Abstract
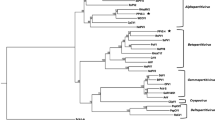
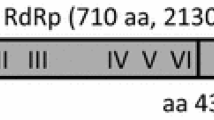
Many bands were detected on an electrophoretic profile of double-stranded (ds) RNA preparation from a single strain of Fusarium poae isolated from wheat. When the purified dsRNA sample was deep-sequenced by a next-generation sequencer, sixteen virus-like assembled contigs with predicted amino acid sequences showing homologies to respective viral RNA-dependent RNA polymerases (RdRps) were found by BLAST analysis. Fourteen out of sixteen sequences showed homologies to RdRps of known mycoviruses, that is, four mitoviruses, two narnaviruses, two partitiviruses, an alternavirus, a fusarivirus, a hypovirus, a victorivirus, and two unclassified mycoviruses, Sclerotinia sclerotiorum dsRNA mycovirus-L and Aspergillus foetidus slow virus 2, respectively. The other two putative viral RdRp sequences showed homologies to those of members of negative-stranded RNA viruses, the Ophiovirus and the Phlebovirus respectively, which mycoviruses had been not ever assigned to. Based on genome structure and phylogenetic analysis, both viruses were thought to be members of novel respective negative-stranded RNA virus groups. The presences of all sixteen viral RdRp sequences identified by BLAST analysis were confirmed by sequencing RT-PCR products generated from the starting dsRNA material using primers designed from the de novo assembled sequences of respective putative mycoviruses. Since the single strain of F. poae was considered to be multiply infected with mycoviruses from novel taxonomical groups in addition to many common mycoviruses, the RNA virome of the strain was found to be highly diverse.






Similar content being viewed by others
References
A.M.Q. King, M.J. Adams, E.B. Carstens, E.J. Lefkowitz, Virus Taxonomy. Ninth Report of the International Committee on Taxonomy of Viruses. (Elsevier Academic Press, London, San Diego, 2011)
H. Kondo, S. Chiba, K. Toyoda, N. Suzuki, Virology 435, 201 (2013)
L. Liu, J. Xie, J. Cheng, Y. Fu, G. Li, X. Yi, D. Jiang, Proc. Natl. Acad. Sci. U.S.A. 111, 12205 (2014)
S.L. Anagnostakis, Science 215, 466 (1982)
S. Chiba, L. Salaipeth, Y.H. Lin, A. Sasaki, S. Kanematsu, N. Suzuki, J. Virol. 83, 12801 (2009)
X. Yu, B. Li, Y. Fu, J. Xie, J. Cheng, S.A. Ghabrial, G. Li, X. Yi, D. Jiang, Proc. Natl. Acad. Sci. U.S.A. 110, 1452 (2013)
H.W. Schroeder, J.J. Christensen, Phytopathology 53, 831 (1963)
S.A. Stenglein, J. Plant Pathol. 91, 25 (2009)
C. Fekete, G. Giczey, I. Papp, L. Szabó, L. Hornok, FEMS Microbiol. Lett. 131, 295–299 (1995)
L. Wang, J. Zhang, H. Zhang, D. Qiu, L. Guo, Int. J. Mol. Sci. 17, 641 (2016)
H.I. Nirenberg, T. Aoki, Mycoscience 38, 329 (1997)
A. Sasaki, M. Miyanishi, K. Ozaki, M. Onoue, K. Yoshida, Arch. Virol. 150, 1069 (2005)
S.F. Altschul, F.W. Gish, W. Miller, E.W. Myers, D.J. Lipman, J. Mol. Biol. 215, 403 (1990)
J. Felsenstein, Evolution 39, 783 (1985)
N. Saitou, M. Nei, Mol. Biol. Evol. 4, 406 (1987)
P. Compel, I. Papp, M. Bibo, C. Fekete, L. Hornok, Virus Genes 18, 49 (1999)
G. Cai, K. Myers, W.E. Fry, B.I. Hillman, Arch. Virol. 157, 165 (2012)
L. Donaire, J. Rozas, M.A. Ayllón, Virology 489, 158 (2016)
S.Y.L. Marzanoa, L.L. Domier, Virus Res. 219, 11 (2016)
P. Martınez-Alvarez, E.J. Vainio, L. Botella, J. Hantula, J.J. Diez, Arch. Virol. 159, 2153 (2014)
B. Paquin, M.J. Laforest, L. Forget, I. Roewer, Z. Wang, J. Longcore, B.F. Lang, Curr. Genet. 31, 380 (1997)
M.L. Nibert, S.A. Ghabrial, E. Maiss, T. Lesker, E.J. Vainio, D. Jiang, N. Suzuki, Virus Res. 188, 128 (2014)
M. Nogawa, T. Kageyama, A. Nakatani, G. Taguchi, M. Shimosaka, M. Okazaki, Biosci. Biotechnol. Biochem. 60, 784 (1996)
H. Osaki, A. Sasaki, K. Nomiyama, H. Sekiguchi, K. Tomioka, T. Takehara, Virus Genes 50, 466 (2015)
Z. Kozlakidis, N. Herrero, S. Ozkan, L. Kanhayuwa, A. Jamal, M.F. Bhatti, R.H.A. Coutts, Arch. Virol. 158, 267 (2013)
S.Y.L. Marzano, B.D. Nelson, O. Ajayi-Oyetunde, C.A. Bradley, T.J. Hughes, G.L. Hartman, D.M. Eastburn, L.L. Domier, J. Virol. 90, 6846 (2016). doi:10.1128/JVI.00357-16
Y.M. Chu, J.J. Jeon, S.J. Yea, Y.H. Kim, S.H. Yun, Y.W. Lee, K.H. Kim, Appl. Environ. Microbiol. 68, 2529 (2002)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Edited by Thomas Hohn.
Rights and permissions
About this article
Cite this article
Osaki, H., Sasaki, A., Nomiyama, K. et al. Multiple virus infection in a single strain of Fusarium poae shown by deep sequencing. Virus Genes 52, 835–847 (2016). https://doi.org/10.1007/s11262-016-1379-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-016-1379-x