including recent<br />
controversies.<br />
85. ^ “A Higher-Level Phylogenetic Classification of the Fungi” by David S. Hibbett,<br />
(.pdf file) Retrieved on 8 March 2007<br />
86. ^ White MM, James TY, O'Donnell K, Cafaro MJ, Tanabe Y, Sugiyama J. (2006).<br />
"Phylogeny of the Zygomycota based on nuclear ribosomal sequence data.".<br />
Mycologia 98: 872-884. PMID 17486964.<br />
87. ^ Esser, Karl; Paul A. Lemke (1994). The Mycota: A Comprehensive Treatise on<br />
Fungi as Experimental Systems for Basic and Applied Research. Springer. ISBN<br />
3540580085.<br />
Further reading<br />
• Alexopoulos, C.J., Charles W. Mims, M. Blackwell et al., Introductory<br />
Mycology, 4 th ed. (John Wiley and Sons, Hoboken NJ, 2004) ISBN 0-471-<br />
52229-5<br />
• Arora, David. (1986). "Mushrooms Demystified: A Comprehensive Guide to<br />
the Fleshy Fungi". 2nd ed. Ten Speed Press. ISBN 0898151694<br />
• Deacon JW. (2005). "Fungal Biology" (4th ed). Malden, MA: Blackwell<br />
Publishers. ISBN 1-4051-3066-0.<br />
• Kaminstein D. (2002). Mushroom poisoning.<br />
External links<br />
Wikimedia Commons has media related to:<br />
Fungi<br />
Look up fungi in<br />
Wiktionary, the free dictionary.<br />
Wikispecies has information related to:<br />
fungi<br />
• The WWW Virtual Library: Mycology<br />
• MykoWeb<br />
• Illinois Mycological Association Mycological Glossary
v • d • e<br />
• Tree of Life web project: Fungi<br />
• Fungal Biology, University of Sydney, School of Biological Sciences, June,<br />
2004. – Online textbook<br />
• The Fifth Kingdom – Online textbook<br />
• CABI Bioscience Databases - Includes Index Fungorum genus and species<br />
names and top-down hierarchy<br />
• Comparative Analysis of Fungal Genomes (at DOE's IMG system)<br />
Earth<br />
[hide]<br />
Elements of nature<br />
History of Earth · Earth science · Structure of the Earth · Plate tectonics ·<br />
Geological history of Earth · Geology<br />
Weather Climate · Earth's atmosphere<br />
Life<br />
Biosphere · Origin of life · Microbe · Plants · Fungus · Fauna · Animals ·<br />
Biology · Evolutionary history of life<br />
Environment Wilderness · Ecology · Ecosystem<br />
Universe Matter · Energy · Outer space<br />
Category · Portal<br />
Retrieved from "http://en.wikipedia.org/wiki/Fungus"<br />
Categories: Fungi<br />
Hidden category: Semi-protected<br />
Views<br />
• Article<br />
• Discussion<br />
• View source<br />
• History<br />
Personal tools<br />
• Log in / create account<br />
Navigation<br />
• Main Page<br />
• Contents<br />
• Featured content<br />
• Current events<br />
• Random article<br />
Interaction
An Agar Plate -- an example of a bacterial growth medium. Specifically, it is a streak<br />
plate; the orange lines and dots are formed by bacterial colonies.<br />
A growth medium or culture medium is a liquid or gel designed to support the<br />
growth of microorganisms or cells. [1] There are different types of media for growing<br />
different types of cells. [2]<br />
There are two major types of growth media: those used for cell culture, which use<br />
specific cell types derived from plants or animals, and microbiological culture, which<br />
are used for growing microorganisms, such as bacteria or yeast. The most common<br />
growth media for microorganisms are nutrient broths and agar plates; specialized<br />
media are sometimes required for microorganism and cell culture growth. [1] Some<br />
organisms, termed fastidious organisms, require specialized environments due to<br />
complex nutritional requirements. Viruses, for example, are obligatory intracellular<br />
parasites and require a growth medium composed of living cells.<br />
Contents<br />
[hide]<br />
• 1 Types of growth mediums<br />
o 1.1 Nutrient media<br />
o 1.2 Minimal media<br />
o 1.3 Selective media<br />
o 1.4 Differential media<br />
• 2 Transport media<br />
• 3 Enriched media<br />
• 4 See also<br />
• 5 References<br />
• 6 External links<br />
[edit] Types of growth mediums<br />
This article needs additional citations for verification.<br />
Please help improve this article by adding reliable references. Unsourced material may be<br />
challenged and removed. (August 2007)
The most common growth mediums for microorganisms are nutrient broths (liquid<br />
nutrient medium) or Luria Bertani medium (LB medium or Lysogeny Broth). Liquid<br />
mediums are often mixed with agar and poured into petri dishes to solidify. These<br />
agar plates provide a solid medium on which microbes may be cultured. Bacteria<br />
grown in liquid cultures often form colloidal suspensions.<br />
The differences between growth mediums used for cell culture and those used for<br />
microbiological culture are due to the fact that cells derived from whole organisms<br />
and grown in culture often cannot grow without the addition of, for instance,<br />
hormones or growth factors which usually occur in vivo. [3] In the case of animal cells,<br />
this difficulty is often addressed by the addition of blood serum to the medium. In the<br />
case of microorganisms, there are no such limitations, as they are often unicellular<br />
organisms. One other major difference is that animal cells in culture are often grown<br />
on a flat surface to which they attach, and the medium is provided in a liquid form,<br />
which covers the cells. In contrast, bacteria such as Escherichia coli may be grown on<br />
solid media or in liquid media.<br />
An important distinction between growth media types is that of defined versus<br />
undefined media. [1] A defined medium will have known quantities of all ingredients.<br />
For microorganisms, they consist of providing trace elements and vitamins required<br />
by the microbe and especially a defined carbon source and nitrogen source. Glucose<br />
or glycerol are often used as carbon sources, and ammonium salts or nitrates as<br />
inorganic nitrogen sources). An undefined medium has some complex ingredients,<br />
such as yeast extract or casein hydrolysate, which consist of a mixture of many, many<br />
chemical species in unknown proportions. Undefined media are sometimes chosen<br />
based on price and sometimes by necessity - some microorganisms have never been<br />
cultured on defined media.<br />
A good example of a growth medium is the wort used to make beer. The wort<br />
contains all the nutrients required for yeast growth, and under anaerobic conditions,<br />
alcohol is produced. When the fermentation process is complete, the combination of<br />
medium and dormant microbes, now beer, is ready for consumption.<br />
[edit] Nutrient media<br />
Undefined media (also known as basal or complex media) is an undefined media that<br />
contains:<br />
• a carbon source such as glucose for bacterial growth<br />
• water<br />
• various salts need for bacterial growth<br />
• a source of amino acids and nitrogen (e.g., beef, yeast extract)<br />
This is an undefined medium because the amino acid source contains a variety of<br />
compounds with the exact composition unknown. Nutrient media contain all the<br />
elements that most bacteria need for growth and are non-selective, so they are used<br />
for the general cultivation and maintenance of bacteria kept in laboratory culture<br />
collections.<br />
Defined media (also known as chemical defined media)
• all the chemicals used are known and<br />
• does not contain any animal, yeast, plant tissue.<br />
Differential medium<br />
• some sort of indicator, typically a dye, is added, that allows for the<br />
differentiation of particular chemical reactions occurring during growth.<br />
[edit] Minimal media<br />
Minimal media are those that contain the minimum nutrients possible for colony<br />
growth, generally without the presence of amino acids, and are often used by<br />
microbiologists and geneticists to grow "wild type" microorganisms. Minimal media<br />
can also be used to select for or against recombinants or exconjugants.<br />
Minimal medium typically contains:<br />
• a carbon source for bacterial growth, which may be a sugar such as glucose, or<br />
a less energy-rich source like succinate<br />
• various salts, which may vary among bacteria species and growing conditions;<br />
these generally provide essential elements such as magnesium, nitrogen,<br />
phosphorus, and sulfur to allow the bacteria to synthesize protein and nucleic<br />
acid<br />
• water<br />
Supplementary minimal media are a type of minimal media that also contains a single<br />
selected agent, usually an amino acid or a sugar. This supplementation allows for the<br />
culturing of specific lines of auxotrophic recombinants.<br />
[edit] Selective media<br />
Blood-free, charcoal-based selective medium agar (CSM) for isolation of<br />
Campylobacter.
Blood agar plates are often used to diagnose infection. On the right is a positive<br />
Streptococcus culture; on the left a positive Staphylococcus culture.<br />
Selective mediums are used for the growth of only select microorganisms. For<br />
example, if a microorganism is resistant to a certain antibiotic, such as ampicillin or<br />
tetracycline, then that antibiotic can be added to the medium in order to prevent other<br />
cells, which do not possess the resistance, from growing. Media lacking an amino acid<br />
such as proline in conjunction with E. coli unable to synthesize it were commonly<br />
used by geneticists before the emergence of genomics to map bacterial chromosomes.<br />
Selective growth media are also used in cell culture to ensure the survival or<br />
proliferation of cells with certain properties, such as antibiotic resistance or the ability<br />
to synthesize a certain metabolite. Normally, the presence of a specific gene or an<br />
allele of a gene confers upon the cell the ability to grow in the selective medium. In<br />
such cases, the gene is termed a marker.<br />
Selective growth media for eukaryotic cells commonly contain neomycin to select<br />
cells that have been successfully transfected with a plasmid carrying the neomycin<br />
resistance gene as a marker. Gancyclovir is an exception to the rule as it is used to<br />
specifically kill cells that carry its respective marker, the Herpes simplex virus<br />
thymidine kinase (HSV TK).<br />
Four types of agar plates demonstrating differential growth depending on bacterial<br />
metabolism.
Some examples of selective media include:<br />
• eosin-methylen blue agar (EMB) that contains methylene blue – toxic to<br />
Gram-positive bacteria, allowing only the growth of Gram negative bacteria<br />
• YM (yeast and mold) which has a low pH, deterring bacterial growth<br />
• blood agar (used in strep tests), which contains beef heart blood that becomes<br />
transparent in the presence of hemolytic Streptococcus<br />
• MacConkey agar for Gram-negative bacteria<br />
• Hektoen Enteric (HE) which is selective for Gram-negative bacteria<br />
• Mannitol Salt Agar (MSA) which is selective for Gram-positive bacteria and<br />
differential for mannitol<br />
• xylose lysine desoxyscholate (XLD), which is selective for Gram-negative<br />
bacteria<br />
• Buffered charcoal yeast extract agar, which is selective for certain gramnegative<br />
bacteria, especially Legionella pneumophila<br />
[edit] Differential media<br />
Differential media or indicator media distinguish one microorganism type from<br />
another growing on the same media. [4] This type of media uses the biochemical<br />
characteristics of a microorganism growing in the presence of specific nutrients or<br />
indicators (such as neutral red, phenol red, eosin y, or methylene blue) added to the<br />
medium to visibly indicate the defining characteristics of a microorganism. This type<br />
of media is used for the detection of microorganisms and by molecular biologists to<br />
detect recombinant strains of bacteria.<br />
Examples of differential media include:<br />
• Eosin methylene blue (EMB), which is differential for lactose and sucrose<br />
fermentation<br />
• MacConkey (MCK), which is differential for lactose fermentation<br />
• Mannitol Salt Agar (MSA), which is differential for mannitol fermentation<br />
• X-gal plates, which are differential for lac operon mutants<br />
[edit] Transport media<br />
These are used for the temporary storage of specimens being transported to the<br />
laboratory for cultivation. Such media ideally maintain the viability of all organisms<br />
in the specimen without altering their concentration. Transport media typically<br />
contain only buffers and salt. The lack of carbon, nitrogen, and organic growth factors<br />
prevents microbial multiplication. Transport media used in the isolation of anaerobes<br />
must be free of molecular oxygen.<br />
[edit] Enriched media<br />
Enriched media contain the nutrients required to support the growth of a wide variety<br />
of organisms, including some of the more fastidious ones. They are commonly used to<br />
harvest as many different types of microbes as are present in the specimen. Blood<br />
agar is an enriched medium in which nutritionally rich whole blood supplements the
asic nutrients. Chocolate agar is enriched with heat-treated blood (40-45°C), which<br />
turns brown and gives the medium the color for which it is named.<br />
[edit] See also<br />
• R2a agar<br />
• MRS agar<br />
• Cell biology<br />
[edit] References<br />
1. ^ a b c Madigan M, Martinko J (editors). (2005). Brock Biology of Microorganisms,<br />
11th ed., Prentice Hall. ISBN 0131443291.<br />
2. ^ Ryan KJ, Ray CG (editors) (2004). Sherris Medical Microbiology, 4th ed.,<br />
McGraw Hill. ISBN 0838585299.<br />
3. ^ Cooper GM (2000). "Tools of Cell Biology", The cell: a molecular approach.<br />
Washington, D.C: ASM Press. ISBN 0-87893-106-6.<br />
4. ^ Washington JA (1996). "Principles of Diagnosis", Baron's Medical Microbiology<br />
(Baron S et al, eds.), 4th ed., Univ of Texas Medical Branch. ISBN 0-9631172-1-1.<br />
INTESTINAL PROTOZOA<br />
http://www.tulane.edu/~wiser/protozoology/notes/inte<br />
s.html<br />
Lumen-Dwelling Protozoa<br />
Flagellates:<br />
Giardia lamblia<br />
Dientamoeba fragilis<br />
Chilomastix mesnili<br />
Enteromonas hominis<br />
Retortamonas intestinalis<br />
Trichomonas hominis<br />
Trichomonas tenax (oral)<br />
Trichomonas vaginalis<br />
(urogenital)<br />
Ameba:<br />
Entamoeba histolytica<br />
Entamoeba dispar<br />
Entamoeba coli<br />
Entamoeba hartmanni<br />
Entamoeba polecki<br />
Entamoeba gingivalis (oral)<br />
Endolimax nana<br />
Iodamoeba bütschlii
Apicomplexa:<br />
Cryptosporidium parvum<br />
Cryptosporidium hominis<br />
Cyclospora cayetanensis<br />
Isospora belli<br />
Microsporidia:<br />
Enterocytozoon bieneusi<br />
Encephalitozoon intestinalis<br />
Other:<br />
Blastocystis hominis<br />
Balantidium coli<br />
Numerous protozoa inhabit the gastrointestinal<br />
tract of humans (see Box).<br />
This list includes representatives from<br />
many diverse protozoan groups. The<br />
majority of these protozoa are nonpathogenic<br />
commensals, or only result<br />
in mild disease. Some of these<br />
organisms can cause severe disease<br />
under certain circumstances. For<br />
example, Giardia lamblia can cause<br />
severe acute diarrhea which may lead<br />
to a chronic diarrhea and nutritional<br />
disorders; Entamoeba histolytica can become a highly virulent and<br />
invasive organism that causes a potentially lethal systemic disease.<br />
Apicomplexa and microsporidia species (discussed elsewhere),<br />
which normally do not evoke severe disease, can cause severe and<br />
life-threatening diarrhea in AIDS patients and other<br />
immunocompromised individuals. Trichomonas vaginalis does not<br />
reside within the gastro-intestinal tract, but is often discussed with<br />
the intestinal flagellates. It infects the urogenital tract and and<br />
causes a sexually-transmitted disease.<br />
Intestinal protozoa are transmitted by the fecal-oral route and tend<br />
to exhibit similar life cycles consisting of a cyst stage and a<br />
trophozoite stage (Figure). Fecal-oral transmission involves the<br />
ingestion of food or water contaminated with cysts. After ingestion<br />
by an appropriate host, the cysts transform into trophozoites which<br />
exhibit an active metabolism and are usually motile. The parasite<br />
takes up nutrients and undergoes asexual replication during the<br />
trophic phase. Some of the trophozoites will develop into cysts<br />
instead of undergoing replication. Cysts are characterized by a<br />
resistant wall and are excreted with the feces. The cyst wall<br />
functions to protect the organism from desiccation in the external<br />
environment as the parasite undergoes a relatively dormant period
waiting to be ingested by the next host. Factors which increase the<br />
likelyhood of ingesting material contaminated with fecal material<br />
play a role in the transmission of this intestinal protozoa (see Box).<br />
In general, situations involving close human-human contact and<br />
unhygenic conditions promote<br />
transmission.<br />
TOPICS:<br />
• Giardiasis<br />
o Life Cycle and<br />
Morphology<br />
Trophozoite<br />
Cyst<br />
o The Adhesive Disk<br />
o Symptoms and<br />
Pathogenesis<br />
o Diagnosis<br />
o Treatment and Control<br />
• Trichomoniasis<br />
• Balantidosis<br />
• Amebiasis<br />
• Non-Pathogenic Commensals<br />
GIARDIASIS<br />
Giardia lamblia (also known as G.<br />
duodenalis, see comments on<br />
taxonomy) is a protozoan parasite<br />
that colonizes the upper portions of<br />
the small intestine. It has a<br />
worldwide distribution and is the<br />
most common protozoan isolated<br />
from human stools. The incidence is<br />
estimated at 200 million clinical<br />
cases per year. In fact, it was<br />
probably the first symbiotic<br />
Fecal-Oral Transmission Factors<br />
poor personal hygiene<br />
• children (eg, day-care<br />
centers)<br />
• institutions (eg, prisons,<br />
mental hospitals, orphanages)<br />
• food handlers<br />
developing countries<br />
• poor sanitation<br />
• lack of indoor plumbing<br />
• endemic<br />
• travelers' diarrhea<br />
water-borne epidemics<br />
• water treatment failures<br />
male homosexuality<br />
• oral-anal contact<br />
zoonosis?<br />
• Entamoeba = no<br />
• Cryptosporidium = yes<br />
• Giardia = controversial<br />
protozoan ever observed. It is quite likely that Van Leeuwenhoek,<br />
the inventor of the microscope, first described Giardia in 1681in his<br />
own stools based upon his description of its characteristic<br />
movement. However, van Leeuwenhoek never submitted drawings<br />
of the organisms and Lambl is usually given credit for the<br />
identification of Giardia in the stools of pediatric patients in Praque<br />
in 1859.<br />
Typically Giardia is non-invasive and often results in asymptomatic<br />
infections. Symptomatic giardiasis is characterized by acute or<br />
chronic diarrhea and/or other gastro-intestinal manifestations.
LIFE CYCLE AND MORPHOLOGY<br />
Giardia exhibits a typical fecal-oral transmission<br />
cycle (see above). The infection is acquired<br />
through the ingestion of cysts. Factors leading to<br />
contamination of food or water with fecal<br />
material are correlated with transmission (Box).<br />
For example, giardiasis is especially prevalent in<br />
children and particularly those children in<br />
institutions or day-care centers. In developing<br />
countries, poor sanitation contributes to the<br />
higher levels of giardiasis, and water-borne<br />
outbreaks due to inadequate water treatment<br />
have also been documented. Backpackers in areas of no human<br />
habitation are believed to acquire from drinking from streams and<br />
some data suggest that beavers are the reservoir. However, the<br />
zoonotic transmission of Giardia is controversial and has not been<br />
unambiguously demonstrated. It is not clear whether Giardia<br />
lamblia represents a single species capable of infecting a wide range<br />
of animals, or whether each host has their own 'pet' Giardia.<br />
Evidence indicating that Giardia transmission between dogs and<br />
humans is quite rare favors the latter. Molecular evidence suggests<br />
that some isolates exhibit narrow host ranges whereas others<br />
exhibit wide host ranges (see notes on taxonomy). Regardless of<br />
whether zoonotic transmission is possible, person-to-person<br />
transmission is the most prevalent mode of transmission and the<br />
risk factors are close human contact combined with unhygienic<br />
conditions.<br />
The ingested cyst passes through the stomach and excystation<br />
takes place in the duodenum. Excystation can be induced in vitro by<br />
a brief exposure of the cysts to acidic pH (~2) or other sources of<br />
hydrogen ions. This exposure to the acidic pH mimics the conditions<br />
of the stomach and probably functions as an environmental cue for<br />
the parasite. Flagellar activity begins within 5-10 minutes following<br />
the acid treatment and the trophozoite emerges through a break in<br />
the cyst wall. The breakdown of the cyst wall is believed to be<br />
mediated by proteases. The trophozoite will undergo cytokinesis<br />
(cell division without nuclear replication) within 30 minutes after<br />
emerging from the cyst resulting in two binucleated trophozoites.<br />
The Giardia trophozoite exhibits a characteristic pear, or tear-drop,<br />
shape with bilateral symmetry when viewed from the top (Figure).<br />
It is typically 12-15 µm long, 5-10 µm wide, and 2-4 µm thick.<br />
Characteristic features of the stained trophozoite include: two nuclei<br />
(Nu) with central karyosomes (k), fibrils running the length of the<br />
parasite, and median bodies (MB). The large karyosome and lack of
peripheral chromatin gives the nuclei a halo<br />
appearance. The fibrils are called axonemes (Ax) and<br />
are formed from the proximal regions of the flagella<br />
(Fg) within the body of the trophozoite. The median<br />
bodies are a pair of curved rod-shaped structures<br />
which lie posterior to the nuclei. At the ultrastructural<br />
level the median bodies contain an array of<br />
microtubules. The function of the median bodies is not<br />
known, but most believe they are somehow involved<br />
with the adhesive disk and its formation. An adhesive disk (AD), not<br />
always visible by light microscopy, occupies the ventral side of the<br />
anterior end.<br />
Giardia trophozoites possess four pairs of flagella and are motile.<br />
Three pairs of flagella emerge from the dorsal surface (anterior,<br />
posterior-lateral, caudal) and one pair emerges from the ventral<br />
surface. Trophozoites exhibit a distinctive erratic twisting motion,<br />
sometimes compared to that of a falling leaf. However, the<br />
trophozoites are predominantly found attached to epithelial cells of<br />
the small intestine (especially the duodenum and jejunum) and are<br />
rarely found in stools, except in the cases of severe diarrhea. This<br />
attachment to the intestinal epithelium is mediated by an organelle<br />
on the ventral side of the parasite referred to as the adhesive disk<br />
(see below). The trophozoite absorbs nutrients from the intestinal<br />
lumen via pinocytosis and no specialized feeding organelles have<br />
been described.<br />
The trophic stage is also characterized by an asexual replication.<br />
Both nuclei divide at about the same time and cytokinesis restores<br />
the binucleated state. Each daughter cell receives one copy of each<br />
nuclei. Both nuclei appear equal in regards to gene expression and<br />
other properties.<br />
As an alternative to replication the trophozoite can encyst. During<br />
encystment the parasite rounds up, detaches from the intestinal<br />
epithelium, and secretes a cyst wall. Encystation can also be carried<br />
out in vitro. Optimal induction of encystment is obtained by<br />
depriving the trophozoites of bile at pH 7 followed by an exposure<br />
to high concentrations of bile at pH 7.8. The lack of bile at neutral<br />
pH mimics the conditions under the mucus blanket adjacent to the<br />
intestinal epithelial cells, whereas exposure to high concentrations<br />
of bile at more alkaline pH is analogous to the intestinal lumen.<br />
These studies highlight the extent to which Giardia has adapted to<br />
life within the gastrointestinal tract.<br />
Molecular and ultrastructural studies reveal the synthesis of cyst<br />
wall proteins and the appearance of large secretory vesicles in the<br />
parasite cytoplasm follow the induction of encystment. After cyst
wall formation the parasite undergoes one round of nuclear division<br />
without cytokinesis resulting in four nuclei. These four nuclei (Nu)<br />
are usually located at the anterior end of the cyst (Figure). The<br />
flagella and adhesive disk are lost as the cyst matures, but the<br />
axonemes (Ax) and median bodies (MB) persist. The distinctive<br />
fibrils (ie, axonemes), which extend across the length of the cyst,<br />
result in Giardia being relatively easy to unambiguously identify.<br />
The cysts are oval shaped and typical measure 11-14 µm in length<br />
and 6-10 µm wide. Other characteristics of Giardia cysts include a<br />
well-defined wall (CW) which is often set apart from the cytoplasm<br />
of the parasite. The cysts are passed in the feces and can survive<br />
for up to three months under appropriate temperature and moisture<br />
conditions. Mature cysts are infective to the next host that happens<br />
to ingest them, thus completing the life cycle.<br />
THE ADHESIVE DISK<br />
A unique ultrastructural feature of Giardia is the adhesive disk (also<br />
called ventral disk, sucking disk, sucker, or striated disk). The<br />
adhesive disk is a concave structure which occupies approximately<br />
two-thirds of the anterior end of the ventral surface (Figure, left<br />
panel). As the names imply, this structure plays a role in the<br />
attachment of the trophozoite to the intestinal epithelium and<br />
ultrastructural studies reveal close associations between the<br />
adhesive disk and the intestinal brush border (Figure, upper right<br />
panel). (Click here for larger image.)<br />
The adhesive disk appears to be<br />
a relatively rigid structure and<br />
striations are evident by<br />
transmission electron<br />
microscopy. These striations are<br />
the result of microtubules (mT)<br />
and a unique cytoskeletal<br />
element called microribbons<br />
(mR). Microribbons are long<br />
flattened structures and each<br />
microribbon is associated with a<br />
microtubule (Figure, middle<br />
right panel). The combined microtubule-microribbon structure are<br />
arranged in concentric rows that form a flatten spiral with minimal<br />
overlap. The outer rim of the adhesive disk, called the lateral crest,<br />
contains components of the actin-myosin cytoskeleton.<br />
A major component of microribbons are proteins called giardins<br />
(aka beta-giardins). These giardins play primarily a structural role in<br />
the formation of the microribbons. Interestingly, the giardins show
a limited homology to a protein called 'striated fibre assemblin' from<br />
Chlamydomonas (a free-living, bi-flagellated unicellular algae). In<br />
Chlamydomonas this protein forms filamentous structures at the<br />
base of the flagella. The giardins have evolved to play a different<br />
functional role in Giardia, but are still associated with microtubule<br />
based cytoskeletal elements.<br />
This association of proteins involved in the generation of contractile<br />
force and other cytoskeletal elements in the adhesive disk suggests<br />
that attachment is mediated by mechanical forces generated by the<br />
parasite. The observation that imprints and circular dome-shaped<br />
lesions remain in the intestinal brush border (ie, microvilli) following<br />
detachment of trophozoites (Figure, lower right panel) is consistent<br />
with contractile forces playing a role in attachment. Other proposed<br />
mechanisms for the attachment of Giardia to the intestinal<br />
epithelium include hydrodynamic forces generated by the ventral<br />
flagella and receptor-mediated binding via lectins on the trophozoite<br />
surface. However, flagellar movement is poorly correlated with<br />
attachment and the surface lectins cover the entire trophozoite and<br />
are not specifically localized to the adhesive disk.<br />
SYMPTOMS AND PATHOGENESIS<br />
The clinical features associated with Giardia infection range from<br />
total latency (ie, asymptomatic), to acute self-resolving diarrhea, to<br />
chronic syndromes associated with nutritional disorders, weight loss<br />
and failure to thrive. Children exhibit clinical symptoms more<br />
frequently that adults and subsequent infections tend to be less<br />
severe than initial infections. The incubation period is generally 1-2<br />
weeks, but ranges of 1-75 days have been reported.<br />
The first signs of acute giardiasis include nausea, loss of appetite<br />
and an upper gastro-intestinal uneasiness. These signs are often<br />
followed or accompanied by a sudden onset of explosive, watery,<br />
foul-smelling diarrhea. Stools associated with Giardia infection are<br />
generally described as loose, bulky, frothy and/or greasy with the<br />
absence of blood or mucus, which may help distinguish giardiasis<br />
from other acute diarrheas. Other gastro-intestinal disturbances<br />
associated with giardiasis include: flatulence, bloating, anorexia,<br />
cramps, and foul sulfuric belching (sometimes called 'purple burbs').<br />
The acute stage usually resolves spontaneously in 3-4 days and is<br />
often not recognized as being giardiasis. Occasionally, though, an<br />
acute infection will persist and lead to malabsorption, steatorrhea<br />
(excessive loss of fat in the feces), debility (loss of strength) and<br />
weight loss. Some of the individuals who resolve the acute<br />
symptoms do not clear the infection, but become asymptomatic cyst
passers without clinical manifestations, whereas others may have a<br />
few sporadic recurrences of the acute symptoms.<br />
Acute infections can also develop into long-standing subacute or<br />
chronic infections which in rare cases last for years. The typical<br />
chronic stage patient presents with recurrent brief episodes of loose<br />
foul stools which may be yellowish, frothy and float, accompanied<br />
by intestinal gurgling, abdominal distention and flatulence. Between<br />
episodes the stools are usually mushy, but normal stools or<br />
constipation can also occur. Cramps are uncommon during chronic<br />
infections, but sulfuric belching is frequent. Anorexia, nausea, and<br />
epigastric uneasiness are additional frequent complaints during<br />
chronic infections. In the majority of chronic cases the parasites and<br />
symptoms spontaneously<br />
disappear.<br />
The specific mechanisms of Giardia<br />
pathogenesis leading to diarrhea<br />
and intestinal malabsorption are not<br />
completely understood and no<br />
Click for larger image<br />
specific virulence factors have been<br />
identified. Attachment of trophozoites to the brush border could<br />
produce a mechanical irritation or mucosal injury. In addition,<br />
normal villus structure is affected in some patients. For example,<br />
villus blunting (atrophy) and crypt cell hypertrophy and an increase<br />
in crypt depth have been observed to varying degrees. The increase<br />
in crypt cells will lead to a repopulation of the intestinal epithelium<br />
by relatively immature enterocytes with reduced absorptive<br />
capacities. An increased inflammatory cell infiltration in the lamina<br />
propria has also been observed and this inflammation may be<br />
associated with the pathology. Giardia infection can also lead to<br />
lactase deficiency (see lactose intolerance below) as well as other<br />
enzyme deficiencies in the microvilli. This reduced digestion and<br />
absorption of solutes may lead to an osmotic diarrhea and could<br />
also explain the malabsorption syndromes. Thus far, no single<br />
virulence factor or unifying mechanism explains the pathogenesis of<br />
giardiasis. [See also Pathophysiology of Diarrhea for a general<br />
discussion of diarrhea.]<br />
Post-Giardia Lactose Intolerance. Some patients may present<br />
with a lactose intolerence during active Giardia infections which can<br />
persist after parasite clearance. This clinical manifestation is due to<br />
the parasite-induced lactase deficiency and is most common in<br />
ethnic groups with a predisposition for lactase deficiency. Lactase is<br />
an enzyme that breaks down lactose, a sugar found in milk, to<br />
monosaccharides which can be absorbed. This lactose intolerence<br />
syndrome should be considered in persons who still present mushy
stools and excessive gas following treatment, but have no<br />
detectable parasites.<br />
DIAGNOSIS<br />
Parasite Detection<br />
Stool Examination<br />
• 3 non-consecutive days<br />
• wet mount or stained<br />
• IFA, copro-antigens<br />
Duodenal Aspirate or Biopsy<br />
• Enterotest®<br />
Diagnosis is confirmed by finding cysts<br />
or trophozoites in feces or in<br />
duodenojejunal aspirates or biopsies.<br />
Detection of the parasites can be difficult<br />
since Giardia does not appear<br />
consistently in the stools of all patients.<br />
Some patients will express high levels of<br />
cysts in nearly all the stools, whereas<br />
others will only exhibit low parasite<br />
counts in some of the stools. A mixed<br />
pattern, in which periods of high cyst<br />
excretion alternate with periods of low<br />
excretion, has also been observed. In<br />
addition, parasites are easier to find during acute infections than<br />
chronic infections. Aspiration and biopsy may also fail to confirm the<br />
infection due to patchy loci of infection, and some question the<br />
usefulness of these invasive procedures.<br />
Stool examination is the preferred method for Giardia diagnosis.<br />
Three stools taken at intervals of at least two days should be<br />
examined. Watery or loose stools may contain motile trophozoites<br />
which are detectable by the immediate examination of wet smears.<br />
Otherwise the specimen should be preserved and stained due to<br />
trophozoite lability. The hardier cysts are relatively easy to<br />
recognize in either direct or stained smears (see cyst morphology).<br />
In addition, diagnostic kits based on immunofluorescence or the<br />
detection of copro-antigens are also available.<br />
Diagnosis can also be made by examining duodenal fluid for<br />
trophozoites. Duodenal fluid is obtained by either intubation or the<br />
Enterotest® (also called 'string test'). The Enterotest® consists of a<br />
gelatin capsule containing a nylon string of the appropriate length.<br />
The free end of the string is taped to the patient's face and the<br />
capsule is swallowed. After four hours to overnight the string is<br />
retrieved and the bile-stained mucus on the distal portion of the<br />
string is scraped off and examined by both wet mount and<br />
permanent staining. A small intestinal biopsy, preferably from<br />
multiple duodenal and jejunal sites, may also reveal trophozoites<br />
attached to the intestinal epithelium. [The small intestine is divided<br />
into 3 sections: the duodenum (first or proximal portion after the<br />
stomach); the jejunum (the middle portion); and the ileum (the<br />
distal or last portion before the large intestine).]
