Abstract
Pestalotiopsis is a taxonomically confused, pathogenic and chemically creative genus requiring a critical re-examination using a multi-gene phylogeny based on ex-type and ex-epitype cultures. In this study 40 isolates of Pestalotiopsis, comprised of 28 strains collected from living and dead plant material of various host plants from China were studied by means of morphology and analysis of ITS, β–tubulin and tef1 gene sequence data. Based on molecular and morphological data we describe 14 new species (Pestalotiopsis asiatica, P. chinensis, P. chrysea, P. clavata, P. diversiseta, P. ellipsospora, P. inflexa, P. intermedia, P. linearis, P. rosea, P. saprophyta, P. umberspora, P. unicolor and P. verruculosa) and three species are epitypified (P. adusta, P. clavispora and P. foedans). Of the 10 gene regions (ACT, β-tubulin, CAL, GPDH, GS, ITS, LSU, RPB 1, SSU and tef1) utilized to resolve cryptic Pestalotiopsis species, ITS, β–tubulin and tef1 proved to be the better markers. The other gene regions were less useful due to poor success in PCR amplification and/or in their ability to resolve species boundaries. As a single gene tef1 met the requirements for an ideal candidate and functions well for species delimitation due to its better species resolution and PCR success. Although β-tubulin showed fairly good differences among species, a combination of ITS, β-tubulin and tef1 gene data gave the best resolution as compared to single gene analysis. This work provides a backbone tree for 22 ex-type/epitypified species of Pestalotiopsis and can be used in future studies of the genus.
Similar content being viewed by others
Introduction
Pestalotiopsis, an appendage-bearing conidial asexual form in the family Amphisphaeriaceae (Barr 1975, 1990; Kang et al. 1998, 1999) is widely distributed throughout the tropical and temperate ecosystems (Bate-Smith and Metcalfe 1957). It is an important plant pathogenic genus (Yasuda et al. 2003; Das et al. 2010; Maharachchikumbura et al. 2011) with more than 235 species, traditionally named according to their host associations (Guba 1961; Steyaert 1949; Venkatasubbaiah et al. 1991; Kohlmeyer and Kohlmeyer 2001). Following the discovery of taxol, the multimillion dollar anti-cancer drug, from P. microspora (Speg.) G.C. Zhao & N. Li, an endophytic strain isolated from Taxus wallachiana (Strobel et al. 1996), the importance of the genus has increased considerably (Strobel et al. 2002; Xu et al. 2010). Chemical exploration of endophytic Pestalotiopsis species subsequently increased in an unprecedented way (Wei and Xu 2004; Tejesvi et al. 2007a, b). Species belonging to the genus Pestalotiopsis are thought to be a rich source for bioprospecting when compared to other fungal genera (Aly et al. 2010; Xu et al. 2010). Xu et al. (2010) reviewed 130 different compounds isolated from species of Pestalotiopsis in the preceding 10 years. These included bioactive alkaloids, terpenoids, isocoumarin derivatives, coumarins, chromones, quinones, semiquinones, peptides, xanthones, xanthone derivatives, phenols, phenolic acids, and lactones with a range of antifungal, antimicrobial, and antitumor activities.
Due to their ability to switch life modes, many endophytic and pathogenic Pestalotiopsis species persist as saprobes (Hyde et al. 2007; Zhou and Hyde 2001). Species of Pestalotiopsis have been isolated as saprobes from dead leaves, bark and twigs (Guba 1961). Several species have been recovered from soil, polluted stream water, wood, paper, fabrics and wool (Guba 1961). For example, P. bicolor (Ellis & Everh.) A.R. Liu, T. Xu & L.D. Guo, P. funerea (Desm.) Steyaert, P. monochaetioides (Doyer) Steyaert, P. montellica (Sacc. & Voglino) Tak. Kobay., P. disseminata (Thüm.) Steyaert, P. foedans (Sacc. & Ellis) Steyaert, P. versicolor (Speg.) Steyaert and P. virgatula (Kleb.) Steyaert are common saprobic species recorded either from decaying leaves or bark. However, there is less recent data on saprobic Pestalotiopsis species (Table 1).
