Abstract
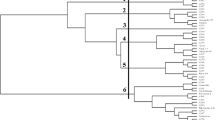
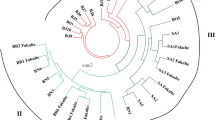
Three molecular markers, including start codon targeted (SCoT) polymorphism, directed amplification of minisatellite-region DNA polymerase chain reaction (DAMD-PCR), and inter simple sequence repeat (ISSR) markers, were compared in terms of their informativeness and efficiency for analysis of genetic relationships among 38 accessions of eight annual Cicer species. The results were as follows: (1) the highest level of detected polymorphism was observed for all three marker types; (2) the rate of diversity for the three marker techniques was approximately equal, and the correlation coefficients of similarity were statistically significant for all three marker systems; (3) the three molecular markers showed relatively similar phylogenetic grouping for examined species. Diversity analysis showed that Cicer reticulatum is the closest wild species to the cultivated chickpea, and this finding supports the hypothesis that C. reticulatum is the most probable progenitor of the cultivated species. C. bijugum, C. judaicum, and C. pinnatifidum were clustered together, and in other clusters C. yamashitae and C. cuneatum were grouped close together. To our knowledge, this is the first detailed comparison of performance among two targeted DNA region molecular markers (SCoT and DAMD-PCR) and the ISSR technique on a set of samples of Cicer. The results provide guidance for future efficient use of these molecular methods in genetic analysis of Cicer.




Similar content being viewed by others
References
Ahmad F, Slinkard AE (1992) Genetic relationships in the genus Cicer L. as revealed by polyacrylamide gel electrophoresis of seed storage proteins. Theor Appl Genet 84:688–692
Ahmad F, Slinkard AE, Scoler GJ (1987) The cytogenetic relationship between Cicer judaicum Boiss and Cicer chroassannicum (Bge). M Pop Genome 29:883–886
Ahmad F, Gaur PM, Slinkard AE (1992) Isozyme polymorphism and phylogenetic interpretations in the genus Cicer L. Theor Appl Genet 83:620–627
Akem C, Caccarelli S, Erkine W, Lenne J (2000) Using genetic diversity for disease resistance in agricultural production. Outlook Agric 29:25–30
Andersen JR, Lubberstedt T (2003) Functional markers in plants. Trends Plant Sci 8:554–560
Anonymous (1995) Kabuli chickpea improvement. Annu rep germplasm program: legumes. ICARDA, Aleppo, pp 59–132
Aruna R, Rao DM, Sirvaramakrishnan S, Reddy LJ, Bramel P, Upadhaya H (2008) Efficiency of three DNA markers in revealing genetic variation among wild Cajanus species. Plant Genet Resour 7(2):113–121
Botstein B, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Ann J Hum Genet 32:314–331
Collard BCY, Mackill DJ (2009) Start Codon Targeted (SCoT) polymorphism: a simple novel DNA marker technique for generating gene-targeted markers in plants. Plant Mol Biol 27:86–93
Excoffier L, Smouse P, Quattro J (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: applications to human mitochondrial DNA restriction data. Genetics 131:479–491
Garcia GM, Stalker HT, Shroeder E, Kochert G (1996) Identification of RAPD, SCAR and RFLP markers tightly linked to nematode resistance genes introgressed from Arachis cardenasii into Arachis hypogaea. Genome 39:836–845
Gascuel O (1997) Concerning the NJ algorithm and its unweighted version, UNJ. In: Mirkin B, McMorris FR, Roberts FS, Rzhetsky A (eds) Mathematical hierarchies and biology. DIMACS series in discrete mathematics and theoretical computer science. Providence, RI. Amer Math Soc 149–170
Gupta PK, Rustgi S (2004) Molecular markers from the transcribed/expressed region of the genome in higher plants. Funct Integr Geonomics 4:139–162
Iruela M, Rubio J, Cubero JI, Gil J, Milan T (2002) Phylogenetic analysis in the genus Cicer and cultivated chickpea using RAPD and ISSR markers. Theor Appl Genet 104:643–651
Kalendar R (2007) FastPCR: a PCR primer design and repeat sequence searching software with additional tools for the manipulation and analysis of DNA and protein. Available at www.biocenter
Kang HW, Park DS, Go SJ, Eun MY (2002) Fingerprinting of diverse genomes using PCR with universal rice primers generated from repetitive sequence of Korean weedy rice. Mol Cells 13:281–287
Karaca M, Saha S, Zipf A, Jenkins JN, Lang DJ (2002) Genetic diversity among forage bermudagrass (Cynodon spp.): evidence from chloroplast and nuclear DNA fingerprinting. Crop Sci 42:2118–2127
Kazan K, Muehlbauer FJ (1991) Allozyme variation and phylogeny in annual species of Cicer (Leguminosae). Plant Syst Evol 175:11–21
Labdi M, Robertson LD, Singh KB, Charrier A (1996) A genetic diversity and phylogenetic relationships among the annual Cicer species as revealed by isozyme polymorphism. Euphytica 88:181–188
Ladizinsky and Adler (1976) Genetic relationships among the annual species of Cicer L. Theor Appl Genet 48:197–203