TREATMENT AND CONTROL<br />
Infected individuals should be treated since Giardia can persist and<br />
lead to severe malabsorption syndromes and weight loss. Treatment<br />
is effective at reducing morbidity and there are no sequelae.<br />
Metronidazole (Flagyl®), although not licensed in the United States<br />
for giardiasis, effectively clears the parasite (cure rates<br />
approximately 85%) and is the drug of choice. The recommended<br />
dosage is 750 mg three times per day for five days (or at least >3<br />
days). For children 15 mg/kg/d in three doses is recommended.<br />
Other effective drugs include: quinacrine (Atabrine®), tinidazole<br />
(Fasigyn®), furazolidone (Furoxone®), and paramomycin<br />
(Humatin®). Tinidazole is effective as a single two gram dose;<br />
paramomycin is not absorbed and may be useful during pregnancy.<br />
The widespread distribution of Giardia and the infectivity of the<br />
cysts make it unlikely that human infection will be completely<br />
eliminated. Control measures to prevent or reduce Giardia infection<br />
will depend on the specific circumstances of transmission, but in<br />
general involve measures which prevent the ingestion of substances<br />
contaminated with fecal material (see fecal-oral transmission<br />
factors). Health promotion and education aimed at improving<br />
personal hygiene, and emphasizing hand washing, sanitation and<br />
food handling, are effective control activities for the reduction of<br />
person-to-person transmission. Special attention to personal<br />
hygiene in high-risk situations such as day-care centers and other<br />
institutions is needed. Treatment of asymptomatic household<br />
members prevents reinfection in non-endemic areas. However, the<br />
value of treating asymptomatic carriers in hyperendemic<br />
communities is questionable since reinfection rates are high. The<br />
socio-economic situation in many developing countries makes it<br />
difficult to prevent infection. Public health measures to protect<br />
water supplies from contamination are required to prevent<br />
epidemics and to reduce endemicity. Tourists should not drink tap<br />
water without additional treatment in places where purity is<br />
questionable. Boiling or iodine treatment kills Giardia cysts, but<br />
standard chlorination does not. There are no safe or effective<br />
chemoprophylatic drugs for giardiasis.<br />
TRICHOMONIASIS<br />
• Tricomonad Morphology and Species<br />
• Transmission and Life Cycle<br />
• Symptoms and Pathogenesis<br />
• Diagnosis, Treatment and Control
The trichomonads are a group of flagellated protozoa. Most of the<br />
members of this group are parasitic and only a few free-living<br />
species have been identified. Generally the trichomonads are nonpathogenic<br />
commensals and only a few species are of importance in<br />
animals and humans. Four species of trichomonads infect humans<br />
(Table). Among these only Trichomonas vaginalis is clearly<br />
pathogenic and it is usually of low virulence. The others exhibit a<br />
questionable pathogenicity.<br />
Trichomonads of Humans The trichomonads of humans<br />
inhabit different anatomical<br />
Species Location locations. T. vaginalis is a<br />
Trichomonas vaginalis uro-genital tract common sexually transmitted<br />
Trichomonas tenax oral cavity disease found in the urogenital<br />
tract. T. tenax, also<br />
Pentatrichomonas hominis intestine<br />
called T. buccalis, is a<br />
Dientamoeba fragilis intestine<br />
commensal of the human oral<br />
cavity, found particularly in<br />
patients with poor oral hygiene and advanced periodontal disease.<br />
T. tenax, or an organism with similar morphology is also<br />
occasionally found in the lungs. Such cases have reported mainly in<br />
patients with underlying cancers or other lung diseases or following<br />
surgery. Pentatrichomonas hominis, formerly known as<br />
Trichomonas hominis, is a non-pathogenic commensal of the large<br />
intestine (see non-pathogenic intestinal flagellates). Some authors<br />
divide the trichomonads into three genera based on the number of<br />
free flagella. Species with three flagella are called Tritrichomonas,<br />
those with four are called Trichomonas, and Pentatrichomonas<br />
refers to trichomonads with five free anterior flagella. Dientamoeba<br />
fragilis was originally believed to be an ameba (see non-pathogenic<br />
intestinal ameba). Now it is know to be a flagellate—however<br />
without flagella—related to the trichomonads.<br />
A distinctive feature of the trichomonads is an axostyle (ax) which<br />
runs the length of the organism and appears to protrude from the<br />
posterior end (Figure). The axostyle is a cytoskeletal element<br />
composed of concentric rows of microtubules and is believed to<br />
function in the attachment of the parasite to epithelial cells.<br />
Trichomonads are also characterized by 4-6 flagella (fg) emerging<br />
from the anterior end. One of the flagella is attached to the body of<br />
the organism and forms a posteriorly-directed undulating<br />
membrane (um), whereas the remaining flagella are free. The<br />
combined basal bodies (bb) and the base of the undulating<br />
membrane, called the costa (cs), are often seen is stained<br />
preparations. Less frequently seen is the cytostomal groove (cy). A<br />
single nucleus (nu) is found at the anterior end of the parasite.
Schematic representation of major structural features of trichmonads (left). Giemsa-stained trophoz<br />
vaginalis from in vitro culture (middle). Electron micrograph of axostyle cross-section showing conce<br />
of microtubules (right).<br />
The trichomonads, like many other intestinal protozoa, exhibit an<br />
anerobic metabolism and lack mitochondria. Part of energy<br />
metabolism of trichomonads involves a unique organelle called the<br />
hydrogenosome. The hydrogenosome has a double membrane and<br />
is distantly related to the mitochondrion. However, it lacks DNA,<br />
cytochromes and many typical mitochnondrial functions such as<br />
enzymes of the tricarboxylic acid cycle and oxidative<br />
phosphorylation. The primary function of the hydrogenosome is the<br />
metabolism of pyruvate, produced during glycolysis within the<br />
cytosol, to acetate and carbon dioxide with the concomitant<br />
production of ATP. The electrons release from the oxidation of<br />
pyruvate are transferred to hydrogen ions to produce molecular<br />
hydrogen, hence the name hydrogenosome.<br />
TRICHOMONAS VAGINALIS<br />
Trichomonas vaginalis was first described from purulent vaginal<br />
discharges in 1836 and by the early part of the twentieth century<br />
was recognized as an etiological agent of vaginitis. Trichomoniasis is<br />
a common sexually transmitted disease with a worldwide<br />
distribution and an estimated 167 million people becoming infected<br />
per year worldwide and 5 million new infections per year in the<br />
United States. Trichomoniasis is believed to be the most common<br />
non-viral sexually transmitted disease. Despite the frequency of
trichomoniasis it has in the past been considered more of a<br />
nuisance parasite rather than a major pathogen. However it is now<br />
recognized a factor in promoting HIV infection (see Box), causing<br />
low-weight and premature births, and predisposing women to<br />
substantial discomfort and stress.<br />
Trichomonas and HIV<br />
The pathology caused by Trichomonas may enhance the efficiency of HIV transmission<br />
(1). T. vaginalis infection typically elicits a local cellular immune response with<br />
inflammation of the vaginal epithelium and cervix in women and the urethra of men. This<br />
inflammatory response includes the infiltration of potential HIV target cells such as CD4+<br />
bearing lymphocytes and macrophages. In addition, T. vaginalis can cause punctate<br />
hemorrhages on the vaginal walls and cervix. This leukocyte infiltration and the genital<br />
lesions may increase the number of target cells for the virus and allowing direct viral<br />
access to the bloodstream through open lesions. In addition, the hemorrhages and<br />
inflammation can increase the level of virus in body fluids and the numbers of HIVinfected<br />
lymphocytes and macrophages present in the genital area in persons already<br />
infected with HIV. This increase of free virus and virus-infected leukocytes can increase<br />
the probability of HIV exposure and transmission to an uninfected partner. Increased<br />
cervical shedding of HIV has been shown to be associated with cervical inflammation, and<br />
substantially increased viral loads in semen have been documented in men with<br />
trichomoniasis. Moreover, since many patients with Trichomonas infection are<br />
asymptomatic, or only mildly symptomatic, they are likely to remain sexually active in<br />
spite of infection.<br />
1. Sorvillo F, Smith L, Kerndt P, Ash L. (2001) Trichomonas vaginalis, HIV, and<br />
African-Americans. Emerg Infect Dis. 7:927-32.<br />
T. vaginalis, despite its name, infects both men and women. In<br />
females the organism primarily inhabits the vagina, and in males it<br />
is usually found in the urethra, prostate or epididymis. The life cycle<br />
consists only of a trophozoite stage which is transmitted by direct<br />
contact during sexual intercourse. Non-venereal transmission is<br />
rare, but possible since the trophozoites can survive 1-2 days in<br />
urine and 2-3 hours on a wet sponge. In addition, neonatals have<br />
been infected during the birth process. The trophozoites live closely<br />
associated or attached to the epithelium of the urogenital tract,<br />
where they replicate by binary fission.
SYMPTOMS AND PATHOGENESIS<br />
Clinical Manifestations<br />
Females Males<br />
• asymptomatic<br />
(15-20%*)<br />
• vaginal discharge<br />
(50-75%*)<br />
• dyspareunia<br />
(50%*)<br />
• pruritus (25-<br />
50%*)<br />
*% of infected; **% of symptomatic<br />
• asymptomatic<br />
(50-90%*)<br />
• urethral discharge<br />
(50-60%**)<br />
• dysuria (12-<br />
25%**)<br />
• urethral pruritus<br />
(25%**)<br />
T. vaginalis causes<br />
different clinical<br />
manifestations in men<br />
and women and women<br />
(Table) are more likely<br />
to exhibit symptoms<br />
which tend to persist<br />
longer. The incubation<br />
period typically ranges<br />
from 4-28 days. In<br />
females the infection<br />
can present as a mild<br />
vaginitis, an acute or<br />
chronic vulvovaginitis,<br />
or urethritis. The onset<br />
or exacerbation of symptoms commonly occurs during or<br />
immediately after menstration. The most common complaint<br />
associated with T. vaginalis infection is a persistent mild vaginitis<br />
associated with a copious, foul-smelling discharge that is often<br />
accompanied by burning or itching. This discharge is most often<br />
gray, but can be yellow or green and is occasionally frothy or blood<br />
tinged. The discharge diminishes as the infection becomes more<br />
chronic. Many women also experience painful or difficult coitus.<br />
Urethral involvement occurs in a large number of cases and is<br />
characterized by dysuria (painful urination) and frequent urination.<br />
The vaginal epithelium is the primary site of infection. Thus the<br />
vaginal walls are usually erythematous (i.e., red) and may show<br />
petechial (a small non-raised spot) hemorrhages. Punctate<br />
hemorrhages of the cervix, called strawberry cervix, are observed in<br />
approximately 2% of the cases. This strawberry cervix is a<br />
distinctive pathological observation associated with trichomonasis<br />
not seen with other sexually transmitted diseases.<br />
Males are likely to be asymptomatic (50-90%) and the infection<br />
tends to be self-limiting. The urethra and prostate are the most<br />
common sites of infection. Common symptoms include: urethral<br />
discharge (ranging from scant to purulent), dysuria, and urethral<br />
pruritus (itching). Some men experience burning immediately after<br />
coitus.<br />
Little is known about the pathophysiology associated with T.<br />
vaginalis infection, but is presumably due to interactions between<br />
the parasite and host epithelial cells. In vitro studies indicate that T.<br />
vaginalis can destroy cells in a contact dependent manner.<br />
Therefore adhesion of the trophozoites to the epithelium is believed
to be a major factor in the pathogenesis. Several adhesion proteins<br />
have been identified on the surface of the trophozoites. In addition,<br />
secreted proteases that could play a role in pathogenesis have also<br />
been identified.<br />
DIAGNOSIS, TREATMENT AND CONTROL<br />
In general, the clinical manifestations are not reliable as sole means<br />
of diagnosis since the clinical presentation is similar to other STDs<br />
and many patients have mild or no symptoms. Diagnosis is<br />
confirmed by the demonstration of trophozoites in vaginal, urethral,<br />
prostatic secretions, or urine sediment (following prostate<br />
massage). Microscopic examination of wet mounts of fresh vaginal<br />
discharge, preferably collected with a speculum on a cotton-tipped<br />
applicator, is the most practical method of diagnosis. Specimens<br />
should be diluted in saline and examined immediately. T. vaginalis<br />
is recognized by its characteristic morphological features (see<br />
above) and its rapid jerky motility. Specimens can also be fixed and<br />
stained with Giemsa or fluorescent dyes. However, the organism<br />
may be difficult to recognize on stained slides.<br />
The sensitivity of direct observation ranges from 40-80%.<br />
Therefore, in vitro culture is considered the gold standard for<br />
diagnosis despite some limitations. For example, access to facilities<br />
is needed and organisms require 2-7 days of growth before they are<br />
detected. The accessibility issue is partly resolved by the<br />
InPouchTV culture system (Biomed Diagnostics). This is a<br />
commercially available self-contained system for the detection of T.<br />
vaginalis in clinical specimens. Antibody and DNA-based tests with<br />
high sensitivity and specificity are being developed.<br />
Metronidazole (Flagyl®) and other nitroimidazoles, such as<br />
tinidazole, are highly effective against trichomoniasis. The<br />
metronidazole is activated by the hydrogensome to a nitro radical<br />
ion intermediate. Either a single two gram dose (85-92% cure rate)<br />
or 250 mg three time daily for 7-10 days (>95% cure rate) can be<br />
used. Sexual partners should be treated at the same time to<br />
prevent reinfection. Some drug resistance has been reported, but<br />
this is not a wide-spread problem. Treatment failures are generally<br />
due to noncompliance or reinfection.<br />
Trichomoniasis as an STD<br />
• 5% females attending<br />
family planning clinics<br />
• 7-32% females attending<br />
venereal disease clinics<br />
• 50-75% prostitutes
The epidemiology of trichomonasis<br />
exhibits features similar to other<br />
sexually transmitted diseases (Box)<br />
and incidence correlates with the<br />
number of sexual partners. In<br />
addition, co-infection with other STDs<br />
is common. It is estimated that up to 25% of sexually active women<br />
will become infected at some point during their lives and the<br />
disease will be transmitted to 30-70% of their male partners.<br />
Measures used in the control of other STD, such as limiting number<br />
of sexual partners and use of condoms, are also effective in<br />
preventing trichomoniasis.<br />
Reviews on Trichomoniasis:<br />
• Lehker, M.W. and Alderete, J.F. (2000) Biology of<br />
trichomonosis. Current Opinion in Infectious Diseases 13, 37-<br />
45.<br />
• Petrin, D., Delgaty, K., Bhatt, R., Garber, G. (1998) Clinical<br />
and microbiological aspects of Trichomonas vaginalis. Clin.<br />
Microbiol. Rev. 11: 300-317.<br />
• Schwebke, J.R. and Burgess, D. (2004) Trichomoniasis.<br />
Clinical Microbiology Reviews 17, 794-803.<br />
DIENTAMOEBA FRAGILIS<br />
• 4% males attending<br />
venereal disease clinics<br />
• 5-15% males with nongonococcal<br />
urethritis<br />
Dientamoeba fragilis was originally described as an ameba based<br />
upon its morphology. However, later it was recognized to exhibit a<br />
morphology more similar to the turkey parasite Histomonas<br />
meleagridis, except for the lack of flagella. Ultrastructural studies<br />
also suggest similarities to the trichomonads, including the<br />
possession of hydrogenosomes and molecular studies have<br />
confirmed a close phylogenetic relationship between Dientamoeba<br />
and Histomonas and a possible more distal relationship to<br />
Trichomonas.<br />
As with other trichomonads, Dientamoeba only exhibits a<br />
trophozoite stage (Figure). This raises some questions about the<br />
mode of transmission in that a cyst stage is usually involved in fecal<br />
oral transmission. In addition, the trophozoites of Dientamoeba<br />
survive outside of the body for a very short time. H. meleagridis<br />
also lacks a cyst stage and has been demonstrated to be<br />
transmitted via the eggs of a nematode. Due to the close<br />
relationship between Histomonas and Dientamoeba, it is proposed<br />
that Dientamoeba is also transmitted via helminth eggs.<br />
Epidemiological and experimental evidence tends to incriminate the<br />
pinworm Enterobius vermicularis as the carrier for Dientamoeba.
Morphology of Dientamoeba fragilis from a stool sample. Trophozoites exhibit an amebalike<br />
morphology and are often bi-nucleated.<br />
Historically Dientamoeba has been considered as a non-pathogenic<br />
commensal. However, clinical symptoms often correlate with the<br />
presence of large numbers of trophozoites and treatment of the<br />
infection resolves the symptoms. The incidence of symptoms is<br />
estimated at 15-30% of infected individuals. Clinical symptoms<br />
associated with Dientamoeba include intermittent diarrhea,<br />
abdominal pain, flatulence, nausea and fatigue. Little is known<br />
about the pathogenesis and Dientamoeba probably acts as a lowgrade<br />
irritant of intestinal mucosal surfaces that may lead to some<br />
inflammation. Iodoquinol is generally the drug of choice for the<br />
treatment of Dientamoeba. Tetracycline, paromomycin, and<br />
metronidazole are also effective.<br />
For a comprehensive review of Dientamoeba see: Johnson et al,<br />
Clin. Microbiol. Rev. 17:553, 2004.
BALANTIDOSIS<br />
Balantidium coli is the only ciliate which infects<br />
humans. It is found world wide, but like many other<br />
fecal-oral transmitted diseases, it is more prevalent<br />
in the tropics. However, prevalence rates rarely<br />
exceed 1%. B. coli also infects a wide variety of<br />
mammals and is especially common in monkeys and<br />
pigs. Prevalence in pigs ranges from 20–100% and<br />
human balantidiosis usually exhibits an increased<br />
prevalence in communities that live in close<br />
association with pigs. For example, in Papua New<br />
Guinea, where pigs are the principal domestic<br />
animals, the prevalence among swine herders and<br />
slaughterhouse workers has been reported to be as<br />
high as 28%. Human-to-human transmission has also been<br />
documented and this mode of transmission is likely to occur in<br />
environments with crowding and poor personal hygiene such as<br />
mental hospitals and prisons. (Skip general ciliate biology)<br />
GENERAL CILIATE BIOLOGY<br />
Ciliates are a large and diverse group of protozoa. Most ciliates are<br />
free-living and are found in a variety of habitats. Well-known<br />
ciliates include Paramecium species, which are found in ponds<br />
throughout the world, and Ichthyophthirius multifiliis, an<br />
ectoparasite of fish that causes white spot disease (also called 'ick').<br />
As the name implies, ciliates possess cilia at some point during their<br />
life cycles. The cilia are generally arranged in longitudinal rows and<br />
typically cover the surface of the organism. Ciliates are also<br />
characterized by nuclear dimorphism in that they have two distinct<br />
nuclei. The large kidney-shaped macronucleus is involved in the<br />
'housekeeping' or somatic functions of the cell, whereas the smaller<br />
spherical micronucleus contains the complete genome. The<br />
macronucleus contains thousands of copies of transcriptionally<br />
active 'minichromosomes' representing 10-20,000 different DNA<br />
molecules. This large number of telomeres (chromosome ends)<br />
resulted in ciliates being an early model system for the study of<br />
telomeres and telomerase (the enzyme that synthesizes telomeres).
Ciliates undergo both an asexual reproduction (ie, binary fission)<br />
and a sexual reproduction involving conjugation (Figure above).<br />
During conjugation, two ciliates of opposite mating types pair and<br />
exchange genetic material. Conjugal contact triggers meiosis in the<br />
micronuclei resulting in 4 haploid micronuclei. Concurrently, the<br />
macronucleus breaks down and disappears. Three of the micronuclei<br />
disintegrate and the remaining micronucleus divides again. Each of<br />
the conjugating organisms donates a micronucleus (gametic or<br />
male) to its mate via a cytoplasmic bridge that connects them. The<br />
gametic micronucleus fuses with the stationary (or female)<br />
micronucleus forming the diploid zygotic micronucleus. The<br />
conjucating pair separates and the zygotic nucluei undergo another<br />
round of division. One of these micronuclei develops into the the<br />
macronucleus, thus completing the cycle. Formation of the<br />
macronucleus involves fragmentation of the chromosomes and loss<br />
of some DNA sequences. The remaining minichromosomes are then<br />
amplified. (See diagram of DNA processing during macronucleus<br />
formation.)<br />
BALANTIDOSIS<br />
B. coli usually lives as a non-pathogenic commensal in the large<br />
intestine and produces no symptoms. Superficial inflammation of<br />
the colonic mucosa may occur which can result in diarrhea and<br />
colicky pain. Mild or chronic infections are characterized by<br />
intermittent diarrhea and constipation, weight loss, and abdominal<br />
pain. On rare occasions the trophozoites will invade the intestinal<br />
epithelium and produce ulceration. Clinically this results in an acute
diarrhea with mucus and blood (ie, dysentery). This balantidial<br />
dysentery is similar to the dysentery produced by Entameoba<br />
histolytica (see below). Rare extra-intestinal infections involving<br />
lungs, vagina, ureter and urinary bladder and intestinal perforations<br />
leading to peritonitis have been reported.<br />
Laboratory diagnosis is made by identifying the organism in feces.<br />
Balantidium exhibits a typical fecal-oral life cycle consisting of<br />
trophozoite and cyst stages. The large size and unique<br />
morphological features of Balantidium (Figure) precludes its<br />
confusion with any other protozoa found in human feces. The<br />
trophozoite is ovoid and has an average size of 70 x 45 µm, but can<br />
range upwards to 150-200 µm. The cyst has a distinctive cyst wall<br />
(CW) and is more spherical with an average diameter of 55 µm. In<br />
stained specimens the most obvious internal structure is the large<br />
macronucleus (maN). The micronucleus (miN) may not always be<br />
apparent because of its close association with the macronucleus.<br />
Contractile vacuoles (CV), which function in osmotic regulation, are<br />
often visible and occasionally the cytostome (Cy) is detectable.<br />
Similar to many other ciliates, Balantidium is covered by rows of<br />
cilia. The cilia give the parasite surface a fuzzy appearance and are<br />
less pronounced in the cyst stage.<br />
The treatment of choice is tetracycline given at 500 mg four times<br />
per day for 10 days. Iodoquinol is the recommended alternate drug.<br />
Metronidazole has not produced consistent results. Preventive<br />
measures are the same as other diseases transmitted by the fecaloral<br />
route (see fecal-oral transmission factors or discussion of<br />
Giardia prevention). In addition, pig sewerage should be kept away<br />
from supplies of drinking water and food.<br />
AMEBIASIS<br />
Several members of the genus Entamoeba infect humans (see<br />
below). Among these only E. histolytica is considered pathogenic<br />
and the disease it causes is called amebiasis or amebic dysentery.<br />
E. dispar is morphologically identical to E. histolytica and the two<br />
were previously considered to be the same species. However,<br />
genetic and biochemical data indicate that the non-pathogenic E.<br />
histolytica is a distinct species (see discussion of criteria). The two<br />
species are found throughout the world, but like many other<br />
intestinal protozoa, they are more common in tropical countries or<br />
other areas with poor sanitary conditions. It is estimated that up to<br />
10% of the world's population may be infected with either E.<br />
histolytica or E. dispar and in many tropical countries the<br />
prevalence may approach 50%. There are an estimated 50 million<br />
cases of amebiasis per year and up to 100,000 deaths.