The use of molecular data in resolving Pestalotiopsis species has been reviewed by Hu et al. (2007), Tejesvi et al. (2007a), Liu et al. (2010) and Maharachchikumbura et al. (2011). These studies have suggested that multi-locus phylogenetic analysis is needed to resolve the cryptic species in the genus. Furthermore, species need to be epitypified so that we have sequence data pinned to names and thus can confidently name species in future (Cai et al. 2011).
We have been studying the genus Pestalotiopsis and testing the use of various genes to resolve species boundaries. In this study, we report on 28 isolates sourced from plant material from China. All isolated species were first morphologically characterised and then sequenced using ITS, β-tubulin and tef1 genes. In order to select suitable gene regions for better species resolution, we analyzed nuclear ribosomal large subunit rDNA (LSU), nuclear ribosomal small subunit rDNA (SSU), partial actin (ACT), glutamine synthase (GS), glyceraldehyde-3-phosphate dehydrogenase (GPDH), RNA polymerase II (RPB1) and calmodulin (CAL) gene regions for several isolates of Pestalotiopsis. We compared the morphological data versus the sequence data from single and combined genes to establish which characters satisfactorily resolve the species. As a result, we epitypified three species and describe 14 new saprobic Pestalotiopsis species. It is our hope that this work will provide a backbone phylogenetic tree for 22 type/epitypified species, which can be used in future taxonomic work on the genus.
Methods and materials
Isolation and identification of pathogen
Dead plant tissues were collected from different sites in China. The samples were placed in separate plastic bags lined with tissue paper, sprayed with sterile water to create humid conditions and incubated at room temperature. The fungi present on the samples were isolated by single spore culture technique (Chomnunti et al. 2011). In short, a conidiomata was immersed in 300 μl of sterile distilled water on a slide and left a few minutes so that the conidia were discharged. A conidial suspension was made, small drops were placed on water agar (WA) in Petri dishes and kept at room temperature for 8–12 h for conidia to germinate; single germinating conidia were transferred to potato dextrose agar (PDA) plates. The plates were incubated at 25 °C for 7 to 10 days. Colonies grown on PDA were transferred to PDA slants, and stored at 4 °C for further study. Sporulation was induced by placing sterilized carnation leaves on the surface of PDA with growing mycelia. The morphology of fungal colonies was recorded following the method of Hu et al. (2007). Fungal mycelia and spores were observed under a light microscope and photographed. All microscopic measurements were done with Tarosoft image framework (v. 0.9.0.7) and 30 conidial measurements were taken for each isolate. Isolates were deposited in Novozymes, Beijing and were also transferred to MFLUCC from Novozymes by Material Transfer Agreement and cannot be distributed to a third party. All other cultures dealt with in this study were obtained from China General Microbiological Culture Collection (CGMCC) and The International Collection of Micro-organisms from Plants (ICMP).
DNA extraction
Total genomic DNA was extracted from fresh cultures using a modified protocol of Guo et al. (2000). Fresh fungal mycelia (500 mg) was scraped from the margin of a PDA plate incubated at 25 °C for 7 to 10 days and transferred into a 1.5 ml centrifuge tube with 100 μl of preheated (60 °C) 2X CTAB extraction buffer (2 % (w/v) CTAB, 100 mM Tris-HCl, 1.4 M NaCl, 20 mM EDTA, pH 8.0), and 200 mg sterilized quartz sand. Mycelia were ground using a glass pestle for 5 min and an extra 500 μl 2X CTAB preheated (60 °C) was added and incubated in a 65 °C water bath for 30 min with occasional shaking. 500 μl of phenol:chloroform (1:1) was added to each tube and shaken thoroughly to form an emulsion. The mixture was spun at 11,900 g for 15 min at 25 °C in a microcentrifuge and the supernatant phase decanted into a fresh 1.5 ml tube. Supernatant containing DNA was re-extracted with phenol: chloroform (1:1) at 4 °C until no interface was visible. 50 μl of 5 M KOAc was added into the supernatant followed by 400 μl of isopropanol and inverted gently to mix. The genomic DNA was precipitated at 9,200 g for 2 min at 4 °C in a microcentrifuge. The DNA pellet was washed with 70 % ethanol twice and dried using SpeedVac® (AES 1010; Savant, Holbrook, NY, USA) until dry. The DNA pellet was then resuspended in 100 μl TE buffer (10 mM Tris-HCl, 1 mM EDTA).