Lassner MW, Peterson P, Yoder JI (1989) Simultaneous amplification of multiple DNA fragments by polymerase chain reaction in the analysis of transgenic plants and their progeny. Plant Mol Biol Rep 7:116–128
Mantel NA (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
McCoy TJ, Echt CS (1993) Potential of trispecies bridge crosses and random amplified polymorphic DNA markers for introgression of Medicago daghestanica and Medicago pironae germplasm into alfalfa (Medicago sativa). Genome 36:594–601
Muehlbauer FJ, Kaiser WJ, Simon CJ (1994) Potential for wild species in cool season food legume breeding. Euphytica 73:109–114
Nguyen TT, Taylor PWJ, Redden RJ, Ford R (2004) Genetic diversity estimates in Cicer using AFLP analysis. Plant Breed 123:173–179
Ocampo B, Venora G, Errico A, Singh KB, Saccardo F (1992) Karyotype analysis in the genus Cicer. J Genet Breed 46:229–240
Patil P, Vrinten PL, Scoles GJ, Slinkard AE (1995) Variation in the ribosomal RNA units of the genera Lens and Cicer. Euphytica 83:33–42
Perrier X, Flori A, Bonnot F (2003) Data analysis methods. In: Hamon P, Seguin M, Perrier X, Glaszmann JC (eds) Genetic diversity of cultivated tropical plants. Science, Enfield, pp 43–76
Pundir RPS, Vander Maesen LJG (1983) Interspecific hybridization in Cicer. Int Chickpea Newsl 8:4–5
Rajesh PN, Sant VJ, Gupta VS, Muehlbauer FJ, Ranjekar PK (2002) Genetic relationships among annual and perennial wild species of Cicer using inter simple sequence repeat (ISSR) polymorphism. Euphytica 129:15–23
Robertson LD, Ocampo B, Singh KB (1997) Morphological variation in wild annual Cicer species in comparison to the cultigens. Euphytica 95:309–319
Rohlf FJ (1997) NTSYS-Pc. Numerical taxonomy and multivariate analysis system version 2.1. Exeter Software, Setauket, p 7
Saini N, Jain N, Jain RK (2004) Assessment of genetic diversity within and among Basmati and non-Basmati rice varieties using AFLP, ISSR and SSR markers. Euphytica 140:133–146
Sant VJ, Sainani MN, Sami-subbu R, Ranjekar PK, Gupta VS (2000) Ty1-copia retrotransposon-like elements in chickpea genome: their identification, distribution and use for diversity analysis. Gene 257:157–166
Singh KB, Ocampo B (1993) Interspecific hybridization in annual Cicer species. J Genet Breed 47:199–204
Singh KB, Ocampo B (1997) Exploitation of wild Cicer species for yield improvement in chickpea. Theor Appl Genet 95:418–423
Singh KB, Malhotra RS, Halila MH, Knights EJ, Verma MM (1994) Current status and future strategy in breeding chickpea for resistance to biotic and abiotic stresses. Euphytica 73:137–149
Sudupak MA (2004) Inter and intra-species Inter Simple Sequence Repeat (ISSR) variations in the genus Cicer. Euphytica 135:229–238
Sudupak MA, Akkaya MS, Kence A (2002) Analysis of genetic relationships among perennial and annual Cicer species growing in turkey using RAPD markers. Theor Appl Genet 105:1220–1228
Talebi R, Jelodar NB, Mardi M, Fayaz F, Furman BJ, Bagheri NA (2009) Phylogenetic diversity relationship among annual Cicer species using random amplified polymorphic DNA. Gener Appl Plant Physiol 35:3–12
Tayyar RI, Lukaszewiski AJ, Waines JG (1994) Chromosome banding patterns in the annual species of Cicer. Genome 37:656–663
Torres AM, Weeden NF, Martin A (1993) Linkage among isozyme, RFLP and RAPD markers in Vicia faba. Theor Appl Genet 85:935–945
Udupa SM, Robertson LD, Weigand F, Baum M, Kahl G (1999) Allelic variation at (TAA) microsatellite loci in a world collection of chickpea (Cicer arietinum L) germplasm. Mol Gen Genet 261:354–363
Vairinhos F, Murray DR (1983) The seed proteins of chickpea comparative studies of Cicer arietinum, C. reticulatum and C.echinospermum (Leguminosae). Plant Syst Evol 142:15–22
van der Maesen LJG (1987) Origin, history and taxonomy of chickpea. In: Saxena MC, Singh KB (eds) The chickpea. CAB Int Publ, UK, pp 11–34
van Rheenen HA, Pundir PRS, Miranda JH (1993) How to accelerate the genetic improvement of a recalcitrant crop species such as chickpea. Curr Sci 65:414–417
Vavilov NI (1926) The origin of the cultivation of ‘primary’ crops, in particular cultivated hemp. In: Studies on the origin of cultivated plants, Institute of applied botany and plant breeding, Leningrad, USSR, pp 221–233
Acknowledgments
We would like to thank Islamic Azad University, Sanandaj Branch for support of this study. Also, we wish to present our special thanks to the International Center for Agricultural Research in the Dry Areas (ICARDA), Aleppo, Syria and the Australian Temperate Field Crops Collection (ATFCC) at the Victorian Institute for Dryland Agriculture (VIDA), Horsham, Australia for kindly supplying seeds of Cicer species.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Amirmoradi, B., Talebi, R. & Karami, E. Comparison of genetic variation and differentiation among annual Cicer species using start codon targeted (SCoT) polymorphism, DAMD-PCR, and ISSR markers. Plant Syst Evol 298, 1679–1688 (2012). https://doi.org/10.1007/s00606-012-0669-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00606-012-0669-6