• Life Cycle and Morphology<br />
• Pathogenesis<br />
• Possible Mechanisms of Pathogenisis<br />
o Schematic Figure of Trophozoite<br />
Invasion<br />
• Clinical Presentation<br />
• Diagnosis, Treatment and<br />
Control<br />
LIFE CYCLE AND MORPHOLOGY<br />
E. histolytica exhibits a typical fecal-oral life cycle consisting of<br />
infectious cysts passed in the feces and trophozoites which replicate<br />
within the large intestine. The infection is acquired through the<br />
ingestion of cysts and the risk factors are similar to other diseases<br />
transmitted by the fecal-oral route (see Table). Contaminated food<br />
and water are probably the primary sources of infection. The higher<br />
prevalence in areas of lower socioeconomic status is likely due to<br />
poor sanitation and a lack of indoor plumbing. However, E.<br />
histolytica is rarely the cause of travelers' diarrhea and is usually<br />
associated with a long-term (>1 month) stay in an endemic area. A<br />
higher prevalence of E. histolytica infection is also observed in<br />
institutions, such as mental hospitals, orphanages and prisons,<br />
where crowding and problems with fecal contamination are<br />
contributing factors. A high prevalence among male homosexuals<br />
has also been noted. Humans are the only host of E. histolytica and<br />
there are no animal reservoirs.<br />
Upon ingestion the cysts pass through the stomach and excyst in<br />
the lower portion of the small intestine. Excystation involves a<br />
disruption of the cyst wall and the quadranucleated ameba emerges<br />
through the opening. The ameba undergoes another round of<br />
nuclear division followed by three successive rounds of cytokinesis<br />
(ie, cell division) to produce eight small uninucleated trophozoites,<br />
sometimes called amebulae. These immature trophozoites colonize<br />
the large intestine, especially the cecal and sigmoidorectal regions,<br />
where they feed on bacteria and cellular debris and undergo<br />
repeated rounds of binary fission.<br />
E. histolytica trophozoites have an amorphous shape and are<br />
generally 15-30 µm in diameter. The trophozoites move by<br />
extending a finger-like pseudopodium (psd) and pulling the rest of<br />
the body forward (called ameboid movement). The pseudopodia,<br />
and sometimes the outer edge of the trophozoite, have a clear<br />
refractile appearance and is referred to as the ectoplasm (ecto). The<br />
rest of the cytoplasm has a granular appearance and is called the<br />
endoplasm (endo). Occasionally a glycogen vacuole (vac) is evident.
Nuclear (Nu) morphology in stained specimens<br />
is characterized by a finely granular ring of<br />
peripheral chromatin and a centrally located<br />
karyosome (ka).<br />
As an alternative to asexual replication<br />
trophozoites can also encyst. The factors<br />
responsible for the induction of encystation<br />
are not known. Encystation begins with the<br />
trophozoites become more spherical and the<br />
appearance of chromatoid bodies in the<br />
cytoplasm. Chromatoid bodies (cb) are stained<br />
elongated structures with round ends and<br />
represent the aggregation of ribosomes. The cyst wall is composed<br />
of chitin and has a smooth refractile appearance. Cyst maturation<br />
involves two rounds of nuclear replication without cell division and<br />
cysts with 1-4 nuclei (Nu) are found in feces. The nuclear<br />
morphology of the cyst is similar to that of the trophozoite except<br />
that the nuclei become progressively smaller following each<br />
division. Sometimes the young cysts (ie, 1-2 nuclei) will have a<br />
glycogen vacuole (vac) which will appear as a clear area in stained<br />
specimens. This vacuole will sometimes displace and alter the<br />
morphology of the nuclei. The chromatoid bodies tend to disappear<br />
as the cyst matures. The cysts are generally 12-15 µm in diameter.<br />
Cysts are immediately infective upon excretion with the feces and<br />
will be viable for weeks-to-months depending on environmental<br />
conditions.<br />
PATHOGENESIS<br />
non-invasive<br />
Amebiasis Progression<br />
• ameba colony on mucosa surface<br />
o asymptomatic cyst passer<br />
o non-dysenteric diarrhea<br />
invasive<br />
• necrosis of mucosa → ulcer<br />
o dysentery<br />
o hematophagous trophozoites<br />
• ulcer enlargement → peritonitis<br />
o occasional ameboma<br />
• metastasis → extraintestinal<br />
amebiasis<br />
o via blood-stream or direct
E. histolytica frequently lives as<br />
a commensal within the large<br />
intestine with no overt clinical<br />
manifestations. However,<br />
trophozoites can invade the<br />
colonic epithelium and produce<br />
ulcers and dysentery (see Box).<br />
extension<br />
o primarily liver → amebic<br />
abscess<br />
o other sites infrequent<br />
o ameba-free stools common<br />
This invasive disease can become progressively worse and lead to a<br />
more serious disease. The amebas can also metastasize to other<br />
organs and produce anextraintestinal amebiasis. In other words, E.<br />
histolytica is a facultative pathogen that exhibits a wide range of<br />
virulence.<br />
The non-invasive disease is often asymptomatic, but can cause<br />
diarrhea or other gastro-intestinal symptoms such as abdominal<br />
pain or cramps. This non-invasive infection can persist or progress<br />
to an invasive disease in which trophozoites penetrate the intestinal<br />
mucosa and kill the epithelial cells. The early lesion is a small area<br />
of necrosis, or ulcer, characterized by raised edges and virtually no<br />
inflammation between lesions (Figure). The ameba will spread<br />
laterally and downward in the submucosa (beneath the epithelium)<br />
and kill host cells as they progress. This results in the classic 'flaskshaped'<br />
ulcer with a small opening and a wide base. Trophozoites<br />
are most numerous at the boundary between the healthy tissue and<br />
the necrotic tissue. These invasive ameba are ingesting host cells<br />
and trophozoites with ingested erythrocytes are often evident.<br />
These hematophagous trophozoites are sometimes found in the<br />
dysenteric feces. Cyst production decreases during the invasive<br />
stage of the infection and cysts are never found in the tissue<br />
lesions.<br />
Left: The lumenal side of the colon from fulminating amebiasis case showing several ulcers. Note<br />
raised edges (arrow). Middle: Histological preparation showing cross-section of ulcer. Note the<br />
high degree of necrosis in center of ulcer. The amebas are advancing laterally under the intact<br />
mucosa as indicated by the microvilli. Right: Higher magnification of ulcer showing several<br />
hematophagous trophozoites. The nucleus (arrow) is evident in one of the amebas. Pictures from
Peters and Gilles (1989), A Colour Atlas of Tropical Medicine and Parasitology (3rd edition).<br />
E. histolytica is found primarily in<br />
the colon where it can live as a<br />
non-pathogenic commensal or<br />
invade the intestinal mucosa<br />
(green). The ameba can<br />
metastasize to other organs via a<br />
hematogenous route (purple);<br />
primarily involving the portal vein<br />
and liver. The ameba can also<br />
spread via a direct expansion<br />
(blue) causing a pulmonary<br />
infection, cutaneous lesions or<br />
perianal ulcers.<br />
The ulcerative process may continue<br />
to expand laterally or downward. If<br />
large numbers of ulcers are present,<br />
they may coalesce which could lead<br />
to a localized sloughing off of the<br />
intestinal wall. Ulcer expansion can<br />
also penetrate the serous layer and<br />
lead to perforation of the intestinal<br />
wall. This perforation can lead to<br />
local abscesses or a generalized<br />
peritonitis. (See also schematic<br />
representation of tissue invasion.)<br />
Amebic ulcers can also become<br />
secondarily infected with bacteria<br />
which may confuse the clinical<br />
picture. In addition, E. histolytica<br />
infection can occasionally lead to the<br />
formation of an amebic granuloma,<br />
also called an ameboma. The<br />
ameboma is an inflammatory<br />
thickening of the intestinal wall<br />
around the ulcer which can be<br />
confused with a tumor.<br />
Amebiasis can also progress to a<br />
systemic, or extraintestinal infection.<br />
Dissemination from the primary<br />
intestinal lesion is predominantly via<br />
the blood stream, but can also occur<br />
by direct extension of the lesion. The<br />
liver is the most commonly affected<br />
organ and this is probably due to the<br />
direct transport of trophozoites from<br />
the large intestine to the liver via the<br />
hepatic portal vein (Figure). Initially<br />
the lesions are small foci of necrosis<br />
which tend to coalesce into a single<br />
abscess as they expand. This hepatic<br />
abscess will continue to enlarge as the trophozoites progressively<br />
destroy and ingest host cells. The center of the abscess, consisting<br />
of lysed hepatocytes, erythrocytes, bile and fat, may liquefy and<br />
this necrotic material (sometimes incorrectly called pus) will range<br />
in color from yellowish to reddish brown. Secondary bacterial<br />
infections in the liver abscess are not common (~2%).
Hematogenous spread of trophozoites to other sites, such as the<br />
lungs or brain, is rare, but does occur. The second most common<br />
extraintestinal site after the liver is the lungs. Pulmonary infections<br />
generally result from a direct extension of the hepatic lesion across<br />
the diaphragm and into the pleura and lungs. Cutaneous lesions<br />
formed as a result of hepatic or intestinal fistula can also occur,<br />
although extremely rare. Other cutaneous lesions include perianal<br />
ulcers and involvement of the genitalia, including the penis of<br />
homosexuals. These later manifestations are likely due to the skin<br />
or mucous membranes coming in contact with invasive<br />
trophozoites.<br />
POSSIBLE MECHANISMS OF<br />
PATHOGENESIS<br />
As discussed above, E. histolytica is<br />
pathogen that exhibits a wide<br />
spectrum of virulence, ranging from<br />
an avirulent commensal to a highly<br />
invasive and destructive organism<br />
(see discussion of pathogenicity vs.<br />
virulence). Some of this difference in<br />
virulence is explained by the<br />
existence of the morphologically<br />
identical, but avirulent, E. dispar. E.<br />
dispar has never been associated with<br />
a symptomatic invasive disease and<br />
infection does not elicit serum<br />
Entamoeba Prevalences<br />
• E. dispar ~10-fold > E.<br />
histolytica<br />
• discrete endemic pockets of<br />
E. histolytica observed<br />
• ~25% seropositive for E.<br />
histolytica in endemic areas<br />
• ~10% infected with E.<br />
histolytica will develop<br />
invasive amebiasis<br />
antibodies. In contrast, anti-ameba humoral responses are<br />
observed in both asymptomatic and symptomatic E. histolytica<br />
infections. This suggests that even in asymptomatic cases there is a<br />
limited amount of invasion. However, infection with E. histolytica<br />
does not always lead to invasive disease, though, in that only about<br />
10% of the infected individuals will develop symptomatic invasive<br />
amebiasis. The factors responsible for the pathogenesis of E.<br />
histolytica are not well understood. One approach to understanding<br />
the pathogenesis is to compare possible virulence factors between<br />
these two closely related species.<br />
Possible Virulence Factors<br />
host factors<br />
• ineffective innate immunity<br />
• inflammatory response
parasite factors<br />
• resistance to host response<br />
(eg, complement resistance)<br />
• adherence properties (eg,<br />
'Eh-lectin')<br />
• cytolytic properties (eg,<br />
adherence + 'amebapore')<br />
• ability to breakdown tissues<br />
(eg, secreted proteases)<br />
Pathology results from host-parasite<br />
interactions, and therefore, host<br />
factors, parasite factors or a<br />
combination of both may contribute<br />
to the disease state. For example, the<br />
development of invasive disease<br />
could be due to quantitative or<br />
qualitative aspects of the host<br />
immune response. Recruitment of<br />
neutrophils and intense inflammation<br />
are noted in the early phases of<br />
amebic invasion. However,<br />
inflammation surrounding established ulcers and abscesses if often<br />
minimal given the degree of tissue damage.<br />
The nature of protective immune responses is not clear. Innate or<br />
nonspecific immunity, as well as acquired immunity, are probably<br />
both important for the prevention of invasive disease. The mucous<br />
layer covering the epitheilial cells can prevent contact between<br />
trophozoite and host cells. In addition, mucosal IgA responses do<br />
occur as a result of infection and fecal IgA against a trophozoite<br />
surface lectin (see Eh-lectin) are associated with a lower incidence<br />
of new E. histolytica infections. High titers of serum antibodies also<br />
develop in patients with liver abscesses. However, since the<br />
invasive disease is often progressive and unremitting, the role of<br />
these anti-ameba antibodies is in question. Cell-mediated responses<br />
appear to play a role in limiting the extent of invasive amebiasis<br />
and protecting the host from recurrence following successful<br />
treatment.<br />
Resistance to the host immune response is another possible<br />
virulence factor which could contribute to the development and<br />
exacerbation of invasive disease. For example, one phenotypic<br />
difference between E. dispar and E. histolytica is the resistance of<br />
the latter to complement mediated lysis (see E. dispar). In addition,<br />
E. histolytica rapidly degrades secretory IgA and possibly<br />
suppresses T-cell responses to E. histolytica antigens. E. histolytica<br />
is also able to kill cells, including neutrophils and other immune<br />
effector cells, in a contact dependent manner. Lysis of neutrophils<br />
could also release toxic products which contribute to the destruction<br />
of host tissue. However, the role of these various phenomena in<br />
pathogenesis is not known.<br />
Invasion of intestinal mucosa by E. histolytica is an active process<br />
mediated by the parasite and distinct steps can be recognized<br />
(Figure, click here for larger image and detailed legend).<br />
Trophozoites adhere to the mucus layer (step 1). This adherence
per se probably does not contribute to pathogenesis and is simply a<br />
mechanism for the ameba to crawl along the substratum. Depletion<br />
of the mucus barrier allows for the trophozoite to come in contact<br />
with epithelial cells. Epithelial cells are killed in a contact dependent<br />
manner leading to a disruption of the intestinal mucosa (step 2).<br />
The trophozoites will continue to kill host cells in the submucosa<br />
and further disrupt the tissue as they advance (step 3). Disruption<br />
of the intestinal wall (step 4) or metastasis via the circulatory<br />
system (step 5) is also possible. Adherence, cytotoxicity, and<br />
disruption of the tissues are important factors in the pathogenesis<br />
of E. histolytica. Parasite proteins which could play a role in these<br />
processes include: the Eh-lectin, amebapore, and proteases.<br />
(Skip detailed discussions of Eh-lectin, amebapore, and proteases and go to<br />
clinical symptoms.)<br />
Eh-lectin. E. histolytica can kill cells within minutes of adhering to<br />
them in the presence of extracellular calcium. Adherence of E.<br />
histolytica trophozoites to host cells and colonic mucins is mediated<br />
by a lectin-activity expressed on the ameba's surface. This lectin<br />
binds galactose or N-acetyl-D-galactosamine (GalNAc) with a high<br />
affinity and is also called the galactose-inhibitable adherence<br />
protein (GIAP) or the Gal/GalNAc lectin. The contact-dependent<br />
killing of target cells is almost completely inhibited by galactose or<br />
GalNAc and target cells lacking terminal galactose residues on their<br />
surface glycoproteins are resistant to trophozoite adherence and<br />
cytotoxicity. This suggests that the Gal/GalNAc lectin is an<br />
important virulence factor. In addition, the Eh-lectin is involved in<br />
resistance to complement mediated lysis. Because of its potential<br />
role in adherence and virulence and since fecal IgA against it<br />
protect against amebic colitis, the Gal/GalNAc is a vaccine candidate<br />
(Petri et al, 2006, Arch. Med. Res. 37:288).
The Eh-lectin is a heterodimer consisting of a 170 kDa heavy chain<br />
and a 31-35 kDa light chain joined by disulfide bonds. An<br />
intermediate subunit of 150 kDa is noncovalently associated with<br />
the heterodimer. The heavy chain has a transmembrane domain<br />
and a carbohydrate binding domain. All of subunits are encoded by<br />
multigene families. There are five members of the heavy chain<br />
family, 6-7 members of the light chain family and 30 members of<br />
the intermediate chain family. The members of the heavy chain<br />
gene family exhibit 89-95% sequence identity at the amino acid<br />
level whereas the light chain family members are less conserved<br />
sharing only 79-85% sequence identity.<br />
E. dispar also expresses Gal/GalNAc lectin on its surface. Both E.<br />
dispar and E. histolytica need to adhere to the mucous layer which<br />
is medicated by the Gal/GalNAc lectin. Mucus is composed of<br />
glycoproteins called mucins. The predominant mucin found on the<br />
intestinal mucosa is Muc2 which is extensively glycosylated with Olinked<br />
GalNAc residues. The sequence of the light and heavy chain<br />
genes from E. dispar are homologous, but not identical, to those of<br />
E. histolytica. Antigenic differences between the GIAP of E. dispar<br />
and E. histolytica have also been described in that only two epitopes<br />
out of six are shared between the two species (see E. dispar). It is<br />
not known whether these sequence differences can account for the<br />
differences in virulence between E. dispar and E. histolytica.<br />
Adherence is obviously important for both species, but it is possible<br />
that the adherence is qualitatively or quantitatively different<br />
between the two species.<br />
[Review on the Eh-lectin: Petri et al (2002) Annu. Rev. Microbiol.<br />
56:39.]<br />
Amebapore. A family of pore-forming polypeptides has been<br />
identified in E. histolytica and E. dispar. The three family members<br />
are designated as amebapore A, B and C with amebapore A being<br />
predominant expressed. The mature polypeptide is 77 amino acids<br />
long and forms dimers at low pH (4-6). Three of these dimers then<br />
assemble into a hollow ring-shaped structure. This hexamer then<br />
can intercalate into membranes and introduce 2 nm pores (i.e.,<br />
holes) which results in cell death. The pore-forming activity is<br />
dependent on this assembly process beginning with the<br />
dimerization. Amebabpore A is 95% identical (i.e., four residues are<br />
different) between E. histolytica and E. dispar. In addition, the E.<br />
dispar amebapore has approximately half of the pore-forming<br />
activity as the E. histolytica amebapore. This difference in poreforming<br />
activity has been attributed to a glutamate residue at<br />
position 2 in the E. histolytica amebapore, as compared to a proline<br />
residue in the E. dispar amebapore. This particular amino acid
esidue is important for the formation of the dimers and it is<br />
believed that the dimers of E. dispar amebapore are less stable.<br />
Amebapore is localized to vacuolar compartments (eg, food<br />
vacuoles) within the trophozoite and is most active at acidic pH<br />
suggesting that the major function of amebapore is to lyse ingested<br />
bacteria. Nonetheless, amebapore is implicated as a virulence factor<br />
in that genetic manipulation of E. histolytica resulting in decreased<br />
expression of amebapore leads to a reduction in pathogenicity<br />
(ability to form liver abscesses) as well as a reduction in<br />
bacteriocidal activity (Bracha et al Mol. Microbiol. 34:363, 1999).<br />
Similarly, modified E. histolytica completely devoid of amebapore<br />
production are unable to form liver abscesses in model systems<br />
(Zhang et al, Inf. Imm. 72:678, 2004). However, these amebas are<br />
able to cause inflammation and tissue damage in models for amebic<br />
colitis.<br />
[Review on amebapore: Leippe et al, Tr. Parasitol. 21:5, 2005.]<br />
Proteases. Proteases are enzymes that degrade other proteins and<br />
could contribute to the pathogenesis cause by E. histolytica. In this<br />
regard, E. histolytica expresses and secretes higher levels of<br />
cysteine proteases, a particular class of protease, than E. dispar.<br />
Cysteine proteases have been shown to disrupt the polymerization<br />
of MUC2, the major component of colonic mucus. This degraded<br />
mucus is less efficient at blocking adherence of trophozoites to<br />
epithelial cells. Destruction of the extracellular matrix (ECM) by<br />
proteases may also facilitate trophozoite invasion. Inhibitors of<br />
cysteine proteases can decrease liver abscess size in experimental<br />
models.<br />
Twenty different cysteine<br />
protease genes have been<br />
identified in E. histolytica.<br />
Orthologs of two of the E.<br />
histolytica cysteine protease<br />
genes are not found in E.<br />
dispar. One of these,<br />
designated CP5, is expressed at<br />
high levels on the trophozoite<br />
surface. Mutants expressing<br />
lower levels of CP5 had a<br />
reduced ability to generate liver<br />
abscesses in a hamster<br />
amebiasis model. However,<br />
these mutants also had a<br />
reduced growth rate and lower<br />
erythrophagocytic activity, thus<br />
Figure from Horstmann et al (1992) Trop. Med.<br />
Parasitol. 43, 213.<br />
Factor histolytica vs dispar<br />
Eh-lectin<br />
sequence and epitope<br />
differences<br />
amebapore Ed has less activity (Pro/Glu)<br />
proteases<br />
Eh has unique genes and<br />
expresses more activity<br />
Figure from Horstmann et al (1992) Trop. Med.<br />
Parasitol. 43, 213.
it is not clear whether CP5 directly participates in the invasiveness<br />
of E. histolytica. Furthermore inhibition of 90% of CP5 activity did<br />
not affect the ability of E. histolytica trophozoites to destroy cell<br />
monolayers in vitro. CP1, CP2, and CP5 are the most abundantly<br />
expressed cysteine proteases in E. histolytica, whereas CP3 is the<br />
most abundant in E. dispar. Interestingly, over expression of CP2 in<br />
E. dispar increased the ability of trophozoites to destroy cell<br />
monolayers in vitro. However, the over expression of CP2 did not<br />
lead to the ability of E. dispar to form liver abscesses in gerbils.<br />
Therefore, it is not clear the precise roles proteases may play in<br />
pathogenesis.<br />
In summary, the pathogenesis associated with E. histolytica<br />
infection is primarily due to its ability to invade tissues and kill host<br />
cells. Several potential virulence factors have been identified (see<br />
Table). However, it is not clear the exact role these various<br />
virulence factors play in the development of invasive disease. One<br />
approach to understanding the pathogenesis is to compare these<br />
factors from E. histolytica and E. dispar. These two species are<br />
closely related and the potential virulence factors are found in both<br />
species. Adherence, cytolytic activity and proteolytic activity are<br />
inherent biological features of both species and these activities do<br />
not necessarily lead to pathology. However, there are qualitative<br />
and quantitative differences between E. histolytica and E. dispar<br />
which may account for the differences in virulence. These genetic<br />
differences between E. histolytica and E. dispar indicate that<br />
pathogenesis is in part an inherent feature of the parasite.<br />
However, pathogenesis is probably due to the combined effects of<br />
several host and parasite factors, and the virulence may represent<br />
the degree to which the host can control trophozoite invasion and<br />
replication.<br />
[See Huston, 2004, Tr. Parasitol. 20:23 for review of<br />
pathogenesis.]<br />
CLINICAL PRESENTATION<br />
Amebiasis presents a wide range of clinical syndromes (Table)<br />
which reflects the potential for E. histolytica to become invasive and<br />
cause a progressive disease. The incubation period can range from<br />
a few days to months or years with 2-4 weeks being the most<br />
common. Transitions from one type of intestinal syndrome to<br />
another can occur and intestinal infections can give rise to<br />
extraintestinal infections.<br />
Clinical Syndromes
Clinical Syndromes<br />
Associated with Amebiasis<br />
Intestinal Disease<br />
• asymptomatic cyst passer<br />
• symptomatic nondysenteric<br />
infection<br />
• amebic dysentery (acute)<br />
• fulminant colitis<br />
o + perforation<br />
(peritonitis)<br />
• ameboma (amebic<br />
granuloma)<br />
• perianal ulceration<br />
Extraintestinal Disease<br />
• liver abscess<br />
• pleuropulmonary amebiasis<br />
• brain and other organs<br />
• cutaneous and genital<br />
diseases<br />
The majority of individuals diagnosed<br />
with E. histolytica (or E. dispar)<br />
exhibit no symptoms or have vague<br />
and nonspecific abdominal<br />
symptoms. This state can persist or<br />
progress to a symptomatic infection.<br />
Symptomatic nondysenteric<br />
infections exhibit variable symptoms<br />
ranging from mild and transient to<br />
intense and long lasting. Typical<br />
symptoms include: diarrhea, cramps,<br />
flatulence, nausea, and anorexia. The<br />
diarrhea frequently alternates with<br />
periods of constipation or soft stools.<br />
Stools sometimes contain mucus, but<br />
there is no visible blood.<br />
Amebic dysentery usually starts<br />
slowly over several days with<br />
abdominal cramps, tenesmus, and<br />
occassional loose stools, but<br />
progresses to diarrhea with blood<br />
and mucus. Blood, mucus and pieces of necrotic tissue become<br />
more evident as the number of stools increases (10-20 or more per<br />
day) and stools will often contain little fecal material. A few patients<br />
may develop fever, vomiting, abdominal tenderness, or dehydration<br />
(especially children) as the severity of the disease increases.<br />
Fulminant, or grangrenous, colitus is a rare but extremely severe<br />
form of intestinal amebiasis. Patients present with severe bloody<br />
diarrhea, fever, and diffuse abdominal tenderness. Most of the<br />
mucosa is involved and mortality exceeds 50%. A chronic<br />
amebiasis, characterized by recurrent attacks of dysentery with<br />
intervening periods of mild or moderate gastrointestinal symptoms,<br />
can also occur.<br />
Amebomas present as painful abdominal masses which occur most<br />
frequently in the cecum and ascending colon. Obstructive symptoms<br />
or hemorrhages may also be associated with an ameboma.<br />
Amebomas are infrequent and can be confused with carcinomas or<br />
tumors. Perianal ulcers are a form of cutaneous amebiasis that<br />
result from the direct spread of the intestinal infection.<br />
Amebic liver abscesses are the most common form of extraintestinal<br />
amebiasis. The onset of hepatic symptoms can be rapid or gradual.<br />
Hepatic infections are characterized by hepatomegaly, liver<br />
tenderness, pain in the upper right quadrant, fever and anorexia.<br />
Fever sometimes occurs on a daily basis in the afternoon or
evening. Liver function tests are usually normal or slightly abnormal<br />
and jaundice is unusual. Liver abscesses will occasionally rupture<br />
into the peritoneum resulting in peritonitis.<br />
Pulmonary amebiasis is generally results from the direct extension<br />
of the liver abscess through the diaphragm. Clinical symptoms most<br />
often include cough, chest pain, dyspnea (difficult breathing), and<br />
fever. The sputum may be purulent or blood-stained and contain<br />
trophozoites. A profuse expectoration (ie, vomica) of purulent<br />
material can also occur. Primary metastasis to the lungs is rare, but<br />
does occur. Similarly, infection of other organs (eg., brain, spleen,<br />
pericardium) is also rare. Clinical symptoms are related to the<br />
affected organ.<br />
Cutaneous amebiasis is the result of skin or mucus membranes<br />
being bathed in fluids containing trophozoites. This contact can be<br />
the result of fistula (intestinal, hepatic, perineal) or an invasion of<br />
the genitalia. Cutaneous lesions have a wet, granular, necrotic<br />
surface with prominent borders and can be highly destructive.<br />
Clinical diagnosis is difficult and is usually considered with<br />
epidemiological risk factors (eg., endemic areas, male<br />
homosexuality, etc.).
DIAGNOSIS, TREATMENT AND CONTROL<br />
Intestinal Disease<br />
Diagnosis<br />
• stool examination<br />
o cysts and/or<br />
trophozoites<br />
• sigmoidoscopy<br />
o lesions, aspirate,<br />
biopsy<br />
• antigen detection<br />
o histolytica/dispar<br />
Extraintestinal (hepatic) Disease<br />
• serology<br />
o current or past?<br />
• imaging<br />
o CT, MRI, ultrasound<br />
• abscess aspiration<br />
o only select cases<br />
o reddish brown liquid<br />
o trophozoites at<br />
abscess wall<br />
Definitive diagnosis of amebiasis<br />
requires the demonstration of E.<br />
histolytica cysts or trophozoites in<br />
feces or tissues. Stool specimens<br />
should be preserved and stained and<br />
microscopically examined. Cysts will<br />
tend to predominate in formed stools<br />
and trophozoites in diarrheic stools<br />
(see morphology). Fresh stools can<br />
also be immediately examined for<br />
motile trophozoites which exhibit a<br />
progressive motility. Sigmoidoscopy<br />
may reveal the characteristic ulcers,<br />
especially in more severe disease.<br />
Aspirates or biopsies should also be<br />
examined microscopically for<br />
trophozoites.<br />
E. histolytica and E. dispar cannot be<br />
distinguished on morphological<br />
criteria. Antigen detection kits are<br />
available for the positive<br />
identification of these species.<br />
Serology is especially useful for the diagnosis of extraintestinal<br />
amebiasis. Greater than 90% of patients with invasive colitis and<br />
liver abscesses exhibit serum antibodies against E. histolytica.<br />
However, the antibodies can persist for years and distinguishing<br />
past and current infections may pose problems in endemic areas.<br />
Non-invasive imaging techniques (eg., ultrasound, CT, MRI) can be<br />
used to detect hepatic abscesses. It is also possible to aspirate<br />
hepatic abscesses. However, this is rarely done and only indicated<br />
in selected cases (eg., serology and imaging not available,<br />
therapeutic purposes). The aspirate is usually a thick reddish brown<br />
liquid that rarely contains trophozoites. Trophozoites are most likely<br />
to be found at the abscess wall and not in the necrotic debris at the<br />
abscess center.<br />
Several drugs are available for the treatment of amebiasis and the<br />
choice of drug(s) depends on the clinical stage of the infection<br />
(Table). The prognosis following treatment is generally good in<br />
uncomplicated cases. In cases where E. histolytica is confirmed or<br />
the species (ie, dispar or histolytica) is unknown, asymptomatic cyst<br />
passers should be treated to prevent the progression to severe<br />
disease and to control the spread of the disease. However, in many<br />
endemic areas, where the rates of reinfection are high and
treatment is expensive, the standard practice is to only treat<br />
symptomatic cases. Metronidazole or tinidazole (if available) is<br />
recommended for all symptomatic infections. This treatment should<br />
be followed by or combined with lumenal antiamebic drugs as<br />
described for asymptomatic patients.<br />
Drugs Uses<br />
Iodoquinol, Paromomycin, or<br />
Diloxanide furoate<br />
Metronidazole or Tinidazole<br />
Dehydroemetine or Emetine<br />
Amebiasis Treatment<br />
Luminal agents to treat asymptomatic cases and as a follow up<br />
treatment after a nitroimidazole.<br />
Treatment of nondysenteric colitis, dysentery, and extraintestinal<br />
infections.<br />
Treatment of severe disease such as necrotic colitis, perforation<br />
of intestinal wall, rupture of liver abscess.<br />
In the cases of fulminant amemic colitis or perforation of the<br />
intestinal wall a broad spectrum antibiotic can also be used to treat<br />
intestinal bacteria in the peritoneum. Necrotic colitis requires urgent<br />
hospitalization to restore fluid and electrolyte balance. In addition,<br />
emetine or dehydroemetine are sometimes co-administered with<br />
the nitroimidazole. This is only done in the most severe cases due<br />
to the toxicity of these drugs. Surgery may also be needed to close<br />
perforations or a partial colostomy. Abscess drainage of hepatic<br />
lesions (ie, needle aspiration or surgical drainage) is now rarely<br />
performed for therapeutic purposes and is only indicated in cases of<br />
large abscesses with a high probability of rupture.<br />
Prevention and control measures are similar to other diseases<br />
transmitted by the fecal-oral route (see Risk Factors or discussion of<br />
Giardia control). The major difference is that humans are the only<br />
host for E. histolytica and there is no possibility of zoonotic<br />
transmission. Control is based on avoiding the contamination of<br />
food or water with fecal material. Health education in regards to<br />
improving personal hygiene, sanitary disposal of feces, and hand<br />
washing are particularly effective. Protecting water supplies will<br />
lower endemicity and epidemics. Like Giardia, Entamoeba cysts are<br />
resistant to standard chlorine treatment, but are killed by iodine or<br />
boiling. Sedimentation and filtration processes are quite effective at<br />
removing Entamoeba cysts. Chemoprophylaxis is not<br />
recommended.<br />
Recent review on amebiasis:<br />
• Haque, R. et al (2003) Amebiasis. N. Engl. J. Med. 348:1565.<br />
• Stanley, S.L. (2003) Amoebiasis. The Lancet 361:1025.