PCR amplification
The ITS and 5.8 S region of rDNA fragment was amplified using primer pairs ITS5 (5′-GGAAGTAAAAGTCGTAACAAGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) (White et al. 1990), partial β-tubulin gene region was amplified with primer pairs BT2A (5′-GGTAACCAAATCGGTGCTGCTTTC-3′) and BT2B (5′ ACCCTCAGTGTAGTGACCCTTGGC-3′) (Glass & Donaldson 1995; O’Donnell & Cigelnik 1997) and tef1 was amplified using the primer pairs EF1-526 F (5′-GTCGTYGTYATY GGHCAYGT-3′) and EF1-1567R (5′-ACHGTRCCRATACCACCRATCTT-3′) (Rehner 2001). In addition to above three gene regions selected LSU, SSU, Actin, GS, GPDH, RPB1and CAL regions were amplified using primer pair/s listed in Table 2.
PCR was performed with the 25 μl reaction system containing 19.5 μl of double distilled water, 2.5 μl of 10× Taq buffer with MgCl2, 0.5 μl of dNTP (10 mM each), 0.5 μl of each primer (10 μM), 0.25 μl Taq DNA polymerase (5 U μl−1), 1.0 μl of DNA template. The thermal cycling program was as follows: For ITS an initial denaturing step of 95 °C for 3 min, followed by 35 amplification cycles of 95 °C for 30 s, 52 °C for 45 s and 72 °C for 90 s, and a final extension step of 72 °C for 10 min. For β-tubulin PCR conditions were an initial step of 3 min at 95 °C, 35 cycles of 1 min at 94 °C , 50 s at 55 °C , and 1 min at 72 °C, followed by 10 min at 72 °C. For tef1, an initial step of 5 min at 94 °C, 10 cycles of 30 s at 94 °C, 55 s at 63 °C or 66 °C (decreasing 1 °C per cycle), 90 s at 72 °C, plus 36 cycles of 30 s at 94 °C, 55 s at 53 °C or 56 °C, 90 s at 72 °C, followed by 7 min at 72 °C. The LSU, SSU, Actin, GS, GPDH, RPB 1and CAL regions were tested under different optimal conditions (not shown). The PCR products were verified by staining with Goldview (Guangzhou Geneshun Biotech, China) on 1 % agarose electrophoresis gels.
Phylogenetic analysis
DNAStar and SeqMan were used to obtain consensus sequences from sequences generated from forward and reverse primers. Single locus dataset and combination of multi-locus dataset of three gene regions were aligned using CLUSTALX (v. 1.83) (Thompson et al. 1997). The sequences were further aligned using default settings of MAFFTv6 (Katoh and Toh 2008; mafft.cbrc.jp/alignment/server/) and manually adjusted using BioEdit (Hall 1999) to allow maximum alignment and minimum gaps. A maximum parsimony analysis (MP) was performed using PAUP (Phylogenetic Analysis Using Parsimony) v. 4.0b10 (Swofford 2002). Ambiguously aligned regions were excluded and gaps were treated as missing data. Trees were inferred using the heuristic search option with TBR branch swapping and 1,000 random sequence additions. Maxtrees were set up to 5,000, branches of zero length were collapsed and all multiple parsimonious trees were saved. Tree length [TL], consistency index [CI], retention index [RI], rescaled consistency index [RC], homoplasy index [HI], and log likelihood [-ln L] (HKY model) were calculated for trees generated under different optimality criteria. The robustness of the most parsimonious trees was evaluated by 1,000 bootstrap replications resulting from maximum parsimony analysis, each with 10 replicates of random stemwise addition of taxa (Felsenstein 1985). The Kishino–Hasegawa tests (Kishino & Hasegawa 1989) were performed to determine whether the trees inferred under different optimality criteria were significantly different. Trees were viewed in Treeview (Page 1996).
Results
Phylogenetic trees were constructed using individual and combined ITS, β-tubulin and tef1 sequences for our 40 isolates of Pestalotiopsis with a Seiridium species as the outgroup taxon and other sequences downloaded from GenBank (Table 3). We tested 10 genes in PCR amplification, alignment and the species delimitation in Pestalotiopsis (Tables 4 and 5) and found that β-tubulin and tef1 were the optimal genes, while ITS is included as it is the accepted barcode for fungi (Schoch et al. 2012). We used the available type ITS sequences from other studies (Pestalotiopsis pallidotheae, P. hainanensis, P. jesteri and P. kunmingensis), for comparison.
Sequence analysis of ITS from Pestalotiopsis strains
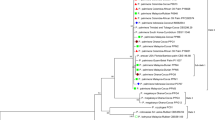
ITS sequences from the types (Pestalotiopsis pallidotheae, P. hainanensis, P. jesteri and P. kunmingensis) for Pestalotiopsis were analysed with our isolates used in this study. The alignment comprised 45 taxa and 527 characters (including gaps) (Fig. 1). Parsimony analysis indicates that 398 characters were constant, 41 variable characters parsimony-uninformative and 88 characters are parsimony-informative. The parsimony analysis of the data matrix resulted in two equally parsimonious trees and the first tree (TL = 243, CI = 0.683, RI = 0.910, HI = 0.317, RC = 0.622) is shown here (Fig. 1).
In the ITS phylogram, the Pestalotiopsis strains separated into three major clades, named A, B and C with high bootstrap support (Fig. 1). Clade A comprised species having pale brown or olivaceous concolorous median conidial cells. Clade B comprised species having versicolorous median conidial cells and Clade C species with dark concolorous median conidial cells, with knobbed apical appendages. The species within each group were not well resolved at the terminal clades. Specifically, all taxa in Clade B did not separate into distinct species but clustered in two subclades. Species resolution was higher in Clade A, although a few species are not well resolved at the terminal ends. Thus, ITS had lower inter-specific variation and, therefore, further gene sequences are needed to determine genetic variation within each biological species.
Sequence analysis of β-tubulin gene data from Pestalotiopsis strains
The aligned dataset for β-tubulin sequences comprised 37 taxa and 487 characters (including gaps). Parsimony analysis indicated that 285 characters were constant, 48 variable characters parsimony-uninformative and 154 characters parsimony-informative. The parsimony analysis of the data matrix resulted in two equally parsimonious trees and the first tree (TL = 410 steps, CI = 0.702, RI = 0.912, HI = 0.298 and RC = 0.640) was shown here (Fig. 2).
Analysis of the β-tubulin gene sequences resulted in a phylogram (Fig. 2) in which the Pestalotiopsis species separated into three major clades, A, B and C with high bootstrap support. Clade A comprised twelve well-resolved species. It was not possible to obtain PCR products from P. chinensis (MFLUCC 12-0273), P. intermedia (MFLUCC 12-0260), P. linearis (MFLUCC 12-0272) and P. verruculosa (MFLUCC 12-0274) using primer pair BT2A and BT2B. Although most of the species were well-resolved in the β-tubulin tree, the success rate of PCR has been low for this gene. Therefore, further molecular loci were needed to resolve the species in this genus.
Sequence analysis of tef1 gene data from of Pestalotiopsis strains
The aligned dataset for tef1 sequence data comprised 39 taxa and 1,005 characters (including gaps). Among these, 723 characters were constant, 87 variable characters parsimony-uninformative and 195 characters parsimony-informative. The parsimony analysis resulted in six equally parsimonious trees and the first tree (TL = 606 steps, CI = 0.670, RI = 0.896, HI = 0.330 and RC = 0.600) is shown here (Fig. 3).