NON-PATHOGENIC COMMENSALS<br />
Numerous protozoa can inhabit the gastro-intestinal tract of<br />
humans. Most of these exhibit little or no overt pathology. Infection<br />
with these protozoa is evidence of fecal contamination and indicates<br />
a risk for more serious infections such as Giardia or E. histolytica.<br />
These non-pathogenic species can also be confused with the<br />
potentially pathogenic Giardia or E. histolytica and result in<br />
unnecessary drug treatment. In addition, such a misdiagnosis is<br />
also problematic in that the true cause of the symptoms may be<br />
missed and the appropriate treatment will be<br />
delayed.<br />
• Other Entamoeba<br />
• Other Intestinal Amebae<br />
• Other Intestinal Flagellates<br />
• Blastocystis<br />
Entamoeba Species Infecting<br />
Humans<br />
Several Entamoeba species infect humans (box). E. histolytica can<br />
cause a severe intestinal disease characterized by dysentery as well<br />
as an invasive disease affecting primarily the liver (see Amebiasis).<br />
E. dispar is morphologically identical to E. histolytica, but does not<br />
produce an invasive disease (see further discussion on E. dispar). A<br />
distinguishing feature of the Entamoeba is their nuclear morphology<br />
which is described as having peripheral chromatin and a small<br />
karyosome. E. histolytica/dispar, E.coli, and E. hartmanni can be<br />
distinguished by size and minor morphological differences (see<br />
Table).<br />
Intestinal Entamoeba Species<br />
E. dispar* E. coli E. hartmanni<br />
Trophozoites Trophozoites Trophozoites<br />
• 15-20 µm**<br />
• extend pseudopodia<br />
• progressive movement<br />
• 20-25 µm<br />
• broad blunt pseudopodia<br />
• sluggish, non-directional<br />
movement<br />
Cysts Cysts Cysts<br />
• 12-15 µm<br />
• 4 nuclei<br />
• blunt chromatoid bodies<br />
• 15-25 µm<br />
• 8 nuclei<br />
• pointed chromatoid<br />
bodies<br />
• E. histolytica<br />
• E. dispar<br />
• E. coli<br />
• E. hartmanni<br />
• E. polecki<br />
• E. gingivalis<br />
• 8-10 µm<br />
• less progressive than E.<br />
dispar<br />
• 6-8 µm<br />
• 4 nuclei<br />
• blunt chromatoid bodies<br />
• CB persist in mature<br />
cysts
• blunt chromatoid bodies • pointed chromatoid<br />
bodies<br />
*=E. histolytica; **invasive E. histolytica can be >20 mm<br />
• blunt chromatoid bodies<br />
• CB persist in mature<br />
cysts<br />
E. coli is the largest and is best distinguished by 8 nuclei in the<br />
mature cyst. The trophozoites of E. coli can be difficult to<br />
distinguish from E. histolytica/dispar since there is some overlap in<br />
the size ranges. E. hartmanni is quite similar to E. histolytica and<br />
was previously considered a 'small race' of E. histolytica. Generally<br />
10 µm is chosen as the boundary between E. histolytica and E.<br />
hartmanni.<br />
E. polecki is usually associated with pigs and monkeys, but human<br />
cases have been occasionally documented. It appears to be<br />
geographically restricted to particular areas such a Papua, New<br />
Guinea. The trophozoites are similar to E. coli, except a little<br />
smaller, and the cysts are similar to E. histolytica except that the<br />
mature cyst has a single nucleus.<br />
E. gingivalis can be recovered from the soft tartar between teeth<br />
and exhibits a similar morphology to E. histolytica except that it has<br />
no cyst stage. E. gingivalis can also multiply in bronchial mucus,<br />
and thus can appear in the sputum. In this case it could be<br />
confused with E. histolytica from a pulmonary abscess. E. gingivalis<br />
trophozoites will often contain ingested leukocytes which can be<br />
used to differentiate it from E. histolytica. The trophozoites are<br />
most often recovered from patients with periodontal disease, but an<br />
etiology between the organism and disease has not been<br />
established and E. gingivalis is considered to be non-pathogenic.<br />
Other Intestinal Amebae<br />
Other non-pathogenic amebae include Endolimax nana and<br />
Iodoamoeba bütschlii. Historically, Dientamoeba fragilis has been<br />
grouped with the ameba, but electron microscopy and molecular<br />
phylogenetics suggests that it is actually a flagellate and may be<br />
closely related to the trichomonads (see above). All three of these<br />
organisms exhibit similar morphologies and have nuclei which do<br />
not have peripheral chromatin and a large karyosome. Minor<br />
morphological differences allow these organims to be distinguished<br />
(Table).<br />
Other Intestinal Amebae
Endolimax nana Iodoamoeba bütschlii Dientamoeba fragilis*<br />
Trophozoites Trophozoites Trophozoites<br />
• 8-10 µm • 12-15 µm • 8-10 µm<br />
• often bi-nucleated<br />
• fragmented karyosome<br />
Cysts Cysts Cysts<br />
• 6-8 µm<br />
• 4 nuclei<br />
• 10-12 µm<br />
• 1 nuclei<br />
• glycogen vacuole<br />
*A flagellate possibly related to the trichomonads.<br />
Other Intestinal Flagellates<br />
• no cysts<br />
Four additional non-pathogenic flagellates recovered from human<br />
stools are: Trichomonas hominis, Chilomastix mesnili, Enteromonas<br />
hominis, and Retortamonas intestinalis. Among these T. hominis,<br />
also called Pentatrichomonas hominis, is the most common and is<br />
often recovered from diarrheic stools. These flagellates exhibit<br />
similar morphologies (Table) and can be difficult to distinguish. The<br />
trophozoites from all of these flagellates are somewhat teardrop<br />
shaped and contain a single nucleus and the cyst tend to be slightly<br />
elongated or oval.<br />
Other Intestinal Flagellates<br />
trophozoites cysts<br />
Size Flagella Size Nuclei<br />
Trichomonas hominis 6-14 µm 4 anterior, 1 posterior No cyst stage<br />
Chilomastix mesnili 10-15 µm 3 anterior, 1 in cytostome 7-9 µm 1<br />
Enteromonas hominis 6-8 µm 3 anterior, 1 posterior 4-8 µm 1-4<br />
Retortamonas intestinalis 4-10 µm 1 anterior, 1 posterior 4-7 µm 1<br />
Blastocystis hominis<br />
Blastocystis hominis is a common organism found in human stools.<br />
Since its initial description approximately 100 years ago, it has been<br />
variously classified as an ameba, a yeast, a sporozoan, and the cyst<br />
stage of a flagellate. Analysis of the small subunit rRNA sequence<br />
indicates that Blastocystis is most closely related to the
stramenopiles, a complex assemblage of unicellular and muticellular<br />
protists. Other stramenopiles include diatoms, brown algae, and<br />
water molds. Many of the characteristics of Blastocystis are<br />
unknown or controversial. The mode of transmission, mechanism of<br />
cell replication, and other features of the life cycle have not<br />
conclusively demonstrated.<br />
Similarly, the status of<br />
Blastocystis as a pathogen,<br />
commensal, or opportunistic<br />
organism is unknown.<br />
Blastocystis is polymorphic in<br />
that a variety of<br />
morphological forms are found<br />
in feces and in vitro culture.<br />
The most widely recognized<br />
form is spherical 10-15 µm in<br />
diameter with a large central<br />
vacuole (Figure). This large<br />
vacuole pushes the nuclei and<br />
other organelles to the<br />
periphery of the cell. The vacuole is sometimes filled with a granular<br />
material. Small resistant cyst-like forms have been identified from<br />
in vitro cultures and occasionally observed in feces. These<br />
presumed cysts are approximately 5 µm and surround by a<br />
multilayered wall. Furthermore, the cysts do not lyse when placed in<br />
water suggesting that they are resistant to environmental<br />
conditions. Presumably Blastocystis is transmitted via a fecal-oral<br />
route. However, this has not been conclusively demonstrated.<br />
There have been several reports suggesting Blastocystis causes<br />
disease, as well as many reports suggesting the opposite. Diarrhea,<br />
cramps, nausea, vomiting and abdominal pain have been associated<br />
with large numbers of organisms in the stool. In addition, some<br />
studies have shown that treatment alleviates the symptoms and<br />
clears the organisms. However, the drugs used against Blastocystis<br />
(eg., metronidazole) also work against many other intestinal<br />
protozoa and bacteria. The inability to rule out other organisms as<br />
the source of symptoms and the observation that many infected<br />
persons exhibit no symptoms makes it difficult to draw any<br />
definitive conclusions about the pathogenesis of Blastocystis.<br />
Furthermore, it could be that Blastocystis is primarily a commensal,<br />
but can exhibit virulence under specific host conditions like<br />
concomitant infections, poor nutrition, or immunosuppression.<br />
Blastocystis is also found in a wide range of animals, including<br />
mammals, birds, reptiles, amphibians and even insects, and exhibits
a wide range of molecular diversity. The genetic distance between<br />
Blastocystis isolates is greater than the genetic distance between E.<br />
histolytica and E. dispar (see discussion on E. dispar). This<br />
complicates the designation of species and historically human<br />
isolates have been designated as B. hominis and isolates for other<br />
hosts as Blastocytis sp.. However, phylogenetic analysis reveals<br />
that there are no exclusively human clades and human isolates are<br />
found in all of the clades. This raises the possibility that Blastocystis<br />
is not host specific and can be transmitted zoonotically. In addition,<br />
the wide range of genetic diversity might explain the controversy<br />
concerning the pathogenecity of Blastocystis in that some<br />
genotypes may be more virulent than others. However, studies<br />
addressing this issue suggest that this is not the case. Resolution of<br />
the confusion about the taxonomy, transmission and virulence of<br />
Blastocystis will require additional studies.<br />
Recent reviews on Blastocystis:<br />
• Stenzel, D.J. and Boreham, P.F.L. (1996) Blastocystis hominis<br />
revisited. Clinical Microbiology Reviews 9, 563-584.<br />
• Tan, K.S.W. (2004) Blastocystis in humans and animals: new<br />
insights using modern methodologies. Veterinary Parasitology<br />
126, 121-144.<br />
• Yoshikawa, H. Morimoto, K., Wu, Z., Singh, M. and<br />
Hashimoto, T. (2004) Problems in speciation in the genus<br />
Blastocystis. Trends in Parasitology 20, 251-255.<br />
LINKS<br />
• Top<br />
• Contents<br />
• Giardiasis<br />
• Trichomoniasis<br />
• Balantidosis<br />
• Amebiasis<br />
• Non-pathogenic Commensals<br />
• Protozoology Home<br />
o Study Guides<br />
o Syllabus<br />
• Other Courses and Lectures<br />
• Wiser Home<br />
• Other Internet Sites<br />
o Parasites, Division of Parasitic Diseases, CDC<br />
o The Medical Letter (treatment recommendations)<br />
http://en.wikipedia.org/wiki/Category:Laboratory_techniques
Category:Laboratory techniques<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
Laboratory techniques, as used in Biology, Biochemistry, Chemistry, Molecular<br />
biology, etc.<br />
Subcategories<br />
This category has the following 8 subcategories, out of 8 total.<br />
B<br />
C<br />
• [+] Biochemistry<br />
methods<br />
• [+]<br />
Chromatography<br />
D<br />
E<br />
M<br />
• [+] Distillation<br />
• [+] Electrophoresis<br />
• [+] Microbiology<br />
techniques<br />
M cont.<br />
P<br />
• [+] Microscopy<br />
Pages in category "Laboratory techniques"<br />
The following 132 pages are in this category, out of 132 total.<br />
A<br />
B<br />
• Acid-base extraction<br />
• Air-free technique<br />
• Allele specific<br />
oligonucleotide<br />
• Ames test<br />
• Ammonium sulfate<br />
precipitation<br />
• Animal testing<br />
• Assay<br />
• Baby Gender Mentor<br />
F cont.<br />
G<br />
• Fractionation<br />
• Gas chromatographymass<br />
spectrometry<br />
• Gas-liquid<br />
chromatography<br />
• Gel electrophoresis<br />
• Gel extraction<br />
• Gene gun<br />
• [+] Polymerase chain<br />
reaction<br />
• [+] Protein-protein<br />
interaction assays<br />
P cont.<br />
• Plant tissue<br />
culture<br />
• Polymerase<br />
chain reaction<br />
• Post harvest<br />
freshness<br />
• Protein<br />
Misfolding<br />
Cyclic<br />
Amplification<br />
• Protein<br />
electrophoresis<br />
• Protein tag<br />
• Protein-
C<br />
D<br />
E<br />
• Blot (biology)<br />
• Borax<br />
• Cell disruption by<br />
nitrogen<br />
decompression<br />
• Cell fractionation<br />
• Centrifugation<br />
• Chemotaxis assay<br />
• Chromosome jumping<br />
• Cooling bath<br />
• Cosmid<br />
• Cot analysis<br />
• Cot filtration<br />
• Crystallization<br />
• Cycling probe<br />
technology<br />
• DNA extraction<br />
• DNA footprinting<br />
• DNA laddering<br />
• DNase footprinting<br />
assay<br />
• Degasification<br />
• Diethylpyrocarbonate<br />
• Differential<br />
centrifugation<br />
• Digital polymerase<br />
chain reaction<br />
• Direct fluorescent<br />
antibody<br />
• Distillation<br />
• Dithioerythritol<br />
• Electrochromatography<br />
• Electropherogram<br />
• Electrophoresis<br />
(disambiguation)<br />
• Electrophoretic<br />
mobility shift assay<br />
• Electrophoretogram<br />
• Ellman's reagent<br />
H<br />
I<br />
L<br />
M<br />
• Hofmeister series<br />
• Homogenization<br />
• Host-Cell Reactivation<br />
• Hydrophilic<br />
interaction liquid<br />
chromatography<br />
• Immunohistochemistry<br />
• Immunomagnetic<br />
separation<br />
• Immunoperoxidase<br />
• In situ hybridization<br />
• Inductively coupled<br />
plasma atomic<br />
emission spectroscopy<br />
• Inductively coupled<br />
plasma mass<br />
spectrometry<br />
• Inverse polymerase<br />
chain reaction<br />
• Isotopic dilution<br />
• Isotopic labeling<br />
• LIESST<br />
• Laboratory automation<br />
• Laboratory centrifuge<br />
• Lamm equation<br />
• Lipofection<br />
• Liquid gas<br />
• Liquid-liquid<br />
extraction<br />
• List of purification<br />
methods in chemistry<br />
• Low copy number<br />
• Lowry protein assay<br />
• Lysis buffer<br />
• Magnetofection<br />
• Mason-Weaver<br />
equation<br />
• Microscopy<br />
Q<br />
R<br />
S<br />
fragment<br />
Complementatio<br />
n Assay<br />
• Quantitative<br />
polymerase<br />
chain reaction<br />
• Real-time<br />
polymerase<br />
chain reaction<br />
• Recrystallization<br />
• Restriction<br />
landmark<br />
genomic<br />
scanning<br />
• Reverse<br />
transcription<br />
polymerase<br />
chain reaction<br />
• SDS-PAGE<br />
• Salting out<br />
• Sedimentation<br />
• Sequencing by<br />
hybridization<br />
• Serial dilution<br />
• Sham operated<br />
group<br />
• Size exclusion<br />
chromatography<br />
• Solid phase<br />
microextraction<br />
• Sonication<br />
• Southern blot<br />
• Southwestern<br />
blot<br />
• Sparging<br />
(chemistry)<br />
• Standard<br />
addition<br />
• Starch indicator<br />
• Sublimation
F<br />
• Ethanol precipitation<br />
• FLAG-tag<br />
• Filtration<br />
• Finisher<br />
• Fluorescent in situ<br />
hybridization<br />
• Fosmid<br />
N<br />
O<br />
P<br />
• Murashige and Skoog<br />
medium<br />
• Nanopore sequencing<br />
• Nanovid microscopy<br />
• Native PAGE<br />
• Nested polymerase<br />
chain reaction<br />
• Nick translation<br />
• Northern blot<br />
• Nucleic acid<br />
hybridization<br />
• Oligonucleotide<br />
synthesis<br />
• Organ culture<br />
• Overlay assay<br />
• Peptide mass<br />
fingerprinting<br />
T<br />
V<br />
W<br />
Z<br />
(chemistry)<br />
• Sublimation<br />
apparatus<br />
• Sucrose gradient<br />
centrifugation<br />
• Suction<br />
filtration<br />
• SuperSAGE<br />
• TMB Liquid<br />
substrate for<br />
ELISA<br />
• TUNEL assay<br />
• Terminal<br />
restriction<br />
fragment length<br />
polymorphism<br />
• Tissue culture<br />
• Touchdown<br />
polymerase<br />
chain reaction<br />
• Twodimensional<br />
gel<br />
electrophoresis<br />
• Visualized<br />
Experimental<br />
Biology<br />
• Western blot<br />
• Wet laboratory<br />
• Winogradsky<br />
column<br />
• Zoo blot<br />
• Zymography<br />
Retrieved from "http://en.wikipedia.org/wiki/Category:Laboratory_techniques"<br />
Categories: Chemistry | Laboratories | Biological techniques and tools<br />
Views
• This page was last modified on 26 December 2007, at 17:35.<br />
• All text is available under the terms of the GNU Free Documentation License.<br />
(See Copyrights for details.)<br />
Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc., a<br />
U.S. registered 501(c)(3) tax-deductible nonprofit charity.<br />
• Privacy policy<br />
• About Wikipedia<br />
• Disclaimers<br />
http://www.who.int/mediacentre/factsheets/fs094/en/<br />
WHO > Programmes and projects > Media centre > Fact sheets<br />
Main content<br />
printable version<br />
Fact sheet N°94<br />
May 2007<br />
Malaria<br />
- Malaria is both preventable and curable.<br />
- A child dies of malaria every 30 seconds.<br />
- More than one million people die of malaria every year, mostly infants, young<br />
children and pregnant women and most of them in Africa.<br />
INFECTION AND TRANSMISSION<br />
Malaria is a disease which can be transmitted to people of all ages. It is caused by<br />
parasites of the species Plasmodium that are spread from person to person through the<br />
bites of infected mosquitoes. The common first symptoms – fever, headache, chills, and<br />
vomiting – appear 10 to 15 days after a person is infected. If not treated promptly with<br />
effective medicines, malaria can cause severe illness that is often fatal.<br />
There are four types of human malaria – Plasmodium falciparum, P.vivax, P.malariae, and<br />
P.ovale. P.falciparum and P.vivax are the most common. P.falciparum is by far the most<br />
deadly type of malaria infection.<br />
Malaria transmission differs in intensity and regularity depending on local factors such as<br />
rainfall patterns, proximity of mosquito breeding sites and mosquito species. Some regions<br />
have a fairly constant number of cases throughout the year – these are malaria endemic –<br />
whereas in other areas there are “malaria” seasons, usually coinciding with the rainy<br />
season.
Malaria transmission differs in intensity and regularity depending on local factors such as<br />
rainfall patterns, proximity of mosquito breeding sites and mosquito species. Some regions<br />
have a fairly constant number of cases throughout the year – these are malaria endemic –<br />
whereas in other areas there are “malaria” seasons, usually coinciding with the rainy<br />
season.<br />
Large and devastating epidemics can occur in areas where people have had little contact<br />
with the malaria parasite, and therefore have little or no immunity. These epidemics can be<br />
triggered by weather conditions and further aggravated by complex emergencies or natural<br />
disasters.<br />
GLOBAL AND REGIONAL RISK<br />
Approximately, 40% of the world’s population, mostly<br />
those living in the world’s poorest countries, are at risk of<br />
malaria. Every year, more than 500 million people become<br />
severely ill with malaria. Most cases and deaths are in sub-<br />
Saharan Africa. However, Asia, Latin America, the Middle<br />
East and parts of Europe are also affected. Travellers from<br />
malaria-free regions going to areas where there is malaria<br />
transmission are highly vulnerable – they have little or no<br />
immunity and are often exposed to delayed or wrong<br />
malaria diagnosis when returning to their home country.<br />
TREATMENT<br />
Early diagnosis and prompt treatment are the basic<br />
elements of malaria control. Early and effective treatment of malaria disease will shorten<br />
its duration and prevent the development of complications and the great majority of deaths<br />
from malaria. Access to disease management should be seen not only as a component of<br />
malaria control but a fundamental right of all populations at risk. Malaria control must be<br />
an essential part of health care development. In contemporary control, treatment is<br />
provided to cure patients rather than to reduce parasite reservoirs.<br />
Antimalarial treatment policies will vary between countries depending on the<br />
epidemiology of the disease, transmission, patterns of drug resistance and political and<br />
economic contexts.<br />
DRUG RESISTANCE<br />
Related links<br />
:: Global Malaria<br />
Programme<br />
:: Roll Back Malaria<br />
Partnership<br />
:: Malaria (Special<br />
Programme for Research<br />
and Training in Tropical<br />
Diseases, TDR)<br />
The rapid spread of antimalarial drug resistance over the past few decades has required<br />
more intensive monitoring of drug resistance to ensure proper management of clinical<br />
cases and early detection of changing patterns of resistance so that national malaria<br />
treatment policies can be revised where necessary. Surveillance of therapeutic efficacy<br />
over time is an essential component of malaria control. Recent efforts to scale-up malaria<br />
control in endemic countries throughout the world including increased support for<br />
commodities and health systems, as well as the proposed price subsidy on artemisininbased<br />
combination therapies (ACTs) is resulting in greater access to and a vastly increased<br />
use of antimalarial medicines, in particular ACTs. This is leading to a much higher degree
of drug pressure on the parasite which will almost certainly increase the likelihood of<br />
selecting for resistant parasite genotypes. There are currently no effective alternatives to<br />
artemisinins for the treatment of P. falciparum malaria either on the market or towards the<br />
end of the development pipeline.<br />
The parasite's resistance to medicines continues to undermine malaria control efforts.<br />
WHO has therefore called for continuous monitoring of the efficacy of recently<br />
implemented ACTs, and countries are being assisted in strengthening their drug resistance<br />
surveillance systems. In order to preserve the efficacy of artemisinins as an essential<br />
component of life-saving ACTs, WHO has called for a ban on the use of oral artemisinin<br />
monotherapies, at various levels, including manufacturers, international drug suppliers,<br />
national health authorities and international aid and funding agencies involved in the<br />
funding of essential antimalarial medicines.<br />
PREVENTION: VECTOR CONTROL AND INTERMITTENT<br />
PREVENTIVE THERAPY IN PREGNANT WOMEN<br />
The main objective of malaria vector control is to significantly reduce both the number<br />
and rate of parasite infection and clinical malaria by controlling the malaria-bearing<br />
mosquito and thereby reducing and/or interrupting transmission. There are two main<br />
operational interventions for malaria vector control currently available: Indoor Residual<br />
Spraying of long-acting insecticide (IRS) and Long-Lasting Insecticidal Nets (LLINs).<br />
These core interventions can be locally complemented by other methods (e.g. larval<br />
control or environmental management) in the context of Integrated Vector Management<br />
(IVM). Effective and sustained implementation of malaria vector control interventions<br />
(IRS or LLINs) requires clear political commitment and engagement from national<br />
authorities as well as long-term support from funding partners.<br />
Pregnant women are at high risk of malaria. Non-immune pregnant women risk both acute<br />
and severe clinical disease, resulting in up to 60% fetal loss and over 10% maternal deaths,<br />
including 50% mortality for severe disease. Semi-immune pregnant women with malaria<br />
infection risk severe anaemia and impaired fetal growth, even if they show no signs of<br />
acute clinical disease. An estimated 10 000 of these women and 200 000 of their infants<br />
die annually as a result of malaria infection during pregnancy. HIV-infected pregnant<br />
women are at increased risk. WHO recommends that all endemic countries provide a<br />
package of interventions for prevention and management of malaria in pregnancy,<br />
consisting of (1) diagnosis and treatment for all episodes of clinical disease and anaemia<br />
and (2) insecticide-treated nets for night-time prevention of mosquito bites and infection.<br />
In highly endemic falciparum malaria areas, this should be complemented by (3)<br />
intermittent preventive treatment with sulfadoxine–pyrimethamine (IPT/SP) to clear the<br />
placenta periodically of parasites.<br />
INSECTICIDE RESISTANCE<br />
In spite of increased national and international efforts to scale up cost-effective malaria<br />
vector control interventions and maximize the protection of populations at risk, significant<br />
challenges continue to threaten these objectives and the sustainability of achievements.<br />
Challenges include increasing resistance of vector mosquitoes to insecticides, the<br />
behaviour and ecology of local malaria vectors – which often change as a result of vector
control interventions -- and the diminishing number of available insecticides that can be<br />
used against malaria vectors (adulticides).<br />
There are currently no alternatives to DDT and pyrethroids and the development of new<br />
insecticides will be an expensive long-term endeavour. Therefore, immediate sound vector<br />
resistance management practices are required to assure the continued utility of the<br />
currently available insecticides. At present there is only limited evidence of the impact of<br />
various resistance mechanisms on the efficacy of vector control interventions, whether<br />
they are implemented singly or in combination.<br />
Recent evidence from Africa indicates that pyrethroid and DDT resistance is more<br />
widespread than anticipated. It is believed that the same level of resistance will have a<br />
more detrimental impact on the efficacy of IRS than on that of LLINs, but evidence for<br />
this is very limited. Networks for vector resistance monitoring still need greater<br />
strengthening in order to make resistance detection a routine operational feature of<br />
national programmes, particularly in countries in Africa and the Eastern Mediterranean<br />
region. Regional level databases feeding into a global database accessible by governments,<br />
scientists and policy-makers would greatly assist in the rational use and deployment of<br />
vector control interventions.<br />
SOCIOECONOMIC IMPACT<br />
Malaria causes an average loss of 1.3% annual economic growth in countries with intense<br />
transmission. When compounded over the years, this loss has lead to substantial<br />
differences in GDP between countries with and without malaria. Malaria traps families<br />
and communities in a downward spiral of poverty, disproportionately affecting<br />
marginalized populations and poor people who cannot afford treatment or who have<br />
limited access to health care. Malaria’s direct costs include a combination of personal and<br />
public expenditures on both prevention and treatment of disease. In some countries with a<br />
very heavy malaria burden, the disease may account for as much as 40% of public health<br />
expenditure, 30-50% of inpatient admissions and up to 60% of outpatient visits. Malaria<br />
has lifelong effects through increased poverty, impaired learning and decreases attendance<br />
in schools and the workplace.<br />
For more information contact:<br />
WHO Media centre<br />
Telephone: +41 22 791 2222<br />
E-mail: mediainquiries@who.int<br />
http://pathmicro.med.sc.edu/mycology/mycology-1.htm<br />
MYCOLOGY - CHAPTER ONE<br />
INTRODUCTION TO MYCOLOGY
Figure 1.<br />
Chaetomium globosum<br />
spores. Chaetomium is an<br />
ascomycete, and in most<br />
species the spores are lemonshaped,<br />
with a single germ<br />
pore<br />
© Dennis Kunkel Microscopy, Inc. Used<br />
with permission<br />
Figure 2.<br />
Bracket fungus basidiocarp<br />
(fruiting body) lower surface<br />
showing generative hyphae<br />
(gill, spore producing).<br />
Reproductive spores are<br />
dispersed through pores in<br />
the surface of the brackets.<br />
© Dennis Kunkel Microscopy, Inc. Used<br />
with permission<br />
Figure 3.<br />
Mucor spp. fruiting structure<br />
with spores. The fruiting<br />
structure (condiophore) has<br />
matured and its outer<br />
membrane is disintegrating<br />
allowing the spores (conidia)<br />
to be released. Mucor is a<br />
common fungus found in<br />
many environments. It is a<br />
Zygomycetes fungus which<br />
may be allergenic and is often<br />
found as saprobes in soils,<br />
dead plant material (such as<br />
hay), horse dung, and fruits. It<br />
is an opportunistic pathogen<br />
and may cause mucorosis in<br />
immuno-compromised<br />
individuals. The sites of<br />
infections are the lung, nasal<br />
sinus, brain, eye, and skin.<br />
Few species have been<br />
isolated from cases of<br />
zygomycosis, but the term<br />
mucormycosis has often been<br />
used. Zygomycosis includes<br />
mucocutaneous and<br />
rhinocerebral infections, as<br />
well as renal infections,<br />
gastritis, and pulmonary<br />
INTRODUCTION<br />
A. CLASSIFICATION<br />
Fungi are eukaryotic organisms that do not contain chlorophyll, but have cell walls,<br />
filamentous structures, and produce spores. These organisms grow as<br />
saprophytes and decompose dead organic matter. There are between 100,000 to<br />
200,000 species depending on how they are classified. About 300 species are<br />
presently known to be pathogenic for man.<br />
There are five kingdoms of living things. The fungi are in the Kingdom Fungi.<br />
KINGDOM CHARACTERISTIC EXAMPLE<br />
Monera Prokaryocyte Bacteria<br />
Actinomycetes<br />
Protista Eukaryocyte Protozoa<br />
Fungi Eukaryocyte * Fungi<br />
Plantae Eukaryocyte Plants, Moss<br />
Animalia Eukaryocyte * Arthropods<br />
Mammals<br />
Man<br />
*This common characteristic is responsible for the therapeutic dilemma in antimycotic<br />
therapy.<br />
The taxonomy of the Kingdom Fungi is evolving and is controversial. Formerly<br />
based on gross and light microscopic morphology, studies of ultra structure,<br />
biochemistry and molecular biology provide new evidence on which to base<br />
taxonomic positions. Medically important fungi are in four phyla:<br />
1. Ascomycota - Sexual reproduction in a sack called an ascus with the production<br />
of ascopspores (figure 1).<br />
2. Basidiomycota -Sexual reproduction in a sack called a basidium with the<br />
production of basidiospores (figure 2).<br />
3. Zygomycota - sexual reproduction by gametes and asexual reproduction with<br />
the formation of zygospores (figure 3).<br />
4. Mitosporic Fungi (Fungi Imperfecti) - no recognizable form of sexual<br />
reproduction. Includes most pathogenic fungi.
Figure 4.<br />
Candida albicans - yeast and<br />
hyphae stages. A yeast-like<br />
fungus commonly occuring on<br />
human skin, in the upper<br />
respiratory, alimentary &<br />
female genital tracts. This<br />
fungus has a dimorphic life<br />
cycle with yeast and hyphal<br />
stages. The yeast produces<br />
hyphae (strands) and<br />
pseudohyphae. The<br />
pseudohyphae can give rise<br />
to yeast cells by apical or<br />
lateral budding. Causes<br />
candidiasis which includes<br />
thrush (an infection of the<br />
mouth & vagina) and vulvovaginitis.<br />
© Dennis Kunkel<br />
Microscopy, Inc. Used with permission<br />
VIDEO<br />
Growth and Division of<br />
Budding Yeast<br />
(Saccharomyces cerevisiae)<br />
High Resolution<br />
Low resolution<br />
© Philip Meaden<br />
Heriot-Watt University<br />
Edinburgh, Scotland and The<br />
MicrobeLibrary<br />
B. MORPHOLOGY<br />
Pathogenic fungi can exist as yeasts or as hyphae (figure 4). A mass of hyphae is<br />
called mycelia. Yeasts are unicellular organisms and mycelia are multicellular<br />
filamentous structures, constituted by tubular cells with cell walls. The yeasts<br />
reproduce by budding. The mycelial forms branch and the pattern of branching is<br />
an aid to the morphological identification. If the mycelia do not have SEPTA, they<br />
are called coenocytic (nonseptate). The terms "hypha" and "mycelium" are<br />
frequently used interchangeably. Some fungi occur in both the yeast and mycelial<br />
forms. These are called dimorphic fungi.<br />
Dimorphic fungi<br />
The dimorphic fungi have two forms (figure 5):<br />
1. YEAST - (parasitic or pathogenic form). This is the form usually seen in tissue,<br />
in exudates, or if cultured in an incubator at 37 degrees C.<br />
2. MYCELIUM - (saprophytic form). The form observed in nature or when cultured<br />
at 25 degrees C. Conversion to the yeast form appears to be essential for<br />
pathogenicity. In the dimorphic fungi. Fungi are identified by several morphological<br />
or biochemical characteristics, including the appearance of their fruiting bodies.<br />
The asexual spores may be large (macroconidia, chlamydospores) or small<br />
(microconidia, blastospores, arthroconidia).<br />
There are four types of mycotic diseases:<br />
1. Hypersensitivity - an allergic reaction to molds and spores.<br />
2. Mycotoxicoses - poisoning of man and animals by feeds and food products<br />
contaminated by fungi which produce toxins from the grain substrate.<br />
3. Mycetismus - the ingestion of toxin (mushroom poisoning).<br />
4. Infection<br />
We shall be concerned only with the last type: pathogenic fungi that cause<br />
infections. Most common pathogenic fungi do not produce toxins but they do show<br />
physiologic modifications during a parasitic infection (e.g., increased metabolic<br />
rate, modified metabolic pathways and modified cell wall structure). The<br />
mechanisms that cause these modifications as well as their significance as a<br />
pathogenic mechanism are just being described. Most pathogenic fungi are also<br />
thermotolerant, and can resist the effects of the active oxygen radicals released<br />
during the respiratory burst of phagocytes. Thus, fungi are able to withstand many<br />
host defenses. Fungi are ubiquitous in nature and most people are exposed to<br />
them. The establishment of a mycotic infection usually depends on the size of the<br />
inoculum and on the resistance of the host. The severity of the infection seems to<br />
depend mostly on the immunologic status of the host. Thus, the demonstration of<br />
fungi, for example, in blood drawn from an intravenous catheter can correspond to
A<br />
Candida albicans is a<br />
dimorphic fungus in that it<br />
grows as a unicellular yeast<br />
under some environmental<br />
conditions and as a<br />
filamentous fungus under<br />
other conditions.<br />
Budding yeast cells. C.<br />
albicans was grown at 37°C<br />
with aeration for 3 h in yeastpeptone-dextrose<br />
(YPD)<br />
medium. In this image,<br />
unstained cells are magnified<br />
x400. The image was taken<br />
with phase- contrast<br />
microscopy.<br />
colonization of the catheter, to transient fungemia (i.e., dissemination of fungi<br />
through the blood stream), or to a true infection. The physician must decide which<br />
is the clinical status of the patient based on clinical parameters, general status of<br />
the patient, laboratory results, etc. The decision is not trivial, since treatment of<br />
systemic fungal infections requires the aggressive use of drugs with considerable<br />
toxicity. Most mycotic agents are soil saprophytes and mycotic diseases are<br />
generally not communicable from person-to-person (occasional exceptions:<br />
Candida and some dermatophytes). Outbreaks of disease may occur, but these<br />
are due to a common environmental exposure, not communicability. Most of the<br />
fungi which cause systemic infections have a peculiar, characteristic ecologic<br />
niche in nature. This habitat is specific for several fungi which will be discussed<br />
later. In this environment, the normally saprophytic organisms proliferate and<br />
develop. This habitat is also the source of fungal elements and/or spores, where<br />
man and animals, incidental hosts, are exposed to the infectious particles. It is<br />
important to be aware of these associations to diagnose mycotic diseases. The<br />
physician must be able to elicit a complete history from the patient including<br />
occupation, avocation and travel history. This information is frequently required to<br />
raise, or confirm, your differential diagnosis. The incidence of mycotic infections is<br />
currently increasing dramatically, due to an increased population of susceptibles.<br />
Examples are patients with AIDS, patients on immunosuppressive therapy, and<br />
the use of more invasive diagnostic and surgical procedures (prosthetic implants).<br />
Fungal diseases are non-contagious and non-reportable diseases in the national<br />
public health statistics. However, in South Carolina most of the important mycotic<br />
(fungal) diseases were notifiable to the public health authorities until 1994.<br />
B<br />
Budding yeast with septum. The septum has formed between the daughter bud and the mother<br />
cell, but separation of the two has not occurred. This image is from a culture of cells grown at 37°<br />
C for 3 h in YPD medium. The unstained cell is magnified x1,000 using phase- contrast<br />
microscopy.<br />
C<br />
Candida albicans mother and daughter cells. Cells were grown under conditions that induced<br />
hypha formation for 30 min. The daughter cell is on the right; the mother cell is on the left. The<br />
daughter cell has not reached a threshold volume and therefore has not yet formed a hypha. The<br />
mother cell has passed the threshold volume and has started forming a germ tube which will<br />
become a hypha. The germ tube seen here is 6 min old. A septum between the germ tube and the
Figure 5 A-E<br />
© Phillip Stafford<br />
Dartmouth Medical School<br />
Hanover, New Hampshire and<br />
The MicrobeLibrary<br />
mother cell has not yet formed. The unstained cells are magnified x1,000 using phase-contrast<br />
microscopy.<br />
D<br />
C. albicans cell at 3 h. Three hours after the appearance of the germ tube, the hypha has septa. A<br />
new germ tube at the distal pole of the<br />
cell is also evident at this time. The unstained cells are magnified x1,000 using phase-contrast<br />
microscopy.<br />
E<br />
C. albicans hyphal cells at 5 h. After 5 h in hypha-inducing medium, many hyphae are evident.<br />
Clumping of the hyphae is<br />
also apparent, and hyphae are beginning to form hypha blastospores, which are new budding<br />
cells.
Figure 6<br />
A Sabouraud’s dextrose agar<br />
plate culture growing a<br />
Mexican isolate of T. rubrum<br />
var. rodhaini. Dermatophytic<br />
members of the genus<br />
Trichophyton are some of the<br />
leading causes of hair, skin,<br />
and nail infections in humans,<br />
known as dermatophytoses.<br />
The genus includes<br />
anthropophilic, zoophilic, and<br />
geophilic species<br />
CDC/Dr. Libero Ajello<br />
MOLECULAR<br />
STRUCTURE<br />
Amphotericin B<br />
Ketoconazole<br />
Griseofulvin<br />
5-fluorocytosine<br />
C. DIAGNOSIS<br />
1. Skin scrapings suspected to contain dermatophytes or pus from a lesion can be<br />
mounted in KOH on a slide and examined directly under the microscope.<br />
2. Skin testing (dermal hypersensitivity) used to be popular as a diagnostic tool,<br />
but this use is now discouraged because the skin test may interfere with<br />
serological studies, by causing false positive results. It may still be used to<br />
evaluate the patient's immunity, as well as a population exposure index in<br />
epidemiological studies.<br />
3. Serology may be helpful when it is applied to a specific fungal disease; there<br />
are no screening antigens for 'fungi' in general. Because fungi are poor antigens,<br />
the efficacy of serology varies with different fungal infections. The serologic tests<br />
will be discussed under each mycosis. The most common serological tests for<br />
fungi are based on latex agglutination, double immunodiffusion, complement<br />
fixation and enzyme immunoassays. While latex agglutination may favor the<br />
detection of IgM antibodies, double immunodiffusion and complement fixation<br />
usually detect IgG antibodies. Some EIA tests are being developed to detect both<br />
IgG and IgM antibodies. There are some tests which can detect specific fungal<br />
antigens, but they are just coming into general use.<br />
4. Direct fluorescent microscopy may be used for identification, even on nonviable<br />
cultures or on fixed tissue sections. The reagents for this test are difficult to<br />
obtain.<br />
5. Biopsy and histopathology. A biopsy may be very useful for the identification<br />
and as a source of the of tissue-invading fungi. Usually the Gomori methenamine<br />
silver (GMS) stain is used to reveal the organisms which stain black against a<br />
green background. The H&E stain does not always tint the organism, but it will<br />
stain the inflammatory cells.<br />
6. Culture. A definitive diagnosis requires a culture and identification. Pathogenic<br />
fungi are usually grown on Sabouraud dextrose agar (figure 6). It has a slightly<br />
acidic pH (~5.6); cyclohexamide, penicillin, streptomycin or other inhibitory<br />
antibiotics are often added to prevent bacterial contamination and overgrowth.<br />
Two cultures are inoculated and incubated separately at 25 degrees C and 37<br />
degrees C to reveal dimorphism. The cultures are examined macroscopically and<br />
microscopically. They are not considered negative for growth until after 4 weeks of<br />
incubation.<br />
D. TREATMENT<br />
Mammalian cells do not contain the enzymes which will degrade the cell wall<br />
polysaccharides of fungi. Therefore, these pathogens are difficult to eradicate by<br />
the animal host defense mechanisms. Because mammals and fungi are both
eukaryotic, the cellular milieu is biochemically similar in both. The cell membranes<br />
of all eukaryotic cells contain sterols; ergosterol in the fungal cell membrane and<br />
cholesterol in the mammalian cell membrane. Thus, most substances which may<br />
impair the invading fungus will usually have serious side effects on the host.<br />
Although one of the first chemotherapeutic agents (oral iodides) was an antimycotic<br />
used in 1903, the further development of such agents has been left far<br />
behind the development of anti-bacterial agents. The selective toxicity necessary<br />
to inhibit the invading organism with minimal damage to the host has been difficult<br />
to establish within eukaryotic cells.<br />
The primary antifungal agents are:<br />
Amphotericin B<br />
A polyene antimycotic. It is usually the drug of choice for most systemic fungal<br />
infections. It has a greater affinity for ergosterol in the cell membranes of fungi<br />
than for the cholesterol in the host's cells; once bound to ergosterol, it causes<br />
disruption of the cell membrane and death of the fungal cell. Amphotericin B is<br />
usually administered intravenously (patient usually needs to be hospitalized), often<br />
for 2-3 months. The drug is rather toxic; thrombo-phlebitis, nephrotoxicity, fever,<br />
chills and anemia frequently occur during administration.<br />
Azoles<br />
The azoles (imidazoles and triazoles), including ketoconazole, fluconazole, and<br />
itraconozole, are being used for muco-cutaneous candidiasis, dermatophytosis,<br />
and for some systemic fungal infections. Fluconazole is presently essential for the<br />
maintenance of AIDS patients with cryptococcosis. The general mechanism of<br />
action of the azoles is the inhibition of ergosterol synthesis. Oral administration<br />
and reduced toxicity are distinct advantages.<br />
Griseofulvin<br />
Griseofulvin is a very slow-acting drug which is used for severe skin and nail<br />
infections. Its effect depends on its accumulation in the stratum corneum where it<br />
is incorporated into the tissue and forms a barrier which stops further fungal<br />
penetration and growth. It is administered orally. The exact mechanism of action is<br />
unknown.<br />
5-fluorocytosine<br />
5-fluorocytosine (Flucytosine or 5-FC) inhibits RNA synthesis and has found its<br />
main application in cryptococcosis (to be discussed later). It is administered orally.<br />
E. CLINICAL CLASSIFICATION OF THE MYCOSES<br />
Fungal diseases may be discussed in a variety of ways. The most practical<br />
method for medical students is the clinical taxonomy which divides the fungi into:<br />
a. Superficial mycoses<br />
b. Subcutaneous mycoses<br />
c. Systemic mycoses
MOLECULAR<br />
STRUCTURE<br />
Ergosterol<br />
Figure 7.<br />
Ringworm on the skin of the<br />
neck due to Trichophyton<br />
rubrum.<br />
CDC/Lucille K. Georg<br />
Return to the Mycology Section of Microbiology and Immunology On-line<br />
This page copyright 2007, The Board of Trustees of the University of South Carolina<br />
This page last changed on<br />
Page maintained by Richard Hunt<br />
Please report any problems to rhunt@med.sc.edu<br />
http://en.wikipedia.org/wiki/Microbial_metabolism<br />
Microbial metabolism<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
Microbial metabolism is the means by which a microbe obtains the energy and<br />
nutrients (e.g. carbon) it needs to live and reproduce. Microbes use many different<br />
types of metabolic strategies and species can often be differentiated from each other<br />
based on metabolic characteristics. The specific metabolic properties of a microbe are<br />
the major factors in determining that microbe’s ecological niche, and often allow for<br />
that microbe to be useful in industrial processes or responsible for biogeochemical<br />
cycles.<br />
Contents
[hide]<br />
• 1 Types of microbial metabolism<br />
• 2 Heterotrophic microbial metabolism<br />
• 3 Fermentation<br />
• 4 Special metabolic properties<br />
o 4.1 Methylotrophy<br />
o 4.2 Syntrophy<br />
• 5 Anaerobic respiration<br />
o 5.1 Denitrification<br />
o 5.2 Sulfate reduction<br />
o 5.3 Acetogenesis<br />
o 5.4 Inorganic electron acceptors<br />
o 5.5 Organic terminal electron acceptors<br />
• 6 Chemolithotrophy<br />
o 6.1 Hydrogen oxidation<br />
o 6.2 Sulfur oxidation<br />
o 6.3 Ferrous iron (Fe 2+ ) oxidation<br />
o 6.4 Nitrification<br />
o 6.5 Anammox<br />
• 7 Phototrophy<br />
• 8 Nitrogen fixation<br />
• 9 See also<br />
• 10 References<br />
Types of microbial metabolism<br />
Flow chart to determine the metabolic characteristics of microorganisms<br />
Main article: Primary nutritional groups<br />
All microbial metabolism can be arranged according to three principles:
1. How the organism obtains carbon for synthesising cell mass:<br />
• autotrophic – carbon is obtained from carbon dioxide (CO2)<br />
• heterotrophic – carbon is obtained from organic compounds<br />
• mixotrophic – carbon is obtained from both organic compounds and by fixing<br />
carbon dioxide<br />
2. How the organism obtains reducing equivalents used either in energy conservation<br />
or in biosynthetic reactions:<br />
• lithotrophic – reducing equivalents are obtained from inorganic compounds<br />
• organotrophic – reducing equivalents are obtained from organic compounds<br />
3. How the organism obtains energy for living and growing:<br />
• chemotrophic – energy is obtained from external chemical compounds<br />
• phototrophic – energy is obtained from light<br />
In practice, these terms are almost freely combined. Typical examples are as follows:<br />
• chemolithoautotrophs obtain energy from the oxidation of inorganic<br />
compounds and carbon from the fixation of carbon dioxide. Examples:<br />
Nitrifying bacteria, Sulfur-oxidising bacteria, Iron-oxidising bacteria,<br />
Knallgas-bacteria<br />
• photolithoautotrophs obtain energy from light and carbon from the fixation<br />
of carbon dioxide, using reducing equivalents from inorganic compounds.<br />
Examples: Cyanobacteria (water as reducing equivalent donor),<br />
Chlorobiaceae, Chromaticaceae (hydrogen sulfide as reducing equivalent<br />
donor), Chloroflexus (hydrogen as reducing equivalent donor)<br />
• chemolithoheterotrophs obtain energy from the oxidation of inorganic<br />
compounds, but can not fix carbon dioxide. Examples: some Nitrobacter spp.,<br />
Wolinella (with H2 as reducing equivalent donor), some Knallgas-bacteria<br />
• chemoorganoheterotrophs obtain energy, carbon and reducing equivalents<br />
for biosynthetic reactions from organic compounds. Examples: most bacteria,<br />
e. g. Escherichia coli, Bacillus spp., Actinobacteria<br />
• photoorganoheterotrophs obtain energy from light, carbon and reducing<br />
equivalents for biosynthetic reactions from organic compounds. Some species<br />
are strictly heterotrophic, many others can also fix carbon dioxide and are<br />
mixotrophic. Examples: Rhodobacter, Rhodopseudomonas, Rhodospirillum,<br />
Rhodomicrobium, Rhodocyclus, Heliobacterium, Chloroflexus (alternatively to<br />
photolithoautotrophy with hydrogen)<br />
[edit] Heterotrophic microbial metabolism<br />
Most microbes are heterotrophic (more precisely chemoorganoheterotrophic), using<br />
organic compounds as both carbon and energy sources. Heterotrophic microbes live<br />
off of nutrients that they scavenge from living hosts (as commensals or parasites) or<br />
find in dead organic matter of all kind (saprophages). Microbial metabolism is the<br />
main contribution for the bodily decay of all organisms after death. Many eukaryotic
microorganisms are heterotrophic by predation or parasitism, properties also found in<br />
some bacteria such as Bdellovibrio (an intracellular parasite of other bacteria, causing<br />
death of its victims) and Myxobacteria such as Myxococcus (predators of other<br />
bacteria which are killed and lysed by cooperating swarms of many single cells of<br />
Myxobacteria). Most pathogenic bacteria can be viewed as heterotrophic parasites of<br />
humans or whatever other eukaryotic species they affect. Heterotrophic microbes are<br />
extremely abundant in nature and are responsible for the breakdown of large organic<br />
polymers such as cellulose, chitin or lignin which are generally indigestible to larger<br />
animals. Generally, the breakdown of large polymers to carbon dioxide<br />
(mineralization) requires several different organisms, with one breaking down the<br />
polymer into its constituent monomers, one able to use the monomers and excreting<br />
simpler waste compounds as by-products and one able to use the excreted wastes.<br />
There are many variations on this theme, as different organisms are able to degrade<br />
different polymers and secrete different waste products. Some organisms are even<br />
able to degrade more recalcitrant compounds such as petroleum compounds or<br />
pesticides, making them useful in bioremediation.<br />
Biochemically, prokaryotic heterotrophic metabolism is much more versatile than that<br />
of eukaryotic organisms, although many prokaryotes share the most basic metabolic<br />
models with eukaryotes, e. g. using glycolysis (also called EMP pathway) for sugar<br />
metabolism and the citric acid cycle to degrade acetate, producing energy in the form<br />
of ATP and reducing power in the form of NADH or quinols. These basic pathways<br />
are well conserved because they are also involved in biosynthesis of many conserved<br />
building blocks needed for cell growth (sometimes in reverse direction). However,<br />
many bacteria and archaea utilise alternative metabolic pathways other than glycolysis<br />
and the citric acid cycle. A well studied example is sugar metabolism via the ketodeoxy-phosphogluconate<br />
pathway (also called ED pathway) in Pseudomonas] instead<br />
of the glycolytic pathway. Moreover, there is even a third alternative sugar-catabolic<br />
pathway used by some bacteria, the pentose-phosphate pathway. This metabolic<br />
diversity and ability of prokaryotes to use a huge variety of organic compounds arises<br />
from the much deeper evolutionary history and diversity of prokaryotes, as compared<br />
to eukaryotes. It is also noteworthy that the mitochondrion, the small membranebound<br />
intracellular organelle that is the site of eukaryotic energy metabolism, arose<br />
from the endosymbiosis of a bacterium related to obligate intracellular Rickettsia, and<br />
also to plant-associated Rhizobium or Agrobacterium. Therefore it is not surprising<br />
that all mitrochondriate eukaryotes share metabolic properties with these<br />
Proteobacteria. Most microbes respire (use an electron transport chain), although<br />
oxygen is not the only terminal electron acceptor that may be used. As discussed<br />
below, the use of terminal electron acceptors other than oxygen has important<br />
biogeochemical consequences.<br />
[edit] Fermentation<br />
Main article: Fermentation (biochemistry)<br />
Fermentation is a specific type of heterotrophic metabolism that uses organic carbon<br />
instead of oxygen as a terminal electron acceptor. This means that these organisms do<br />
not use an electron transport chain to oxidize NADH to NAD + and therefore must<br />
have an alternative method of using this reducing power and maintaining a supply of<br />
NAD + for the proper functioning of normal metabolic pathways (e.g. glycolysis). As
oxygen is not required, fermentative organisms are anaerobic. Many organisms can<br />
use fermentation under anaerobic conditions and respiration when oxygen is not<br />
present. These organisms are facultative anaerobes. To avoid the overproduction of<br />
NADH obligately fermentative organisms usually do not have a complete citric acid<br />
cycle. Instead of using an ATPase as in respiration, ATP in fermentative organisms is<br />
produced by substrate-level phosphorylation where a phosphate group is transferred<br />
from a high-energy organic compound to ADP to form ATP. As a result of the need to<br />
produce high energy phosphate-containing organic compounds (generally in the form<br />
of CoA-esters) fermentative organisms use NADH and other cofactors to produce<br />
many different reduced metabolic by-products, often including hydrogen gas (H2).<br />
These reduced organic compounds are generally small organic acids and alcohols<br />
derived from pyruvate, the end product of glycolysis. Examples include ethanol,<br />
acetate, lactate and butyrate. Fermentative organisms are very important industrially<br />
and are used to make many different types of food products. The different metabolic<br />
end products produced by each specific bacterial species are responsible for the<br />
different tastes and properties of each food.<br />
Not all fermentative organisms use substrate-level phosphorylation. Instead, some<br />
organisms are able to couple the oxidation of low-energy organic compounds directly<br />
to the formation of a proton (or sodium) motive force and therefore ATP synthesis.<br />
Examples of these unusual forms of fermentation include succinate fermentation by<br />
Propionigenium modestum and oxalate fermentation by Oxalobacter formigenes.<br />
These reactions are extremely low energy-yielding. Humans and other higher animals<br />
also use fermentation to use excess NADH to produce lactate, although this is not the<br />
major form of metabolism as it is in fermentative microorganisms.<br />
[edit] Special metabolic properties<br />
[edit] Methylotrophy<br />
Methylotrophy refers to the ability of an organism to use C1-compounds as energy<br />
sources. These compounds include methanol, methyl amines, formaldehyde and<br />
formate. Several other, less common substrates may also be used for metabolism, all<br />
of which lack carbon-carbon bonds. Examples of methylotrophs include the bacteria<br />
Methylomonas and Methylobacter. Methanotrophs are a specific type of methylotroph<br />
that are also able to use methane (CH4) as a carbon source by oxidizing it sequentially<br />
to methanol (CH3OH), formaldehyde (CH2O), formate (HCOO - ) and finally carbon<br />
dioxide CO2 initially using the important enzyme methane monooxygenase. As<br />
oxygen is required for this process, all (conventional) methanotrophs are obligate<br />
aerobes. Reducing power in the form of quinones and NADH is produced during<br />
these oxidations to produce a proton motive force and therefore ATP generation.<br />
Methylotrophs and methanotrophs are not considered as autotrophic, because they are<br />
able to incorporate some of the oxidized methane (or other metabolites) into cellular<br />
carbon before it is completely oxidised to CO2 (at the level of formaldehyde), using<br />
either the serine pathway (Methylosinus, Methylocystis) or the ribulose<br />
monophosphate pathway (Methylococcus), depending on the species of methylotroph.<br />
In addition to aerobic methylotrophy, methane can also be oxidized anaerobically.<br />
This occurs by a consortium of sulfate-reducing bacteria and relatives of
methanogenic Archaea working syntrophically (see below). Little is currently known<br />
about the biochemistry and ecology of this process.<br />
Methanogenesis is the biological production of methane. It is carried out by<br />
methanogens, strictly anaerobic archaea such as Methanococcus,<br />
Methanocaldococcus, Methanobacterium, Methanothermus, Methanosarcina,<br />
Methanosaeta and Methanopyrus. The biochemistry of methanogenesis is unique in<br />
nature in its use of a number of unusual cofactors to sequentially reduce<br />
methanogenic substrates to methane. These cofactors are responsible (among other<br />
things) for the establishment of a proton gradient across the outer membrane thereby<br />
driving ATP synthesis. Several different types of methanogenesis occurs, which differ<br />