In the tef1 phylogram (Fig. 3), the Pestalotiopsis strains separated into three major Clades, A, B and C with high bootstrap support. In comparison to ITS and β-tubulin, the tef1 gene clearly separated all species used in this study at the species level, with high bootstrap support. The branch lengths of neighboring clades are longest in the tef1 gene region and thus signifies speciation in Pestalotiopsis. It was not successful in the amplification of species P. chinensis (MFLUCC 12-0273) and P. unicolor (MFLUCC 12-0276).
Combined sequence analysis of ITS, β-tubulin and tef1 gene data from Pestalotiopsis strains
The aligned data matrix for combined ITS, β-tubulin and tef1 sequences consisted of 41 taxa and 2,047 characters (including gaps). Parsimony analysis indicated that 1,450 characters were constant, 170 variable characters parsimony-uninformative and 427 characters parsimony-informative. The parsimony analysis of the data matrix resulted in a single parsimonious tree (TL = 1,193 steps, CI = 0.685, RI = 0.907, HI = 0.315, RC = 0.621) (Fig. 4).
In the analysis of the combined dataset from ITS, β-tubulin and tef1 genes, all species separated into three major clades A, B and C with high bootstrap support. Combined sequence analysis successfully resolved most of the Pestalotiopsis species used in this study with high bootstrap supports. The bootstrap support value of terminal and internal node has been increased as compared to the single gene phylogenetic trees.
In this study we attempted to obtain sequence data from 10 genes. In contrast to the other genes, ITS, β-tubulin and tef1 were relatively easy to amplify, sequence and align. β-tubulin and tef1 also contained considerably more phylogenetic informative characters. ITS sequence data has relatively poor species resolution for the genus Pestalotiopsis, even though it is now standardized as the universal DNA barcode marker for the fungi (Schoch et al. 2012). Therefore, ITS can be used as rough identification guide for some species in Pestalotiopsis. β-tubulin and tef1 successfully resolved most of the strains analyzed in this study to species within Pestalotiopsis, although tef1 had a higher PCR success rate when compared to β-tubulin. Thus, due to its better species resolution and PCR success rate, we suggest that tef1 is an additional barcode for Pestalotiopsis species.
Although 235 species have been described in the genus, only those few with sequence data are included in this study. The new species described below are based on molecular data and distinct morphological characteristics. At the terminal ends of the clades, most species can be differentiated from closely related species in the β-tubulin and tef1 and combined ITS, β-tubulin and tef1 phylograms. All designated epitypes have spore characters fitting those of the holotype and are supported as distinct based on molecular data.
Taxonomy
Pestalotiopsis adusta (Ellis & Everh.) Steyaert, Trans. Br. mycol. Soc. 36: 82 (1953)
Basionym: Pestalotia adusta Ellis & Everh., J. Mycol. 4(6): 51 (1888)
MycoBank: MB302600
Description from holotype (Fig. 5a–h)
Conidiomata 80–150 μm diam., acervulus, subepidermal in origin, with basal stroma, with lateral wall 2–4 cells thick comprising hyaline to pale brown cells of textura angularis. Conidiophores indistinct. Conidiogenous cells discrete, simple, short, filiform. Conidia 16–20 × 5–7 μm (\( \overline x = {18}.{7} \times {6}.{2}\;\mu {\text{m}} \)), fusiform to ellipsoid, straight to slightly curved, 4-septate, with short basal cell, obtuse, hyaline, thin-walled and verruculose, 2.7–3.8 μm long (\( \overline x = {3}.{2}\;\mu {\text{m}} \)); with three median cells, doliiform to subcylindrical, concolorous, olivaceous, with septa and periclinal walls darker than the rest of the cell, together 12.4–13.8 μm long (\( \overline x = {13}.{2}\,\mu {\text{m}} \)) second cell from base 4.3–5.3 μm (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); third cell 4–4.7 μm (\( \overline x = {4}.{2}\;\mu {\text{m}} \)); fourth cell 3.8–4.4 μm (\( \overline x = {4}\;\mu {\text{m}} \)); apical cell hyaline, conic, 2.4–3.4 μm long (\( \overline x = {3}\;\mu {\text{m}} \)); with two to three appendages, 7–15 μm long (\( \overline x = {1}0\;\mu {\text{m}} \)), arising from the apex of the apical cell; filiform basal appendage.