in the starting compounds oxidized. Some methanogens reduce carbon dioxide (CO2)<br />
to methane (CH4) using electrons (most often) from hydrogen gas (H2)<br />
chemolithoautotrophically. These methanogens can often be found in environments<br />
containing fermentative organisms. The tight association of methanogens and<br />
fermentative bacteria can be considered to be syntrophic (see below) because the<br />
methanogens, which rely on the fermentors for hydrogen, relieve feedback inhibition<br />
of the fermentors by the build-up of excess hydrogen that would otherwise inhibit<br />
their growth. This type of syntrophic relationship is specifically known as interspecies<br />
hydrogen transfer. A second group of methanogens use methanol (CH3OH) as a<br />
substrate for methanogenesis. These are chemoorganotrophic, but still autotrophic in<br />
using CO2 as only carbon source. The biochemistry of this process is quite different<br />
from that of the carbon dioxide reducing methanogens. Lastly, a third group of<br />
methanogens produce both methane and carbon dioxide from acetate (CH3COO - ) with<br />
the acetate being literally split between the two carbons. These acetate-cleaving<br />
organisms are the only chemoorganoheterotrophic methanogens. All<br />
autotrophicmethanogens use a variation of the acetyl-CoA pathway to fix CO2 and<br />
obtain cellular carbon.<br />
[edit] Syntrophy<br />
Syntrophy, in the context of microbial metabolism, refers to the pairing of multiple<br />
species to achieve a chemical reaction that, on its own, would be energetically<br />
unfavorable. The best studied example of this process is the oxidation of fermentative<br />
end products (such as acetate, ethanol and butyrate) by organisms such as<br />
Syntrophomonas. Alone, the oxidation of butyrate to acetate and hydrogen gas is<br />
energetically unfavorable. However, when a hydrogenotrophic (hydrogen using)<br />
methanogen is present the use of the hydrogen gas will significantly lower the<br />
concentration of hydrogen (down to 10 -5 atm) and thereby shift the equilibrium of the<br />
butyrate oxidation reaction under standard conditions (ΔGº’) to non-standard<br />
conditions (ΔG’). Because the concentration of one product is lowered, the reaction is<br />
"pulled" towards the products and shifted towards net energetically favorable<br />
conditions (for butyrate oxidation: ΔGº’= +48.2 kJ/mol, but ΔG' = -8.9 kJ/mol at 10 -5<br />
atm hydrogen and even lower if also the initially produced acetate is further<br />
metabolised by methanogens). Conversely, the available free energy from<br />
methanogenesis is lowered from ΔGº’= -131 kJ/mol under standard conditions to ΔG'<br />
= -17 kJ/mol at 10 -5 atm hydrogen. This is an example of intraspecies hydrogen<br />
transfer. In this way, low energy-yielding carbon sources can be used by a consortium<br />
of organisms to achieve further degradation and eventual mineralization of these<br />
compounds. These reactions help prevent the excess sequestration of carbon over
geologic time scales, releasing it back to the biosphere in usable forms such as<br />
methane and CO2.<br />
[edit] Anaerobic respiration<br />
In aerobic organisms, oxygen is used as a terminal electron acceptor during<br />
respiration. This is largely because oxygen has a very low reduction potential<br />
allowing for aerobic organisms to utilize their electron transport systems most<br />
efficiently. In anaerobic organisms, terminal electron acceptors other than oxygen are<br />
used. These inorganic compounds have a higher reduction potential compared to<br />
oxygen, meaning that respiration is less efficient in these organisms generally leading<br />
to slower growth rates compared to aerobes. Many facultative anaerobes can use<br />
either oxygen or alternative terminal electron acceptors for respiration depending on<br />
the environmental conditions. Most respiring anaerobes are heterotrophs, although<br />
some do live autotrophically. All of the processes described below are dissimilative,<br />
meaning that they are used during energy production and not to provide nutrients for<br />
the cell (assimilative). Assimilative pathways for many forms of anaerobic respiration<br />
are also known.<br />
[edit] Denitrification<br />
Main article: Denitrification<br />
Denitrification is the utilization of nitrate (NO3 - ) as a terminal electron acceptor. It is a<br />
widespread process that is used by many members of the Proteobacteria. Many<br />
facultative anaerobes use denitrification because nitrate, like oxygen, has a low<br />
reduction potential. Many denitrifying bacteria can also use ferric iron (Fe 3+ ) and<br />
some organic electron acceptors. Denitrification involves the stepwise reduction of<br />
nitrate to nitrite (NO2 - ), nitric oxide (NO), nitrous oxide (N2O) and dinitrogen (N2) by<br />
the enzymes nitrate reductase, nitrite reductase, nitric oxide reductase and nitrous<br />
oxide reductase, respectively. Protons are transported across the membrane by the<br />
initial NADH reductase, quinones and nitrous oxide reductase to produce the<br />
electrochemical gradient critical for respiration. Some organisms (e.g. E. coli) only<br />
produce nitrate reductase and therefore can accomplish only the first reduction<br />
leading to the accumulation of nitrite. Others (e.g. Paracoccus denitrificans or<br />
Pseudomonas stutzeri) reduce nitrate completely. Complete denitrification is an<br />
environmentally significant process because some intermediates of denitrification<br />
(nitric oxide and nitrous oxide) are important greenhouse gases that react with<br />
sunlight and ozone to produce nitric acid, a component of acid rain. Denitrification is<br />
also important in biological wastewater treatment where it is used to reduce the<br />
amount of nitrogen released into the environment thereby reducing eutrophication.<br />
[edit] Sulfate reduction<br />
Sulfate reduction is a relatively energetically poor process used by many Gram<br />
negative bacteria found within the δ-Proteobacteria, Gram positive organisms relating<br />
to Desulfotomaculum or the archaeon Archaeoglobus. Hydrogen sulfide (H2S) is<br />
produced as a metabolic end product. Many sulfate reducers are heterotrophic, using<br />
carbon compounds such as lactate and pyruvate (among many others) as electron
donors while others are autotrophic, using hydrogen gas (H2) as an electron donor.<br />
Some unusual autotrophic sulfate reducing-bacteria can use phosphite (HPO3 - ) as an<br />
electron donor (e.g. Desulfotignum phosphitoxidans) or are capable of sulfur<br />
disproportionation (splitting one compound into two different compounds, in this case<br />
an electron donor and an electron acceptor) using thiosulfate (S2O3 2- e.g.<br />
Desulfovibrio sulfodismutans). All sulfate reducing-organisms are strict anaerobes.<br />
Because sulfate is energetically stable before it can be metabolized it must first be<br />
activated by adenylation to form APS (adenosine 5’-phosphosulfate) thereby<br />
consuming ATP. The APS is then reduced by the enzyme APS reductase to form<br />
sulfite (SO3 2- and AMP. In organisms that use carbon compounds as electron donors,<br />
the ATP consumed is accounted for by fermentation of the carbon substrate. The<br />
hydrogen produced during fermentation is actually what drives respiration during<br />
sulfate reduction. Electrons are passed from the hydrogenase enzyme eventually to the<br />
APS reductase, which along with sulfite reductase completes the reduction of sulfate<br />
to hydrogen sulfide. A proton motive force is established due to a fact that the<br />
hydrogenase, which converts H2 to 2H + is located in the periplasm (or extracellularly<br />
in Gram positive bacteria).<br />
[edit] Acetogenesis<br />
Main article: Acetogenesis<br />
Acetogenesis is a type of microbial metabolism that uses hydrogen (H2) as an electron<br />
donor and carbon dioxide (CO2) as an electron acceptor to produce acetate. This is<br />
similar to methanogenesis (see above) in having the same electron donors and<br />
acceptors. Bacteria that can autotrophically synthesize acetate are called<br />
homoacetogens. Carbon dioxide reduction in all homoacetogens occurs by the acetyl-<br />
CoA pathway. This pathway is also used for carbon fixation by autotrophic sulfatereducing<br />
bacteria and hydrogenotrophic methanogens. Often homoacetogens can also<br />
be fermentative, using the hydrogen and carbon dioxide produced as a result of<br />
fermentation to produce acetate, which is secreted as an end product.<br />
[edit] Inorganic electron acceptors<br />
Ferric iron (Fe 3+ ) is a widespread anaerobic terminal electron acceptor both for<br />
autotrophic and heterotrophic organisms. Electron flow in these organisms is similar<br />
to those in electron transport ending in oxygen or nitrate except that in ferric ironreducing<br />
organisms the final enzyme in this system is a ferric iron reductase. Model<br />
organisms include Shewanella putrifaciens and Geobacter metallireducens. Since<br />
some ferric iron-reducing bacteria (e.g. G. metallireducens) can use toxic<br />
hydrocarbons such as toluene as a carbon source there is significant interest in using<br />
these organisms as bioremediation agents in ferric iron-rich contaminated aquifers.<br />
Although Ferric iron is the most prevalent inorganic electron acceptor, a number of<br />
organisms (including the iron-reducing bacteria mentioned above) can use other<br />
inorganic ions in anaerobic respiration. While these processes may often be less<br />
significant ecologically, they are of considerable interest for bioremediation,<br />
especially when heavy metals or radionuclides are used as electron acceptors.<br />
Examples include:
• Manganic ion (Mn 4+ ) reduction to manganous ion (Mn 2+ )<br />
• Selenate (SeO4 2- ) reduction to selenite (SeO3 2- ) and selenite reduction to<br />
inorganic selenium (Se 0 )<br />
• Arsenate (AsO4 3- ) reduction to arsenite (AsO3 3- )<br />
• Uranyl ion ion (UO2 2+ ) reduction to uranium dioxide (UO2)<br />
[edit] Organic terminal electron acceptors<br />
An number of organisms, instead of using inorganic compounds as terminal electron<br />
acceptors are able to use organic compounds to accept electrons from respiration.<br />
Examples include:<br />
• Fumarate reduction to succinate<br />
• Trimethylamine N-oxide (TMAO) reduction to trimethylamine (TMA)<br />
• Dimethyl sulfoxide (DMSO) reduction to Dimethyl sulfide (DMS)<br />
• Reductive dechlorination<br />
TMAO is a chemical commonly produced by fish, and when reduced to TMA<br />
produces a strong odor. DMSO is a common marine and freshwater chemical which is<br />
also odiferous when reduced to DMS. Reductive dechlorination is the process by<br />
which chlorinated organic compounds are reduced to form their non-chlorinated<br />
endproducts. As chlorinated organic compounds are often important (and difficult to<br />
degrade) environmental polutants, reductive dechlorination is an important process in<br />
bioremediation.<br />
[edit] Chemolithotrophy<br />
Chemolithotrophy is a type of metabolism where energy is obtained from the<br />
oxidation of inorganic compounds. Most chemolithotrophic organisms are also<br />
autotrophic. There are two major objectives to chemolithotrophy: the generation of<br />
energy (ATP) and the generation of reducing power (NADH).<br />
[edit] Hydrogen oxidation<br />
Many organisms are capable of using hydrogen (H2) as a source of energy. While<br />
several mechanisms of anaerobic hydrogen oxidation have been mentioned previously<br />
(e.g. sulfate reducing- and acetogenic bacteria) hydrogen can also be used as an<br />
energy source aerobically. In these organisms hydrogen is oxidized by a membranebound<br />
hydrogenase causing proton pumping via electron transfer to various quinones<br />
and cytochromes. In many organisms, a second cytoplasmic hydrogenase is used to<br />
generate reducing power in the form of NADH, which is subsequently used to fix<br />
carbon dioxide via the Calvin cycle. Hydrogen oxidizing organisms, such as<br />
Cupriavidus necator (formerly Ralstonia eutropha)--Louegger (talk) 16:11, 3<br />
February 2008 (UTC), often inhabit oxic-anoxic interfaces in nature to take advantage<br />
of the hydrogen produced by anaerobic fermentative organisms while still maintaining<br />
a supply of oxygen.<br />
[edit] Sulfur oxidation
Sulfur oxidation involves the oxidation of reduced sulfur compounds (such as sulfide<br />
(H2S), inorganic sulfur (S 0 ) and thiosulfate (S2O2 2- ) ) to form sulfuric acid (H2SO4). A<br />
classic example of a sulfur oxidizing bacterium is Beggiatoa, a microbe originally<br />
described by Sergei Winogradsky, one of the founders of microbiology. Generally,<br />
the oxidation of sulfide occurs in stages, with inorganic sulfur being stored either<br />
inside or outside of the cell until needed. This two step process occurs because<br />
energetically sulfide is a better electron donor than inorganic sulfur or thiosulfate,<br />
allowing for a greater number of protons to be translocated across the membrane.<br />
Sulfur oxidizing organisms generate reducing power for carbon dioxide fixation via<br />
the Calvin cycle using reverse electron flow, an energy-requiring process that pushes<br />
the electrons against their thermodynamic gradient to produce NADH. Biochemically,<br />
reduced sulfur compounds are converted to sulfite (SO3 2- ) and subsequently converted<br />
to sulfate by the enzyme sulfite oxidase. Some organisms, however, accomplish the<br />
same oxidation using a reversal of the APS reductase system used by sulfate-reducing<br />
bacteria (see above). In all cases the energy liberated is transferred to the electron<br />
transport chain for ATP and NADH production. In addition to aerobic sulfur<br />
oxidation, some organisms (e.g. Thiobacillus denitrificans) use nitrate (NO3 2- ) as a<br />
terminal electron acceptor and therefore grow anaerobically.<br />
[edit] Ferrous iron (Fe 2+ ) oxidation<br />
Ferrous iron is a soluble form of iron that is stable at extremely low pHs or under<br />
anaerobic conditions. Under aerobic, moderate pH conditions ferrous iron is oxidized<br />
spontaneously to the ferric (Fe 3+ ) form and is hydrolyzed abiotically to insoluble<br />
ferric hydroxide (Fe(OH)3). There exists, therefore, three distinct types of ferrous<br />
iron-oxidizing microbes. The first are acidophiles, such as the bacteria<br />
Acidithiobacillus ferooxidans and Leptospirrillum ferrooxidans, as well as the<br />
archaeon Ferroplasma. These microbes oxidize iron in environments that have a very<br />
low pH and are important in acid mine drainage. The second type of microbes oxidize<br />
ferrous iron at neutral pH along oxic-anoxic interfaces. Both these bacteria, such as<br />
Gallionella ferruginea and Sphaerotilus natans, and the acidophilic iron oxidizingbacteria<br />
are aerobes. The third type of iron-oxidizing microbes are anaerobic<br />
photosynthetic bacteria such as Chlorobium, which use ferrous iron to produce<br />
NADH for autotrophic carbon dioxide fixation. Biochemically, aerobic iron reduction<br />
is a very energetically poor process which therefore requires large amounts of iron to<br />
be oxidized by the enzyme rusticyanin to facilitate the formation of proton motive<br />
force. Like during sulfur oxidation reverse electron flow must be used to form the<br />
NADH used for carbon dioxide fixation via the Calvin cycle.<br />
[edit] Nitrification<br />
Nitrification is the process by which ammonia (NH3) is converted to nitrate (NO3 - ).<br />
Nitrification is actually the net result of two distinct processes: oxidation of ammonia<br />
to nitrite (NO2 - ) by nitrosifying bacteria (e.g. Nitrosomonas) and oxidation of nitrite to<br />
nitrate by the nitrite-oxidizing bacteria (e.g. Nitrobacter). Both of these processes are<br />
extremely poor energetically leading to very slow growth rates for both types of<br />
organisms. Biochemically, ammonia oxidation occurs by the stepwise oxidation of<br />
ammonia to hydroxylamine (NH2OH) by the enzyme ammonia monooxygenase in the<br />
cytoplasm followed by the oxidation of hydroxylamine to nitrite by the enzyme<br />
hydroxylamine oxidoreductase in the periplasm.
Electron and proton cycling are very complex but as a net result only one proton is<br />
translocated across the membrane per molecule of ammonia oxidized. Nitrite<br />
reduction is much simpler, with nitrite being oxidized by the enzyme nitrite<br />
oxidoreductase coupled to proton translocation by a very short electron transport<br />
chain, again leading to very low growth rates for these organisms. In both ammonia-<br />
and nitrite-oxidation oxygen is required, meaning that both nitrosifying and nitriteoxidizing<br />
bacteria are aerobes. As in sulfur and iron oxidation, NADH for carbon<br />
dioxide fixation using the Calvin cycle is generated by reverse electron flow, thereby<br />
placing a further metabolic burden on an already energy-poor process.<br />
[edit] Anammox<br />
Anammox stands for anaerobic ammonia oxidation and is a relatively recently (late<br />
1990’s) discovered process. It occurs in members of the Planctomycetes (e.g.<br />
Candidatus Brocadia anammoxidans) and involves the coupling of ammonia<br />
oxidation to nitrite reduction. As oxygen is not required for this process these<br />
organisms are strict anaerobes. Amazingly, hydrazine (N2H4-rocket fuel) is produced<br />
as an intermediate during anammox metabolism. To deal with the high toxicity of<br />
hydrazine, anammox bacteria contain an hydrazine-containing intracellular organelle<br />
called the anammoxasome surrounded by highly compact (and unusual) ladderane<br />
lipid membrane. These lipids are unique in nature, as is the use of hydrazine as a<br />
metabolic intermediate. Anammox organisms are autotrophs although the mechanism<br />
for carbon dioxide fixation is unclear. Because of this property, these organisms have<br />
been applied industrially to remove nitrogen in wastewater treatment processes.<br />
Anammox has also been shown have widespread occurrence in anaerobic aquatic<br />
systems and has been speculated to account for approximately 50% of nitrogen gas<br />
production in some marine environments.<br />
[edit] Phototrophy<br />
Many microbes are capable of using light as a source of energy (phototrophy). Of<br />
these, algae are particularly significant because they are oxygenic, using water as an<br />
electron donor for electron transfer during photosynthesis. [citation needed] Phototrophic<br />
bacteria are found in the phyla Cyanobacteria, Chlorobi, Proteobacteria, Chloroflexi<br />
and Firmicutes. [1] Along with plants these microbes are responsible for all biological<br />
generation of diatomic oxygen on Earth. Because chloroplasts were derived from a<br />
lineage of the Cyanobacteria, the general principles of metabolism in these<br />
endosymbionts can also be applied to chloroplasts. In addition to oxygenic<br />
photosynthesis, many bacteria can also photosynthesize anaerobically, typically using<br />
sulfide (H2S) as an electron donor to produce sulfate. Inorganic sulfur (S 0 ), thiosulfate<br />
(S2O3 2- ) and ferrous iron (Fe 2+ ) can also be used by some organisms.<br />
Phylogenetically, all oxygenic photosynthetic bacteria are Cyanobacteria, while<br />
anoxygenic photosynthetic bacteria belong to the purple bacteria (Proteobacteria),<br />
Green sulfur bacteria (e.g. Chlorobium), Green non-sulfur bacteria (e.g. Chloroflexus)<br />
or the heliobacteria (Low %G+C Gram positives). In addition to these organisms,<br />
some microbes (e.g. the archaeon Halobacterium or the bacterium Roseobacter,<br />
among others) can utilize light to produce energy using the enzyme<br />
bacteriorhodopsin, a light-driven proton pump. This type of metabolism is not
considered to be photosynthesis but rather photophosphorylation, since it generates<br />
energy, but does not directly fix carbon.<br />
As befits the large diversity of photosynthetic bacteria, there exist many different<br />
mechanisms by which light is converted into energy for metabolism. All<br />
photosynthetic organisms locate their photosynthetic reaction centers within a<br />
membrane, which may be invaginations of the cytoplasmic membrane (purple<br />
bacteria), thylakoid membranes (Cyanobacteria), specialized antenna structures called<br />
chlorosomes (Green sulfur and non-sulfur bacteria) or the cytoplasmic membrane<br />
itself (heliobacteria). Different photosynthetic bacteria also contain different<br />
photosynthetic pigments such as chlorophylls and carotenoids allowing them to take<br />
advantage of different portions of the electromagnetic spectrum and thereby inhabit<br />
different niches. Some groups of organisms contain more specialized light-harvesting<br />
structures e.g. phycobilisomes in Cyanobacteria and chlorosomes in Green sulfur and<br />
non-sulfur bacteria, allowing for increased light utilization efficiency.<br />
Biochemically, anoxygenic photosynthesis is very different from oxygenic<br />
photosynthesis. Cyanobacteria (and by extension chloroplasts) use the Z scheme of<br />
electron flow in which electrons eventually are used to form NADH. Two different<br />
reaction centers (photosystems) are used and proton motive force is generated both by<br />
using cyclic electron flow and the quinone pool. In anoxygenic photosynthetic<br />
bacteria electron flow is cyclic, with all electrons used in photosynthesis eventually<br />
being transferred back to the single reaction center. A proton motive force is<br />
generated using only the quinone pool. In heliobacteria, Green sulfur and non-sulfur<br />
bacteria NADH is formed using the protein ferredoxin, an energetically favorable<br />
reaction. In purple bacteria NADH is formed by reverse electron flow due to the<br />
lower chemical potential of this reaction centre. In all cases, however, a proton motive<br />
force is generated and used to drive ATP production via an ATPase.<br />
Most photosynthetic microbes are autotrophic, fixing carbon dioxide via the Calvin<br />
cycle. Some photosynthetic bacteria (e.g. Chloroflexus) are photoheterotrophs,<br />
meaning that they use organic carbon compounds as a carbon source for growth.<br />
Some photosynthetic organisms also fix nitrogen (see below).<br />
[edit] Nitrogen fixation<br />
Main article: Nitrogen fixation<br />
Nitrogen is an element required for growth by all biological systems. While extremely<br />
common (80% by volume) in the atmosphere, dinitrogen gas (N2) is generally<br />
biologically inaccessible due to its high activation energy. Throughout all of nature,<br />
only specialized bacteria are capable of nitrogen fixation, converting dinitrogen gas<br />
into ammonia (NH3), which is easily assimilated by all organisms. These bacteria,<br />
therefore are very important ecologically and are often essential for the survival entire<br />
ecosystems. This is especially true in the ocean, where nitrogen-fixing cyanobacteria<br />
are often the only sources or fixed nitrogen and in soils where specialized symbioses<br />
exist between legumes and their nitrogen-fixing partners to provide the nitrogen<br />
needed by these plants for growth.
Nitrogen fixation can be found distributed throughout nearly all bacterial lineages and<br />
physiological classes but is not a universal property. Because the enzyme nitrogenase,<br />
responsible for nitrogen fixation, is very sensitive to oxygen which will inhibit it<br />
irreversibly, all nitrogen-fixing organisms must possess some mechanism to keep the<br />
concentration of oxygen low. Examples include:<br />
• heterocyst formation (cyanobacteria e.g. Anabaena) where one cell does not<br />
photosynthesize but instead fixed nitrogen for its neighbors which in turn<br />
provide it with energy<br />
• root nodule symbioses (e.g. Rhizobium) with plants that supply oxygen to the<br />
bacteria bound to molecules of leghaemoglobin<br />
• anaerobic lifestyle (e.g. Clostridium pasteurianum)<br />
• very fast metabolism (e.g. Azotobacter vinelandii)<br />
The production and activity of nitrogenases is very highly regulated, both because<br />
nitrogen fixation is an extremely energetically expensive process (16-24 ATP are used<br />
per N2 fixed) and due to the extreme sensitivity of the nitrogenase to oxygen.<br />
[edit] See also<br />
• lipophilic bacteria, a minority of bacteria with lipid metabolism<br />
[edit] References<br />
1. ^ D.A. Bryant & N.-U. Frigaard (Nov 2006). "Prokaryotic photosynthesis and<br />
phototrophy illuminated". Trends Microbiol. 14 (11): 488.<br />
doi:doi:10.1016/j.tim.2006.09.001.<br />
• Madigan, M. T., Martinko, J. M. "Brock Biology of Microorganisms, 11th<br />
Ed." (2005) Pearson<br />
http://en.wikipedia.org/wiki/Category:Microbiology_techniques<br />
Category:Microbiology techniques<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
Pages in category "Microbiology techniques"<br />
There are 19 pages in this section of this category.<br />
A<br />
• Antibiogram<br />
C<br />
• Clonogenic assay<br />
M cont.<br />
• Microscopy
B<br />
• Aseptic technique<br />
• Auraminerhodamine<br />
stain<br />
• Axenic<br />
• Bacterial water<br />
analysis<br />
• Blood culture<br />
G<br />
I<br />
K<br />
M<br />
• Cryogenic grinding<br />
• Guthrie test<br />
• Indole test<br />
• Industrial fermentation<br />
• Kirby-Bauer antibiotic<br />
testing<br />
• Microbiological<br />
culture<br />
O<br />
R<br />
S<br />
Z<br />
• Miles and Misra<br />
method<br />
• Oxidase test<br />
• Replica plating<br />
• Streaking<br />
(microbiology)<br />
• Ziehl-Neelsen<br />
stain<br />
Retrieved from "http://en.wikipedia.org/wiki/Category:Microbiology_techniques"<br />
Categories: Microbiology | Laboratory techniques<br />
Microscopy mi·cros·co·py (Pronunciation[mahy-kros-kuh-pee, mahy-kruh-skoh-pee])<br />
is the technical field of using microscopes to view samples or objects. There are three<br />
well-known branches of microscopy, optical, electron and scanning probe<br />
microscopy.<br />
Optical and electron microscopy involve the diffraction, reflection, or refraction of<br />
electromagnetic radiation incident upon the subject of study, and the subsequent<br />
collection of this scattered radiation in order to build up an image. This process may<br />
be carried out by wide field irradiation of the sample (for example standard light<br />
microscopy and transmission electron microscopy) or by scanning of a fine beam over<br />
the sample (for example confocal microscopy and scanning electron microscopy).<br />
Scanning probe microscopy involves the interaction of a scanning probe with the<br />
surface or object of interest. The development of microscopy revolutionized biology<br />
and remains an essential tool in that science, along with many others.