Description from epitype (Fig. 6. a–g)
Conidiophores indistinct. Conidiogenous cells discrete, simple, short, filiform. Conidia 17–20 × 5.2–6.6 μm (\( \overline x = {19} \times {6}\;\mu {\text{m}} \)), fusiform to ellipsoid, straight to slightly curved, 4-septate, basal cell short, obtuse, hyaline, thin-walled and verruculose, 3–3.8 μm long (\( \overline x = {3}.{3}\;\mu {\text{m}} \)); with three median cells, doliiform to subcylindrical, concolorous, olivaceous, septa and periclinal walls darker than the rest of the cell, together 12.5–14.2 μm long (\( \overline x = {13}.{6}\;\mu {\text{m}} \)) second cell from base 4.4–5.5 μm (\( \overline x = {4}.{9}\;\mu {\text{m}} \)); third cell 4.3–5 μm (\( \overline x = {4}.{5}\;\mu {\text{m}} \)); fourth cell 4–4.8 μm (\( \overline x = {4}.{3}\;\mu {\text{m}} \)); apical cell hyaline, conic, 2.7–3.7 μm long (\( \overline x = {3}.{2}\;\mu {\text{m}} \)); with two to three appendages 6–14 (\( \overline x = {1}0\;\mu {\text{m}} \)) μm long, arising from the apex of the apical cell; filiform basal appendage.
Colonies on PDA attaining 7 cm diam. after 7 days at 25 °C, with undulate edge, whitish, with dense, aerial mycelium on surface; fruiting bodies black, gregarious; reverse of the colony yellowish.
Habitat/Distribution: Inhabiting leaves of Prunus cerasus, USA, refrigerator door PVC gasket, Fiji and Syzygium sp., Thailand.
Material examined: USA, Newfield, New Jersey, on leaves of Prunus cerasus L., cultivated plum, 20 July 1887 (NY 00937391, holotype); FIJI, on refrigerator door PVC gasket, 1 June 1978, E.H.C. McKenzie (MFLU12-0425, epitype designated here; ex-type living culture ICMP 6088 = PDDCC 6088).
Additional culture examined: Thailand, Chiang Rai, on leaves of Syzygium sp., 06 Febuary 2010, S.S.N. Maharachchikumbura SS008 (MFLUCC 10-0146).
Notes: Pestalotiopsis adusta was described from cultivated plum in New Jersey (Steyaert 1949) and recently phenolic compounds isolated from one putative isolate of P. adusta showed antimicrobial activity against Fusarium culmorum, Gibberella zeae and Verticillium aiboatrum (Li et al. 2008). Pestalotiopsis adusta is characterized by its small conidia (16–20 × 5–7 μm) and two to three relatively short apical appendages (7–15 μm) (Fig. 5 e–g). According to Guba (1961), P. adusta occurs on various hosts and has a cosmopolitan distribution. Guba (1961) listed it from Acer platanoides in Point Pleasant, New Jersey; on stems of Barringtonia speciosa in Bermuda; from circular spots on leaves of Bischofia javanica in Taiwan; on leaves of Carpinus betulus in Italy; as causing fruit rot and grey leaf spot in Eriobotrya japonica in Japan; on leaves of Homalomena philippinensis in the Philippines; and on spots and dead areas of leaves of Pavonia multiflora in Brazil. Living specimens from cultivated plum or from the USA would have been desirable when epitypifying this taxon. The sample collected from Fiji, however, is characteristic of P. adusta, a distinct species in the genus. The epitype has identical conidiogenous cells and morphology, including three apical appendages and a spore size fitting that of the holotype. As we want to advance the understanding of this poorly defined species rich genus, the Fiji collection is designated here as an epitype of P. adusta.