Scanning electron microscope image of pollen.<br />
Contents<br />
[hide]<br />
• 1 Optical microscopy<br />
o 1.1 Limitations of optical microscopy<br />
o 1.2 Optical microscopy techniques<br />
1.2.1 Bright field optical microscopy and what it means<br />
1.2.2 Oblique illumination and what it means<br />
1.2.3 Dark field optical microscopy and what it means<br />
1.2.4 Phase contrast optical microscopy<br />
1.2.5 Differential interference contrast microscopy<br />
1.2.6 Fluorescence microscopy<br />
1.2.7 Confocal laser scanning microscopy<br />
1.2.8 Deconvolution microscopy<br />
o 1.3 Sub-diffraction optical microscopy techniques<br />
1.3.1 NSOM<br />
1.3.2 Local enhancement / ANSOM / bowties<br />
1.3.3 STED<br />
1.3.4 Fitting the PSF<br />
1.3.5 PALM & STORM<br />
1.3.6 Structured illumination<br />
o 1.4 Extensions of the optical microscope<br />
o 1.5 Other optical microscope enhancements<br />
o 1.6 X-ray microscopy<br />
o 1.7 Electron Microscopy<br />
o 1.8 Atomic de Broglie microscope<br />
• 2 Scanning probe microscopy<br />
o 2.1 Ultrasonic force microscopy<br />
• 3 Infrared microscopy<br />
• 4 Amateur Microscopy
• 5 See also<br />
• 6 References<br />
o 6.1 Further reading<br />
• 7 External links<br />
o 7.1 Organizations<br />
[edit] Optical microscopy<br />
See also: Optical microscope<br />
Optical or light microscopy involves passing visible light transmitted through or<br />
reflected from the sample through a single or multiple lenses to allow a magnified<br />
view of the sample. [1] The resulting image can be detected directly by the eye, imaged<br />
on a photographic plate or captured digitally. The single lens with its attachments, or<br />
the system of lenses and imaging equipment, along with the appropriate lighting<br />
equipment, sample stage and support, makes up the basic light microscope.<br />
[edit] Limitations of optical microscopy<br />
See also: Microscopy#Super-Resolution Optical Microscopy Techniques<br />
Limitations of standard optical microscopy (bright field microscopy) lie in three<br />
areas;<br />
• The technique can only image dark or strongly refracting objects effectively.<br />
• Diffraction limits resolution to approximately 0.2 micrometre (see:<br />
microscope).<br />
• Out of focus light from points outside the focal plane reduces image clarity.<br />
Live cells in particular generally lack sufficient contrast to be studied successfully,<br />
internal structures of the cell are colourless and transparent. The most common way to<br />
increase contrast is to stain the different structures with selective dyes, but this<br />
involves killing and fixing the sample. Staining may also introduce artifacts, apparent<br />
structural details that are caused by the processing of the specimen and are thus not a<br />
legitimate feature of the specimen.<br />
These limitations have, to some extent, all been overcome by specific microscopy<br />
techniques which can non-invasively increase the contrast of the image. In general,<br />
these techniques make use of differences in the refractive index of cell structures. It is<br />
comparable to looking through a glass window: you (bright field microscopy) don't<br />
see the glass but merely the dirt on the glass. There is however a difference as glass is<br />
a more dense material, and this creates a difference in phase of the light passing<br />
through. The human eye is not sensitive to this difference in phase but clever optical<br />
solutions have been thought out to change this difference in phase into a difference in<br />
amplitude (light intensity).<br />
[edit] Optical microscopy techniques
[edit] Bright field optical microscopy and what it means<br />
Main article: Bright field microscopy<br />
Bright field microscopy is the simplest of all the light microscopy techniques. Sample<br />
illumination is via transmitted white light, i.e. illuminated from below and observed<br />
from above. Limitations include low contrast of most biological samples and low<br />
apparent resolution due to the blur of out of focus material. The simplicity of the<br />
technique and the minimal sample preparation required are significant advantages.<br />
[edit] Oblique illumination and what it means<br />
The use of oblique (from the side) illumination gives the image a 3-dimensional<br />
appearance and can highlight otherwise invisible features. A more recent technique<br />
based on this method is Hoffmann's modulation contrast, a system found on inverted<br />
microscopes for use in cell culture. Oblique illumination suffers from the same<br />
limitations as bright field microscopy (low contrast of many biological samples; low<br />
apparent resolution due to out of focus objects), but may highlight otherwise invisible<br />
structures.<br />
[edit] Dark field optical microscopy and what it means<br />
Main article: Dark field microscopy<br />
Dark field microscopy is a technique for improving the contrast of unstained,<br />
transparent specimens. [2] Darkfield illumination uses a carefully aligned light source<br />
to minimise the quantity of directly-transmitted (un-scattered) light entering the image<br />
plane, collecting only the light scattered by the sample. Darkfield can dramatically<br />
improve image contrast—especially of transparent objects—while requiring little<br />
equipment setup or sample preparation. However, the technique does suffer from low<br />
light intensity in final image of many biological samples, and continues to be affected<br />
by low apparent resolution.<br />
Rheinberg illumination is a special variant of dark field illumination in which<br />
transparent, colored filters are inserted just before the condenser so that light rays at<br />
high aperture are differently colored than those at low aperture (i.e. the background to<br />
the specimen may be blue while the object appears self-luminous yellow). Other color<br />
combinations are possible but their effectiveness is quite variable. [3]<br />
[edit] Phase contrast optical microscopy<br />
Main articles: Phase contrast microscope and Phase contrast microscopy<br />
In electron microscopy: Phase-contrast imaging<br />
More sophisticated techniques will show differences in optical density in proportion.<br />
Phase contrast is a widely used technique that shows differences in refractive index<br />
as difference in contrast. It was developed by the Dutch physicist Frits Zernike in the<br />
1930s (for which he was awarded the Nobel Prize in 1953). The nucleus in a cell for<br />
example will show up darkly against the surrounding cytoplasm. Contrast is excellent;<br />
however it is not for use with thick objects. Frequently, a halo is formed even around<br />
small objects, which obscures detail. The system consists of a circular annulus in the<br />
condenser which produces a cone of light. This cone is superimposed on a similar
sized ring within the phase-objective. Every objective has a different size ring, so for<br />
every objective another condenser setting has to be chosen. The ring in the objective<br />
has special optical properties: it first of all reduces the direct light in intensity, but<br />
more importantly, it creates an artificial phase difference of about a quarter<br />
wavelength. As the physical properties of this direct light have changed, interference<br />
with the diffracted light occurs, resulting in the phase contrast image.<br />
[edit] Differential interference contrast microscopy<br />
Main article: Differential interference contrast microscopy<br />
Superior and much more expensive is the use of interference contrast. Differences in<br />
optical density will show up as differences in relief. A nucleus within a cell will<br />
actually show up as a globule in the most often used differential interference<br />
contrast system according to Georges Nomarski. However, it has to be kept in mind<br />
that this is an optical effect, and the relief does not necessarily resemble the true<br />
shape! Contrast is very good and the condenser aperture can be used fully open,<br />
thereby reducing the depth of field and maximizing resolution.<br />
The system consists of a special prism (Nomarski prism, Wollaston prism) in the<br />
condenser that splits light in an ordinary and an extraordinary beam. The spatial<br />
difference between the two beams is minimal (less than the maximum resolution of<br />
the objective). After passage through the specimen, the beams are reunited by a<br />
similar prism in the objective.<br />
In a homogeneous specimen, there is no difference between the two beams, and no<br />
contrast is being generated. However, near a refractive boundary (say a nucleus within<br />
the cytoplasm), the difference between the ordinary and the extraordinary beam will<br />
generate a relief in the image. Differential interference contrast requires a polarized<br />
light source to function; two polarizing filters have to be fitted in the light path, one<br />
below the condenser (the polarizer), and the other above the objective (the analyzer).<br />
Note: In cases where the optical design of a microscope produces an appreciable<br />
lateral separation of the two beams we have the case of classical interference<br />
microscopy, which does not result in relief images, but can nevertheless be used for<br />
the quantitative determination of mass-thicknesses of microscopic objects.<br />
[edit] Fluorescence microscopy<br />
Main article: Fluorescence microscopy<br />
When certain compounds are illuminated with high energy light, they then emit light<br />
of a different, lower frequency. This effect is known as fluorescence. Often specimens<br />
show their own characteristic autofluorescence image, based on their chemical<br />
makeup.<br />
This method is of critical importance in the modern life sciences, as it can be<br />
extremely sensitive, allowing the detection of single molecules. Many different<br />
fluorescent dyes can be used to stain different structures or chemical compounds. One<br />
particularly powerful method is the combination of antibodies coupled to a<br />
fluorochrome as in immunostaining. Examples of commonly used fluorochromes are
fluorescein or rhodamine. The antibodies can be made tailored specifically for a<br />
chemical compound. For example, one strategy often in use is the artificial production<br />
of proteins, based on the genetic code (DNA). These proteins can then be used to<br />
immunize rabbits, which then form antibodies which bind to the protein. The<br />
antibodies are then coupled chemically to a fluorochrome and then used to trace the<br />
proteins in the cells under study.<br />
Highly-efficient fluorescent proteins such as the green fluorescent protein (GFP) have<br />
been developed using the molecular biology technique of gene fusion, a process<br />
which links the expression of the fluorescent compound to that of the target<br />
protein.Piston DW, Patterson GH, Lippincott-Schwartz J, Claxton NS, Davidson MW<br />
(2007). Nikon MicroscopyU: Introduction to Fluorescent Proteins. Nikon<br />
MicroscopyU. Retrieved on 2007-08-22. This combined fluorescent protein is<br />
generally non-toxic to the organism and rarely interferes with the function of the<br />
protein under study. Genetically modified cells or organisms directly express the<br />
fluorescently-tagged proteins, which enables the study of the function of the original<br />
protein in vivo.<br />
Since fluorescence emission differs in wavelength (color) from the excitation light, a<br />
fluorescent image ideally only shows the structure of interest that was labelled with<br />
the fluorescent dye. This high specificity led to the widespread use of fluorescence<br />
light microscopy in biomedical research. Different fluorescent dyes can be used to<br />
stain different biological structures, which can then be detected simultaneously, while<br />
still being specific due to the individual color of the dye.<br />
To block the excitation light from reaching the observer or the detector, filter sets of<br />
high quality are needed. These typically consist of an excitation filter selecting the<br />
range of excitation wavelengths, a dichroic mirror, and an emission filter blocking the<br />
excitation light. Most fluorescence microscopes are operated in the Epi-illumination<br />
mode (illumination and detection from one side of the sample) to further decrease the<br />
amount of excitation light entering the detector.<br />
See also total internal reflection fluorescence microscope.<br />
[edit] Confocal laser scanning microscopy<br />
Main article: Confocal laser scanning microscopy<br />
Generates the image by a completely different way than the normal visual bright field<br />
microscope. It gives slightly higher resolution, but most importantly it provides<br />
optical sectioning without disturbing out-of-focus light degrading the image.<br />
Therefore it provides sharper images of 3D objects. This is often used in conjunction<br />
with fluorescence microscopy.<br />
[edit] Deconvolution microscopy<br />
Fluorescence microscopy is extremely powerful due to its ability to show specifically<br />
labelled structures within a complex environment but also because of its inherent<br />
ability to provide three dimensional information of biological structures.<br />
Unfortunately this information is blurred by the fact, that upon illumination all
fluorescently labeled structures emit light no matter if they are in focus or not. This<br />
means, that an image of a certain structure is always blurred by the contribution of<br />
light from structures which are out of focus. This phenomenon becomes apparent as a<br />
loss of contrast especially when using objectives with a high resolving power,<br />
typically oil immersion objectives with a high numerical aperture.<br />
Fortunately though, this phenomenon is not caused by random processes such as light<br />
scattering but can be relatively well defined by the optical properties of the image<br />
formation in the microscope imaging system. If one considers a small fluorescent<br />
light source (essentially a bright spot), light coming from this spot spreads out the<br />
further out of focus one is. Under ideal conditions this produces a sort of "hourglass"<br />
shape of this point source in the third (axial) dimension. This shape is called the point<br />
spread function (PSF) of the microscope imaging system. Since any fluorescence<br />
image is made up of a large number of such small fluorescent light sources the image<br />
is said to be "convolved by the point spread function".<br />
Knowing this point spread function means, that it is possible to reverse this process to<br />
a certain extent by computer based methods commonly known as deconvolution<br />
microscopy. [4] There are various algorithms available for 2D or 3D Deconvolution.<br />
They can be roughly classified in non restorative and restorative methods. While the<br />
non restorative methods can improve contrast by removing out of focus light from<br />
focal planes, only the restorative methods can actually reassign light to it proper place<br />
of origin. This can be an advantage over other types of 3D microscopy such as<br />
confocal microscopy, because light is not thrown away but reused. For 3D<br />
deconvolution one typically provides a series of images derived from different focal<br />
planes (called a Z-stack) plus the knowledge of the PSF which can be either derived<br />
experimentally or theoretically from knowing all contributing parameters of the<br />
microscope.<br />
[edit] Sub-diffraction optical microscopy techniques<br />
It is well known that there is a spatial limit to which light can focus: approximately<br />
half of the wavelength of the light you are using. But this is not a true barrier, because<br />
this diffraction limit is only true in the far-field and localization precision can be<br />
increased with many photons and careful analysis (although two objects still cannot<br />
be resolved); and like the sound barrier, the diffraction barrier is breakable. This<br />
section explores some approaches to imaging objects smaller than ~250 nm. Most of<br />
the following information was gathered (with permission) from a chemistry blog's<br />
review of sub-diffraction microscopy techniques Part I and Part II. For a review, see<br />
also reference [5] .<br />
[edit] NSOM<br />
Probably the most conceptual way to break the diffraction barrier is to use a light<br />
source and/or a detector that is itself nanometer in scale. Diffraction as we know it is<br />
truly a far-field effect: the light from an aperture is the Fourier transform of the<br />
aperture in the far-field. [6] But in the near-field, all of this is not necessarily the case.<br />
Near-field scanning optical microscopy (NSOM) forces light through the tiny tip of a<br />
pulled fiber—and the aperture can be on the order of tens of nanometers. [7] When the<br />
tip is brought to nanometers away from a molecule, the resolution is not limited by
diffraction but by the size of the tip aperture (because only that one molecule will see<br />
the light coming out of the tip). An image can be built by a raster scan of the tip over<br />
the surface to create an image.<br />
The main down-side to NSOM is the limited number of photons you can force out a<br />
tiny tip, and the minuscule collection efficiency (if you are trying to collect<br />
fluorescence in the near-field). Other techniques such as ANSOM (see below) try to<br />
avoid this drawback.<br />
[edit] Local enhancement / ANSOM / bowties<br />
Instead of forcing photons down a tiny tip, some techniques create a local bright spot<br />
in an otherwise diffraction-limited spot. ANSOM is apertureless NSOM: it uses a tip<br />
very close to a fluorophore to enhance the local electric field the fluorophore sees. [8]<br />
Basically, the ANSOM tip is like a lightning rod which creates a hot spot of light.<br />
Bowtie nanoantennas have been used to greatly and reproducibly enhance the electric<br />
field in the nanometer gap between the tips two gold triangles. Again, the point is to<br />
enhance a very small region of a diffraction-limited spot, thus improving the<br />
mismatch between light and nanoscale objects—and breaking the diffraction barrier. [9]<br />
[edit] STED<br />
A recent favorite is STED—stimulated emission depletion. Stefan Hell at the Max<br />
Planck Institute developed this method, which uses two laser pulses. The first pulse is<br />
a diffraction-limited spot that is tuned to the absorption wavelength, so excites any<br />
fluorophores in that region; an immediate second pulse is red-shifted to the emission<br />
wavelength and stimulates emission back to the ground state before, thus depleting<br />
the excited state of any fluorophores in this depletion pulse. The trick is that the<br />
depletion pulse goes through a phase modulator that makes the pulse illuminate the<br />
sample in the shape of a donut, so the outer part of the diffraction limited spot is<br />
depleted and the small center can still fluoresce. By saturating the depletion pulse, the<br />
center of the donut gets smaller and smaller until they can get resolution of tens of<br />
nanometers. [10]<br />
This technique also requires a raster scan like NSOM and standard confocal laser<br />
scanning microscopy.<br />
[edit] Fitting the PSF<br />
The methods above (and below) use experimental techniques to circumvent the<br />
diffraction barrier, but one can also use crafty analysis to increase the ability to know<br />
where a nanoscale object is located. The image of a point source on a charge-coupled<br />
device camera is called a point-spread function (PSF), which is limited by diffraction<br />
to be no less than approximately half the wavelength of the light. But it is possible to<br />
simply fit that PSF with a Gaussian to locate the center of the PSF—and thus the<br />
location of the fluorophore. The precision by which this technique can locate the<br />
center depends on the number of photons collected (as well as the CCD pixel size and<br />
other factors). [11] Regardless, groups like the Selvin lab and many others have
employed this analysis to localize single fluorophores to a few nanometers. This, of<br />
course, requires careful measurements and collecting many photons.<br />
[edit] PALM & STORM<br />
What fitting a PSF is to localization, photo-activated localization microscopy (PALM)<br />
is to "resolution"—this term is here used loosely to mean measuring the distance<br />
between objects, not true optical resolution. Eric Betzig and colleagues developed<br />
PALM; [12] Xiaowei Zhuang at Harvard used a similar techniques and calls it STORM:<br />
stochastic optical reconstruction microscopy. [13] The basic premise of both techniques<br />
is to fill the imaging area with many dark fluorophores that can be photoactivated into<br />
a fluorescing state by a flash of light. Because photoactivation is stochastic, only a<br />
few, well separated molecules "turn on." Then Gaussians are fit to their PSFs to high<br />
precision (see section above). After the few bright dots photobleach, another flash of<br />
the photoactivating light activates random fluorophores again and the PSFs are fit of<br />
these different well spaced objects. This process is repeated many times, building up<br />
an image molecule-by-molecule; and because the molecules were localized at<br />
different times, the "resolution" of the final image can be much higher than that<br />
limited by diffraction.<br />
The major problem with these techniques is that to get these beautiful pictures, it<br />
takes on the order of hours to collect the data. This is certainly not the technique to<br />
study dynamics (fitting the PSF is better for that).<br />
[edit] Structured illumination<br />
There is also the wide-field structured-illumination (SI) approach to breaking the<br />
diffraction limit of light. [14][15] SI—or patterned illumination—relies on both specific<br />
microscopy protocols and extensive software analysis post-exposure. But, because SI<br />
is a wide-field technique, it is usually able to capture images at a higher rate than<br />
confocal-based schemes like STED. (This is only a generalization, because SI isn't<br />
actually super fast. I'm sure someone could make STED fast and SI slow!) The main<br />
concept of SI is to illuminate a sample with patterned light and increase the resolution<br />
by measuring the fringes in the Moiré pattern (from the interference of the<br />
illumination pattern and the sample). "Otherwise-unobservable sample information<br />
can be deduced from the fringes and computationally restored." [16]<br />
SI enhances spatial resolution by collecting information from frequency space outside<br />
the observable region. This process is done in reciprocal space: the Fourier transform<br />
(FT) of an SI image contains superimposed additional information from different<br />
areas of reciprocal space; with several frames with the illumination shifted by some<br />
phase, it is possible to computationally separate and reconstruct the FT image, which<br />
has much more resolution information. The reverse FT returns the reconstructed<br />
image to a super-resolution image.<br />
But this only enhances the resolution by a factor of 2 (because the SI pattern cannot<br />
be focused to anything smaller than half the wavelength of the excitation light). To<br />
further increase the resolution, you can introduce nonlinearities, which show up as<br />
higher-order harmonics in the FT. In reference [16] , Gustafsson uses saturation of the<br />
fluorescent sample as the nonlinear effect. A sinusoidal saturating excitation beam
produces the distorted fluorescence intensity pattern in the emission. This<br />
nonpolynomial nonlinearity yields a series of higher-order harmonics in the FT.<br />
Each higher-order harmonic in the FT allows another set of images that can be used to<br />
reconstruct a larger area in reciprocal space, and thus a higher resolution. In this case,<br />
Gustafsson achieves less than 50-nm resolving power, more than five times that of the<br />
microscope in its normal configuration.<br />
The main problems with SI are that, in this incarnation, saturating excitation powers<br />
cause more photodamage and lower fluorophore photostability, and sample drift must<br />
be kept to below the resolving distance. The former limitation might be solved by<br />
using a different nonlinearity (such as stimulated emission depletion or reversible<br />
photoactivation, both of which are used in other sub-diffraction imaging schemes); the<br />
latter limits live-cell imaging and may require faster frame rates or the use of some<br />
fiducial markers for drift subtraction. Nevertheless, SI is certainly a strong contender<br />
for further application in the field of super-resolution microscopy.<br />
[edit] Extensions of the optical microscope<br />
Most modern instruments provide simple solutions for micro-photography and image<br />
recording electronically. However such capabilities are not always present and the<br />
more experienced microscopist will, in many cases, still prefer a hand drawn image<br />
rather than a photograph. This is because a microscopist with knowledge of the<br />
subject can accurately convert a three dimensional image into a precise two<br />
dimensional drawing . In a photograph or other image capture system however, only<br />
one thin plane is ever in good focus.<br />
The creation of careful and accurate micrographs requires a microscopical technique<br />
using a monocular eyepiece. It is essential that both eyes are open and that the eye<br />
that is not observing down the microscope is instead concentrated on a sheet of paper<br />
on the bench besides the microscope. With practice, and without moving the head or<br />
eyes, it is possible to accurately record the observed details by tracing round the<br />
observed shapes by simultaneously "seeing" the pencil point in the microscopical<br />
image.<br />
Practising this technique also establishes good general microscopical technique. It is<br />
always less tiring to observe with the microscope focussed so that the image is seen at<br />
infinity and with both eyes open at all times.<br />
[edit] Other optical microscope enhancements<br />
stereomicroscope<br />
[edit] X-ray microscopy<br />
Main article: X-ray microscopy<br />
As resolution depends on the wavelength of the light. Electron microscopy has been<br />
developed since the 1930s that use electron beams instead of light. Because of the<br />
much lower wavelength of the electron beam, resolution is far higher.
Though less common, X-ray microscopy has also been developed since the late<br />
1940s. The resolution of X-ray microscopy lies between that of light microscopy and<br />
the electron microscopy.<br />
[edit] Electron Microscopy<br />
For light microscopy the wavelength of the light limits the resolution to around 0.2<br />
micrometers. In order to gain higher resolution, the use of an electron beam with a far<br />
smaller wavelength is used in electron microscopes.<br />
• Transmission electron microscopy (TEM) is principally quite similar to the<br />
compound light microscope, by sending an electron beam through a very thin<br />
slice of the specimen. The resolution limit nowadays (2005) is around 0.05<br />
nanometer.<br />
• Scanning electron microscopy (SEM) visualizes details on the surfaces of cells<br />
and particles and gives a very nice 3D view. It gives results much like the<br />
stereo light microscope and akin to that its most useful magnification is in the<br />
lower range than that of the transmission electron microscope.<br />
[edit] Atomic de Broglie microscope<br />
Main article: Atomic de Broglie microscope<br />
The atomic de Broglie microscope is an imaging system which is expected to provide<br />
resolution at the nanometer scale using neutral He atoms as probe particles. [17][18] .<br />
Such a device could provide the resolution at nanometer scale and be absolutely nondestructive,<br />
but it is not developed so well as optical microscope or an electron<br />
microscope.<br />
[edit] Scanning probe microscopy<br />
This is a sub-diffraction technique. Examples of scanning probe microscopes are the<br />
atomic force microscope (AFM), the Scanning tunneling microscope and the photonic<br />
force microscope. All such methods imply a solid probe tip in the vicinity (near field)<br />
of an object, which is supposed to be almost flat. For more detail, see Scanning probe<br />
microscopy.<br />
[edit] Ultrasonic force microscopy<br />
Ultrasonic Force Microscopy (UFM) has been developed in order to improve the<br />
details and image contrast on "flat" areas of interest where the AFM images are<br />
limited in contrast. The combination of AFM-UFM allows a near field acoustic<br />
microscopic image to be generated. The AFM tip is used to detect the ultrasonic<br />
waves and overcomes the limitation of wavelength that occurs in acoustic<br />
microscopy. By using the elastic changes under the AFM tip, an image of much<br />
greater detail than the AFM topography can be generated.<br />
Ultrasonic force microscopy allows the local mapping of elasticity in atomic force<br />
microscopy by the application of ultrasonic vibration to the cantilever or sample. In an
attempt to analyse the results of ultrasonic force microscopy in a quantitative fashion,<br />
a force-distance curve measurement is done with ultrasonic vibration applied to the<br />
cantilever base, and the results are compared with a model of the cantilever dynamics<br />
and tip-sample interaction based on the finite-difference technique.<br />
[edit] Infrared microscopy<br />
The term infrared microscope covers two main types of diffraction-limited<br />
microscopy. The first provides optical visualisation plus IR spectroscopic data<br />
collection. The second (more recent and more advanced) technique employs focal<br />
plane array detection for infrared chemical imaging, where the image contrast is<br />
determined by the response of individual sample regions to particular IR wavelengths<br />
selected by the user.<br />
IR versions of sub-diffraction microscopy (see above) exist also. These include IR<br />
NSOM [19] and photothermal microspectroscopy.<br />
[edit] Amateur Microscopy<br />
Amateur Microscopy is the investigation and observation of biological and nonbiological<br />
specimens for recreational purposes using an optical microscope (light<br />
microscopes). Collectors of minerals, insects, seashells and plants may use<br />
microscopes as tools to uncover features that help them classify their collected items.<br />
Other amateurs may be interested in observing the life found in pond water and of<br />
other samples. Microscopes may also prove useful for the water quality assessment<br />
for people that keep a home aquarium. Photographic documentation and drawing of<br />
the microscopic images are additional tasks that augment the spectrum of tasks of the<br />
amateur. There are even competitions for photomicrograph art. Participants of this<br />
pastime may either use commercially prepared microscopic slides or may engage in<br />
the task of specimen preparation.<br />
While microscopy is a central tool in the documentation of biological specimens, it is<br />
rarely sufficient to justify the discovery of a new species based on microscopic<br />
investigations alone. Often genetic and biochemical tests are necessary to confirm the<br />
discovery of a new species. A fully equipped laboratory may be necessary, something<br />
often not available to amateurs. For this reason it may be unlikely that amateur<br />
microscopists are capable of substantiating their find to the extent to yield a scientific<br />
publication.<br />
In the late 1800's amateur microscopy became a popular hobby in the United States<br />
and Europe. Professor John Phin published "Practical Hints on the Selection and Use<br />
of the Microscope (Second Edition, 1878)," and was also the editor of the “American<br />
Journal of Microscopy.”<br />
[edit] See also<br />
• Köhler illumination<br />
• Two-photon excitation microscopy
[edit] References<br />
1. ^ Abramowitz M, Davidson MW (2007). Introduction to Microscopy. Molecular<br />
Expressions. Retrieved on 2007-08-22.<br />
2. ^ Abramowitz M, Davidson MW (2007). Darkfield Illumination. Retrieved on 2007-<br />
08-22.<br />
3. ^ Abramowitz M, Davidson MW (2007). Rheinberg Illumination. Retrieved on 2007-<br />
08-22.<br />
4. ^ Wallace W, Schaefer LH, Swedlow JR (2001). "A workingperson's guide to<br />
deconvolution in light microscopy". BioTechniques 31 (5): 1076-8, 1080, 1082<br />
passim. PMID 11730015.<br />
5. ^ WEM News and Views<br />
6. ^ Fresnel Diffraction Applet (Java applet). Retrieved on 2007-08-22.<br />
7. ^ Cummings JR, Fellers TJ, Davidson MW (2007). Specialized Microscopy<br />
Techniques - Near-Field Scanning Optical Microscopy. Olympus Microscopy<br />
Resource Center. Retrieved on 2007-08-22.<br />
8. ^ Sánchez EJ, Novotny L, Xie XS (1999). "Near-Field Fluorescence Microscopy<br />
Based on Two-Photon Excitation with Metal Tips". Phys Rev Lett 82: 4014-7.<br />
doi:10.1103/PhysRevLett.82.4014.<br />
9. ^ Schuck PJ, Fromm DP, Sundaramurthy A, Kino GS, Moerner WE (2005).<br />
"Improving the Mismatch between Light and Nanoscale Objects with Gold Bowtie<br />
Nanoantennas". Phys Rev Lett 94: 017402. doi:10.1103/PhysRevLett.94.017402.<br />
10. ^ STED<br />
11. ^ Webb paper<br />
12. ^ PALM<br />
13. ^ STORM<br />
14. ^ Bailey, B.; Farkas, D. L.; Taylor, D. L.; Lanni, F. Enhancement of axial resolution<br />
in fluorescence microscopy by standing-wave excitation. Nature 1993, 366, 44–48.<br />
15. ^ Gustafsson, M. G. L. Surpassing the lateral resolution limit by a factor of two using<br />
structured illumination microscopy. J. of Microsc. 2000, 198(2), 82–87.<br />
16. ^ a b Gustafsson, M. G. L. http://dx.doi.org/10.1073/pnas.0406877102 Nonlinear<br />
structured-illumination microscopy: Wide-field fluorescence imaging with<br />
theoretically unlimited resolution. PNAS 2005, 102(37), 13081–13086.<br />
17. ^ D.Kouznetsov; H. Oberst, K. Shimizu, A. Neumann, Y. Kuznetsova, J.-F. Bisson,<br />
K. Ueda, S. R. J. Brueck (2006). "Ridged atomic mirrors and atomic nanoscope".<br />
JOPB 39: 1605-1623.<br />
18. ^ Atom Optics and Helium Atom Microscopy. Cambridge University, http://wwwsp.phy.cam.ac.uk/research/mirror.php3<br />
19. ^ H M Pollock and D A Smith, The use of near-field probes for vibrational<br />
spectroscopy and photothermal imaging, in Handbook of vibrational spectroscopy,<br />
J.M. Chalmers and P.R. Griffiths (eds), John Wiley & Sons Ltd, Vol. 2, pp. 1472 -<br />
1492 (2002)<br />
[edit] Further reading<br />
• Advanced Light Microscopy vol. 1 Principles and Basic Properties by Maksymilian<br />
Pluta, Elsevier (1988)<br />
• Advanced Light Microscopy vol. 2 Specialised Methods by Maksymilian Pluta,<br />
Elsevier (1989)<br />
• Introduction to Light Microscopy by S. Bradbury, B. Bracegirdle, BIOS Scientific<br />
Publishers (1998)<br />
• Video Microscopy by Shinya Inoue, Plenum Press (1986)
• A review of sub-diffraction microscopy techniques Part I and Part II - a blog post<br />
with helpful information, some of which appears in this article<br />
[edit] External links<br />
• Microscopy Techniques Various Techniques Used In Microscopy<br />
• Carl Zeiss "Microscopy from the very beginning", a step by step tutorial into<br />
the basics of microscopy.<br />
• Interactive Fluorescence Dye and Filter Database Carl Zeiss Interactive<br />
Fluorescence Dye and Filter Database.<br />
• Nikon MicroscopyU - The source for microscopy education<br />
• Olympus Microscopy Resource Center<br />
• Microscopy in Detail - A resource with many illustrations elaborating the most<br />
common microscopy techniques<br />
• Images formed by simple microscopes - examples of observations with singlelens<br />
microscopes.<br />
• Portraits of life, one molecule at a time, a feature article on sub-diffraction<br />
microscopy from the March 1, 2007 issue of Analytical Chemistry<br />
[edit] Organizations<br />
v • d • e<br />
• Royal Microscopical Society (RMS)<br />
• Microscopy Society of America (MSA)<br />
• European Microscopy Society (EMS)<br />
Instrumentation<br />
Techniques<br />
Sampling<br />
Prominent<br />
publications<br />
[hide]<br />
Analytical chemistry<br />
Atomic absorption spectrometer · Flame emmission spectrometer ·<br />
Gas chromatograph · High performance liquid chromatograph ·<br />
Infrared Spectrometer · Mass spectrometer · Melting point apparatus ·<br />
Microscope · Spectrometer · Spectrophotometer<br />
Calorimetry · Chemometrics · Chromatography · Electrochemistry ·<br />
Gravimetric analysis<br />
Coning and quartering · Dilution · Dissolution · Filtration · Masking ·<br />
Pulverization · Sample preparation · Separation process · Subsampling<br />
Analytical chemistry<br />
http://mycology.cornell.edu/fteach.html<br />
Fungi Perfecti (Olympia, Washington, USA) supplies a plethora of mushroomgrowing<br />
equipment, spawn and kits, books, and dried edible and medicinal<br />
mushrooms. Their online catalog and information about Paul Stamets' mushroom
cultivation seminars and consultation services can be found here. This elegant web<br />
site includes many impressive images of mushrooms and other products, including<br />
scanning electron micrographs of mushroom ultrastructure.<br />
George Barron's website<br />
This website includes some lovely images of fungi, including Entomophthora,<br />
Spinellus, and some nematode parasites. It also includes information on Barron's book<br />
"Mushrooms of Northeast North America" (in Canada entitled "Mushrooms of<br />
Ontario and Eastern Canada").<br />
Glossary of Technical Terms in Plant Pathology<br />
This useful Glossary of technical terms in Plant Pathology was created by Phil<br />
Arneson of Cornell University. It includes definitions, illustrations, and sound files by<br />
Richard Korf to aid pronunciation.<br />
Irish Potato Famine<br />
A compilation of information on the Irish Potato Famine of the 1840s, during which<br />
time over 3 million Irish died, and many others (including some of my own ancestors)<br />
emigrated to other parts of the world. The Famine resulted from an outbreak of late<br />
blight, caused by Phytophthora infestans.<br />
John C. Tacoma Mushroom Slide Collection<br />
Many, many scanned images of mushrooms and allies, from photographs taken by<br />
John C. Tacoma, 1968-1978. Maintained by the Library of Indiana University-Purdue<br />
University Indianapolis.<br />
LichenLand<br />
Lichenland provides a fine introduction to lichens for both professionals and<br />
amateurs. Synoptic keys to taxa and to terms lead to many fine images of lichens, a<br />
compilation of their characteristics, and pertinent literature.<br />
Meredith Blackwell's Lab<br />
Meredith Blackwell's lab at Louisiana State University provides information on<br />
current research on insect-fungus associations, history of mycology, the genealogy of<br />
American mycologists, teaching resources, the LSU herbarium, and other tidbits.<br />
Microfungal home page<br />
Color images of many microfungi taken under the microscope. Over 100 genera of<br />
molds are represented.
Moulds: their isolation, cultivation and identification<br />
An online version of David Malloch's excellent guide to moulds (University of<br />
Toronto Press, 1981), complete with keys, media recipes, and illustrations of common<br />
genera. This book makes a great introduction to hyphomycetes for those with access<br />
to a microscope.<br />
Mushroom Toxins<br />
This discussion of mushroom toxins and the symptoms they produce forms a chapter<br />
of the "Bad Bug Book" by the US Food and Drug Administration. Other mycotoxins<br />
(aflatoxin and ilk) are discussed in a subsequent chapter.<br />
Mushrooms and Magic<br />
The Mycotheology Home Page provides an interesting discussion of the role of fungi<br />
in magic, folklore, and religion.<br />
Mushrooms of North Carolina<br />
Mycology students at Duke University (NC, USA) have prepared this site<br />
documenting the mushrooms of North Carolina. Their excellent photographs are<br />
available here.<br />
Mycologue Publications<br />
Mycologue is a publishing company founded by W. Bryce Kendrick. It provides<br />
books, teaching materials, and computerized keys to fungi (Canada). The site also<br />
includes information and many illustrations of fungi that complement Dr. Kendrick's<br />
textbook, The Fifth Kingdom (q.v.).<br />
Mycology class at Arizona State University<br />
Home page of the General Mycology class at Arizona State University, USA.<br />
Mycology class at Oregon State University<br />
Home page of the mycology class at Oregon State University, USA.<br />
Mycology class at Towson University<br />
The home page of the Mycology class at Towson University, in Maryland, USA.<br />
Mycology classes at Humboldt University<br />
Home page of Mycology classes at Humboldt University, California, USA.
Mycology Course at the University of Illinois at Urbana/Champaign<br />
This web site for Dr. Carol Shearer's Mycology class includes a syllabus, lab<br />
exercises, and many excellent lecture illustrations.<br />
Mycology Online<br />
Mycology Online is a guide to fungal pathogens of humans, the diseases they cause,<br />
and selected case studies. This Australian site is searchable, nicely illustrated (not for<br />
the squeamish!), and replete with information.<br />
Mycorrhiza information exchange<br />
The Mycorrhiza Information Exchange covers everything you need: literature<br />
databases, job ads, teaching tips, images, inoculum sources, links, etc. Participation is<br />
invited.<br />
Mycorrhizae and Plant Phylogeny<br />
A website devoted to mycorrhizae and plant systematics, and the evolution of<br />
mycorrhizal symbiosis.<br />
Mycorrhizas webpage<br />
This guide to mycorrhizal associations (adapted and excerpted from a larger book) is<br />
provided by Mark Brundett at CSIRO (Australia). It details the structure and<br />
development of mycorrhizae, with handsome images and good textual explanation. It<br />
makes a wonderful teaching tool.<br />
Mycorrhizospheres of boreal forest trees<br />
This site from the Biocenter at the University of Helsinki (Finland) includes scientific<br />
publications documenting the diversity, interactions and functions of forest tree<br />
mycorrhizae.<br />
Mycotoxin homepage<br />
A unit of the US Dept. of Agriculture, Agricultural Research Service that focuses on<br />
mycotoxin research. 3-dimensional molecular structures of a few mycotoxins<br />
produced by molds are available here.<br />
MyxoWeb<br />
This web site devoted to myxomycetes provides information on the plasmodial slime<br />
molds, including some impressively gooey images.
Natural Perspective's introduction to fungi<br />
Natural Perspective's nicely illustrated introduction to the fungal kingdom.<br />
North American Lichen Project<br />
The North American Lichen Project includes essays on lichen biology and the uses of<br />
lichens by people and animals, as well as excerpts and lovely photographs from the<br />
forthcoming book Lichens of North America, by I.M. Brodo, S.D. Sharnoff, and S.<br />
Sharnoff (Yale University Press).<br />
North American Mycological Association<br />
NAMA is a great group for amateur mycologists. It provides a national mushroom<br />
poisoning registry, sponsors an annual foray, and publishes a fine annual journal,<br />
McIlvainea, and a bimonthly newsletter, The Mycophile. Also available through<br />
NAMA are suggestions for teaching K-12 students about fungi, and other tidbits.<br />
Penn State Mushroom Spawn Lab<br />
Pennsylvania State University's strong program in mushroom cultivation presents fact<br />
sheets and other information about commercial mushroom production on these pages.<br />
PSU's mushroom growers' information pages are part of this site.<br />
Plant Pathogenic Fungi<br />
The University of Kentucky's course in plant pathogenic fungi has web pages that<br />
include the syllabus and other information.<br />
Plant Pathology<br />
The Plant Pathology courses at the University of Nebraska-Lincoln (USA). Most<br />
materials are for registered students only; a distance learning course is offered.<br />
Plant Pathology Internet Guidebook<br />
The Plant Pathology Internet Guidebook is a comprehensive source for Plant<br />
Pathology resources online. It is available through the Institute of Plant Diseases and<br />
Plant Protection in Hannover, Germany.<br />
Plant Pathology Simulations<br />
Computer simulations for teaching aspects of plant pathology and epidemiology.
Plasmodiophorid Home Page<br />
These are pages devoted to the Plasmodiophorales that include information about life<br />
histories, cytology, and biology of this interesting group of fungus-like protists. The<br />
site is no longer being updated.<br />
Pythium insidiosum<br />
Pythiosis is a disease of humans and animals that can be caused by the subject of this<br />
web page, Pythium insidiosum. The site includes graphic images and information on<br />
biology, epidemiology, diagnosis, and treatment.<br />
Spongospora Homepage<br />
Spongospora subterranea is a plasmodiophorid pathogen of potatoes (and other plants)<br />
and an emerging pathogen in some regions. This workshop site introduces the biology<br />
and control of S. subterranea and related species, and includes images and a<br />
discussion board.<br />
The Fifth Kingdom<br />
W.B. Kendrick's delightful introductory mycology textbook, The Fifth Kingdom, is<br />
partly available online. This site includes over 800 lavish, colorful illustrations as a<br />
supplement to the text, which is available from Mycologue Publications (q.v.). The<br />
text of sample chapters is available, too. Dr. Kendrick's website also includes other<br />
publications for sale.<br />
The Rhynie Chert and its Flora<br />
The Rhynie Chert is a fossilized Devonian lake shore in Scotland that includes some<br />
of the oldest fossils of plants and their associated fungi. This nice site introduces the<br />
botanical and mycological finds of the Rhynie Chert, and provides photos of the<br />
oldest known lichen and early arbuscular mycorrhizal fungi.<br />
This is Not Just Plant Pathogenic Fungi!<br />
Students at Texas A M University have prepared a guide to plant pathogenic (and<br />
other) fungi.<br />
Tom Volk's web pages<br />
One stop shopping for mycology. These pages feature a "fungus of the month"<br />
column, with entertaining text and nice photos, in addition to a plethora of other<br />
information about fungi. Tom is a professor at the University of Wisconsin-La Crosse,<br />
USA.
Tree of Life<br />
This phylogenetic navigator provides a tree that shows the evolutionary relationships<br />
of living organisms, including fungi. It also supplies descriptive pages on selected<br />
terminal taxa. Like biological systematics itself, it's a work in progress.<br />
UC Berkeley's Introduction to Fungi<br />
The Museum of Paleontology at the University of California, Berkeley provides a<br />
well-prepared introduction to the kingdom Fungi, and also to two groups that have<br />
historically been studied by mycologists, the Oomycota and slime molds. Similar<br />
introductions are available for all other taxa. This link makes a valuable addition to<br />
any teaching program.<br />
University of Tennessee Mycology Labs<br />
Drs. Ron Petersen and Karen Hughes maintain a nice set of web pages that include a<br />
primer on Botanical Nomenclature, a synopsis of molecular phylogenetic techniques.<br />
These pages also provide an important resources on color standards used by<br />
mycologists: a synopsis of Fries' color terminology, and a concordance of colors in<br />
the Ridgway and Methuen color handbooks. Lots of information is also provided on<br />
the projects of staff and students.<br />
Views of the Famine<br />
An illustrated history of news coverage of the Irish Potato Famine that occurred in the<br />
1840s due to Phytophthora infestans, causal agent of late blight of potato.<br />
Wayne's Word on the fungal kingdom<br />
A delightful introduction to selected members of the kingdom Fungi from the e-zine,<br />
Wayne's Word.<br />
Western Montana Mycological Association (USA): Fungal Jungal<br />
The Western Montana Mycological Association maintains this nice site. It includes<br />
photos of Montana mushrooms, recipes, an oyster mushroom cultivation project, a<br />
mushroom "trunk" for teachers, a morel information site, and information on the<br />
WMMA's current activities.<br />
World-Wide Web Virtual Library<br />
You're there now! This is a distributed library of resources maintained at many<br />
different sites all over the world. Unlike some of the big search engines, VL site<br />
maintainers personally select and evaluate the links they recommend, with the result<br />
that VL sites generally have a high signal to noise ratio. The WWW VL is a good<br />
place to start when looking for electronic information on all kinds of different topics.
HOME -- ABOUT -- COLLECTIONS -- DIRECTORIES -- DISCUSSIONS -- GENERAL -- GENETICS -- GUIDES -- MUSHROOMS<br />
-- SUPPLIES -- TAXONOMY -- TEACHING -- INDEX<br />
Robert Koch<br />
From Wikipedia, the free encyclopedia<br />
Jump to: navigation, search<br />
Robert Koch<br />
Born<br />
Died<br />
Robert Koch<br />
December 11, 1843<br />
Clausthal, Hanover<br />
May 27, 1910 (aged 66)<br />
Baden-Baden, Germany<br />
Field Microbiology<br />
Institutions Imperial Health Office, Berlin, University of Berlin<br />
Alma mater University of Göttingen<br />
Academic advisor Friedrich Gustav Jakob Henle<br />
Known for<br />
Co-founder of bacteriology,<br />
Koch's postulates of germ theory,<br />
Isolator of anthrax, tuberculosis and cholera<br />
Notable awards Nobel Prize in Medicine, 1905<br />
For the American lobbyist, see Bobby Koch.<br />
Robert Koch (December 11, 1843 – May 27, 1910) was a German physician. He<br />
became famous for isolating Bacillus anthracis (1877), the tuberculosis bacillus<br />
(1882) and the cholera vibrio (1883) and for his development of Koch's postulates.
He was awarded the Nobel Prize in Physiology or Medicine for his tuberculosis<br />
findings in 1905. He is considered one of the founders of microbiology - he inspired<br />
such major figures as Paul Ehrlich and Gerhard Domagk.<br />
Contents<br />
[hide]<br />
• 1 Biography<br />
• 2 References<br />
• 3 Consult<br />
• 4 See also<br />
• 5 External links<br />
[edit] Biography<br />
Robert Koch was born in Clausthal, Germany as the son of a mining official. He<br />
studied medicine under Friedrich Gustav Jakob Henle at the University of Göttingen<br />
and graduated in 1866. He then served in the Franco-Prussian War and later became<br />
district medical officer in Wollstein (now Wolsztyn, Poland). Working with very<br />
limited resources, he became one of the founders of bacteriology, the other major<br />
figure being Louis Pasteur.<br />
After Casimir Davaine showed the direct transmission of the anthrax bacillus between<br />
cows, Koch studied anthrax more closely. He invented methods to purify the bacillus<br />
from blood samples and grow pure cultures. He found that, while it could not survive<br />
outside a host for long, anthrax built persisting endospores that could last a long time.<br />
These endospores, embedded in soil, were the cause of unexplained "spontaneous"<br />
outbreaks of anthrax. Koch published his findings in 1876, and was rewarded with a<br />
job at the Imperial Health Office in Berlin in 1880. In 1881, he urged the sterilization<br />
of surgical instruments using heat.<br />
In Berlin, he improved the methods he used in Wollstein, including staining and<br />
purification techniques, and bacterial growth media, including agar plates (thanks to<br />
the advice of Angelina and Walther Hesse) and the Petri dish, named after its<br />
inventor, his assistant Julius Richard Petri. These devices are still used today. With<br />
these techniques, he was able to discover the bacterium causing tuberculosis<br />
(Mycobacterium tuberculosis) in 1882 (he announced the discovery on March 24).<br />
Tuberculosis was the cause of one in seven deaths in the mid-19th century.<br />
In 1883, Koch worked with a French research team in Alexandria, Egypt, studying<br />
cholera. Koch identified the vibrio bacterium that caused cholera, though he never<br />
managed to prove it in experiments. The bacterium had been previously isolated by<br />
Italian anatomist Filippo Pacini in 1854, but his work had been ignored due to the<br />
predominance of the miasma theory of disease. Koch was unaware of Pacini's work<br />
and made an independent discovery, and his greater preeminence allowed the
discovery to be widely spread for the benefit of others. In 1965, however, the<br />
bacterium was formally renamed Vibrio cholera Pacini 1854.<br />
In 1885, he became professor of hygiene at the University of Berlin, and later, in<br />
1891, director of the newly formed Institute of Infectious Diseases, a position which<br />
he resigned from in 1904. He started traveling around the world, studying diseases in<br />
South Africa, India, and Java.<br />
Probably as important as his work on tuberculosis, for which he was awarded a Nobel<br />
Prize (1905), are Koch's postulates, which say that to establish that an organism is the<br />
cause of a disease, it must be:<br />
• found in all cases of the disease examined<br />
• prepared and maintained in a pure culture<br />
• capable of producing the original infection, even after several generations in<br />
culture<br />
• be retrievable from an inoculated animal and cultured again.<br />
After Koch's success the quality of his own research declined (especially with the<br />
fiasco over his ineffective TB cure "tuberculin"), although his pupils found the<br />
organisms responsible for diphtheria, typhoid, pneumonia, gonorrhoea, cerebrospinal<br />
meningitis, leprosy, bubonic plague, tetanus, and syphilis, among others, by using his<br />
methods.<br />
He died on 27 May 1910 of a heart-attack in Baden-Baden, aged 66. [1]<br />
Koch crater on the Moon was named after him. The Robert Koch Prize and Medal<br />
were created to honour Microbiologists who make groundbreaking discoveries or who<br />
contribute to global health in a unique way. The first non-German to be awarded the<br />
medal was Professor Bill Hutchison of Strathclyde University in Glasgow. [2]<br />
[edit] References<br />
1. ^ Robert Koch Institute<br />
2. ^ Parasitology in Scotland<br />
[edit] Consult<br />
• Thomas Brock, Robert Koch: A Life in Medicine and Bacteriology,<br />
Washington D.C. (1999)<br />
[edit] See also<br />
• History of medicine<br />
• Microbiology<br />
• Timeline of medicine and medical technology<br />
[edit] External links
v • d • e<br />
• Biography at the Nobel Foundation website<br />
• Biography and bibliography in the Virtual Laboratory of the Max Planck<br />
Institute for the History of Science<br />
[hide]<br />
Nobel Laureates in Physiology or Medicine<br />
Emil Behring (1901) · Ronald Ross (1902) · Niels Finsen (1903) · Ivan Pavlov (1904) ·<br />
Robert Koch (1905) · Camillo Golgi / Santiago Ramón y Cajal (1906) · Alphonse<br />
Laveran (1907) · Ilya Mechnikov / Paul Ehrlich (1908) · Emil Kocher (1909) · Albrecht<br />
Kossel (1910) · Allvar Gullstrand (1911) · Alexis Carrel (1912) · Charles Robert Richet<br />
(1913) · Robert Bárány (1914) · Jules Bordet (1919) · August Krogh (1920) · Archibald<br />
Hill / Otto Meyerhof (1922) · Frederick Banting / John Macleod (1923) · Willem<br />
Einthoven (1924)<br />
Complete roster · 1901–1925 · 1926–1950 · 1951–1975 · 1976–2000 · 2001–present<br />
Retrieved from "http://en.wikipedia.org/wiki/Robert_Koch"<br />
Categories: 1843 births | 1910 deaths | German biologists | German physicians |<br />
German inventors | German microbiologists | Tuberculosis | Nobel laureates in<br />
Physiology or Medicine | German Nobel laureates | German military personnel of the<br />
Franco-Prussian War | University of Göttingen alumni | Deaths by myocardial<br />
infarction | People from the Kingdom of Hanover<br />
Views<br />
• Article<br />
• Discussion<br />
• Edit this page<br />
• History<br />
Personal tools<br />
• Log in / create account<br />
Navigation<br />
• Main Page<br />
• Contents<br />
• Featured content<br />
• Current events<br />
• Random article<br />
Interaction
interested in:<br />
by Name:<br />
List of Virus<br />
Families<br />
viral families with<br />
example viruses<br />
List of Individual<br />
Viruses<br />
all named viruses,<br />
listed with their<br />
assigned family<br />
by<br />
Structure/Genome:<br />
Virus Families by<br />
genome type<br />
RNA, DNA, positive,<br />
negative, etc.<br />
Virus Families<br />
Grouped by the<br />
Baltimore Method<br />
Listed with example<br />
viruses and hosts<br />
by Host:<br />
Virus Families by<br />
Host<br />
includes broad host<br />
categories (algae,<br />
fungi, etc.) with<br />
example viruses that<br />
infect them<br />
by Disease:<br />
Virus Families by<br />
serve as both a catalog of virus pictures on the<br />
Internet and as an educational resource to those<br />
seeking more information about viruses. To this<br />
end, it is intimately linked to All the Virology on<br />
the WWW, and our collection of Virology Courses<br />
and Tutorials.<br />
There are several ways to access the information<br />
in the Big Picture Book of Viruses. All viruses are<br />
listed according to the family to which they have<br />
been assigned by the International Committee on<br />
Taxonomy of Viruses (ICTV). The images and<br />
other data can be obtained by the routes listed at<br />
the left.<br />
What you can find here:<br />
On this page, several types of information about<br />
viruses can be found. First and foremost, we<br />
show you what they look like, either by electron<br />
microscopy or by computerassisted imaging. The<br />
viral images are listed by their taxonomic groups.<br />
Images listed have been gathered from several<br />
well known sources on the web. Each image is<br />
presented in a miniature format on this page, with<br />
the original fullsized image being available by<br />
simply clicking on the image of interest. (Note: if<br />
the original image has moved, please contact me<br />
for assistance) Also listed on the page are links to<br />
detailed information (Tutorials and On-Line<br />
Courses) about the viruses, and links to other<br />
WWW sites (All the Virology on the WWW) with<br />
additional information.<br />
Other Virology Resources:<br />
• All the Virology on the WWW<br />
The full table of contents of this site's<br />
parent with links to all the virology web<br />
sites.<br />
• On-line Virology Courses<br />
Some of the best note sets, diagrams,<br />
tutorials and video available online.
Infectious Disease<br />
includes a list of<br />
infectious diseases in<br />
humans, and links to<br />
pictures of the viruses<br />
that cause them<br />
Guides to Virus<br />
Structure:<br />
Principles of Virus<br />
Architecture<br />
contains descriptions<br />
and diagrams of virus<br />
structure (and a little<br />
history).<br />
Virus Structure from<br />
ATV's own on-line<br />
virology courses.<br />
New!<br />
ICTV Net is an email<br />
network of ICTV<br />
members that provides<br />
the virology<br />
community with the<br />
opportunity to interact<br />
with ICTV, ask<br />
questions, and even<br />
make proposals...<br />
Check it out!<br />
An Introduction to Viral Taxonomy:<br />
(from the National Library of Medicine)<br />
• The Universal System of Virus Taxonomy<br />
• Virus Nomenclature<br />
• Replicative Properties of Viruses Used in<br />
Taxonomy<br />
• References<br />
How to Add your Favorite Image to this Site:<br />
If you know of a virus picture that is not listed<br />
here, or would like to update the listing of a site,<br />
please use our virology site submission form or<br />
email me with the address. We are counting on a<br />
continuation of community support to keep this<br />
site up to date.<br />
Are you interested in more information, or<br />
assistance with your organization's Web site?<br />
If you are interested in developing a WWW site<br />
for your lab or organization, please feel free to<br />
contact me for any needed advice and/or<br />
assistance.<br />
David M. Sander, Ph.D. (david.sander@virology.net<br />
)<br />
Don't forget to sign our Guestbook!<br />
ATV Home | Table of Contents | Submit a Site | Search<br />
Tulane University | Garry Lab Contact Info | FAQ | Garry Lab Home | Tulane Medical<br />
Center
© 1995-2007. D. Sander Established 5/95.<br />
Dedication to Hans Zinsser<br />
http://www.textbookofbacteriology.net/<br />
Welcome to Todar's Online Textbook of Bacteriology textbookofbacteriology.net.<br />
This textbook has evolved from online and live-in-person lectures presented in my<br />
bacteriology courses at the University of Wisconsin-Madison. Its contents are suitable<br />
for reading or presentation in courses or course modules concerning general<br />
microbiology and medical bacteriology at the college and advanced high school levels<br />
of education. As an electronic text, new material is constantly being added, and<br />
current material is constantly being revised and updated. This is an inherent advantage<br />
of the web-based text over the tree-burner.<br />
The textbook will never be complete, as the rate of production of new information in<br />
microbiology far outruns the author's ability to acquire and properly present it. If you<br />
have suggestions, comments or criticisms regarding the textbook or its contents, or the<br />
idea of this type of textbook, please send email to me at the address below.<br />
Kenneth Todar<br />
University of Wisconsin<br />
Department of Bacteriology<br />
Madison, Wisconsin 53706<br />
kgtodar@facstaff.wisc.edu<br />
To search the entire book, enter a term or phrase in the form below<br />
General Bacteriology<br />
Overview of Bacteriology<br />
Search WWW Search Textbook of Bacteriology
The Impact of Microbes on the Environment and Human Activities<br />
Structure and Function of Procaryotes<br />
Nutrition and Growth of Bacteria<br />
Growth of Bacterial Populations<br />
Control of Microbial Growth<br />
The Diversity of Procaryotic Metabolism<br />
Regulation and Control of Metabolic Activities<br />
Bacteriophage<br />
Procaryotes in the Environment<br />
Important Groups of Procaryotes<br />
Bacterial Relationships with Animals<br />
The Nature of Host-Parasite Interactions<br />
The Bacterial Flora of Humans<br />
Mechanisms of Bacterial Pathogenicity<br />
Bacteria of Medical Importance<br />
Immune Defense against Microbial Pathogens: Innate Immunity<br />
Immune Defense against Microbial Pathogens: Adaptive Immunity<br />
Principles of Bacterial Pathogenesis<br />
Bacterial Structure in Relationship to Pathogenicity<br />
Colonization and Invasion by Bacterial Pathogens<br />
Bacterial Defense against Phagocytosis<br />
Bacterial Defense against Immune Responses<br />
Bacterial Protein Toxins
Bacterial Endotoxin<br />
Antimicrobial Agents Used in the Treatment of Infectious Disease<br />
Bacterial Resistance to Antimicrobial Agents<br />
Bacterial Pathogens and Diseases of Humans<br />
Staphylococcus and Staphylococcal Disease<br />
Streptococcus and Streptococcal Disease<br />
Streptococcus pneumoniae<br />
Listeria monocytogenes and Listeriosis<br />
Neisseria: Gonorrhea and Meningitis<br />
Haemophilus influenzae including Hib Meningitis<br />
Opportunistic Infections Caused by Pseudomonas aeruginosa<br />
Whooping Cough (Pertussis)<br />
E. coli: Gastroenteritis, Urinary Tract Infections and Neonatal Meningitis<br />
Cholera<br />
Salmonella and Salmonellosis<br />
Shigella and Shigellosis<br />
Pathogenic Clostridia, including Tetanus and Botulism<br />
Bacillus cereus Food Poisoning<br />
Bacillus anthracis and Anthrax<br />
Diphtheria<br />
Tuberculosis<br />
Rickettsial Diseases, including Rocky Mountain Spotted Fever<br />
Emerging Pathogens
Borrelia burgdorferi<br />
Vibrio vulnificus<br />
Important Groups of Procaryotes In progress: Enteric bacteria; Lactic acid bacteria;<br />
Plant pathogenic bacteria<br />
Bacillus and Related Endospore-forming Bacteria<br />
Kenneth Todar has taught microbiology to undergraduate students at The University of Texas,<br />
University of Alaska and University of Wisconsin since 1969. He received a PhD in Microbiology<br />
in 1972 from The University of Texas-Austin. His main teaching interests are in general<br />
microbiology, bacterial diversity, microbial ecology and pathogenic bacteriology. Currently, he is<br />
an emeritus lecturer at the University of Wisconsin-Madison, where he teaches Microbiology 100,<br />
"The Microbial World". He resides in Madison, Wisconsin and Silvergate, Montana.<br />
WEB TEXT REVIEW (SCIENCE Magazine Vol 304: 1421)<br />
"The Good, the Bad, and the Deadly"<br />
The pearly droplets in this photo are colonies of Bacillus<br />
anthracis, the bacterium that causes anthrax. The bugs exude a<br />
goopy coating that repels immune system assaults and allows<br />
them to establish a foothold in the body. Learn more about the<br />
tricks bacteria use to prosper almost everywhere on Earth in<br />
this Web text from microbiologist Kenneth Todar of the<br />
University of Wisconsin, Madison. High school and college<br />
students can absorb the basics of bacterial structure, physiology,<br />
classification, and ecology.The book emphasizes medical<br />
microbiology, exploring how bacteria hitch a ride from host to<br />
host, how the body tries to corral invading microbes, and how<br />
the bugs elude these defenses. For example, the cholera<br />
bacterium releases a toxin that induces intestinal cells to spill<br />
ions and water, producing potentially lethal diarrhea.
OTHER CITATIONS, REVIEWS, ADAPTATIONS<br />
textbookofbacteriology.net<br />
© 2008 Kenneth Todar University of Wisconsin-Madison Department of<br />
Bacteriology.<br />
Written and edited by Kenneth Todar University of Wisconsin-Madison<br />
Department of Bacteriology. All rights reserved.