Pestalotiopsis asiatica Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800529
Figure 7 a–j.
Etymology: The specific epithet is based on the geographical region (Asia), in reference to where fungus was isolated.
Conidiophores indistinct. Conidiogenous cells hyaline, simple, filiform, 3–12 μm long. Conidia 20–26 × 5–7 μm (\( \overline x = {22}.{6} \times {6}.{25}\;\mu {\text{m}} \)), fusiform, straight to slightly curved, 4-septate; basal cell conical, hyaline, thin and verruculose, 3–5 μm long (\( \overline x = {4}\;\mu {\text{m}} \)); three median cells 13–15.5 μm long (\( \overline x = {14}\;\mu {\text{m}} \)), dark brown, verruculose, septa and periclinal walls darker than the rest of the cell, versicoloured, second cell from base pale brown, 4–5.5 μm (\( \overline x = {4}.{5}\;\mu {\text{m}} \)); third cell darker brown, 4–5 μm (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); fourth cell darker, 4–5 μm (\( \overline x = {4}.{7}\;\mu {\text{m}} \)); apical cell 3.5–5 μm long (\( \overline x = {3}.{35}\;\mu {\text{m}} \)), hyaline, conical to cylindrical, comprising 2–4 appendages (mainly 3); apical appendages 20–30 μm long (\( \overline x = {25}.{6}\;\mu {\text{m}} \)), tubular, arising from the apex of the apical cell; basal appendage, 4–8 μm long (\( \overline x = {5}.{65}\;\mu {\text{m}} \)), filiform.
Colonies on PDA reaching 7 cm diam. after 6 days at 25 °C, with crenate edge, whitish, with aerial mycelium on surface; fruiting bodies black, gregarious; reverse of culture whitish to pale yellow.
Habitat/Distribution: Endophyte on unidentified tree, Yizhang County, Hunan Province China.
Material examined: CHINA, Hunan Province, Yizhang County, Mangshan, isolated from living leaves of unidentified tree, 12 April 2002, Wenping Wu HN51-1 (HMAS047638, holotype; MFLU12-0422, isotype; ex-type living culture NN047638 = MFLUCC 12-0286).
Notes: Pestalotiopsis asiatica is a distinct species in the versicolour group (Clade B) and clearly distinguishable from P. chrysea and P. umberspora in the β-tubulin, tef1 and combined genes phylogram. Pestalotiopsis asiatica (20–26 × 5–7 μm) is morphologically similar to P. pauciseta (Sacc.) Y.X. Chen (conidia 20–24 × 4.5–5 μm) (Saccardo 1914) and P. gracilis (Kleb.) Steyaert (conidia 19–23 × 6–7 μm) (Saccardo 1931). However, P. asiatica differs from P. pauciseta by its wider conidia and from P. gracilis in having long apical appendages (in P. gracilis 10–26 μm).
Pestalotiopsis chinensis Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800522
Figure 8 a–h.
Etymology: The specific epithet is referring to China, the country from where the taxon was isolated.
Conidiophores most often indistinct, septate, hyaline, smooth, rarely branched. Conidiogenous cells discrete, ampulliform to lageniform, smooth, thin-walled, hyaline or pale brown, with 2–3 proliferations. Conidia fusoid to ellipsoid, straight to slightly curved, 4–septate, 23–32 × 7–9 μm (\( \overline x = {29} \times {8}.{3}\;\mu {\text{m}} \)), basal cell conic to obconic, hyaline or slightly olivaceous, thin-walled and verruculose, 5–7 μm long (\( \overline x = {5}.{7}\;\mu {\text{m}} \)), with three median cells, doliiform to cylindrical, constricted at the septa, concolorous, olivaceous, septa and periclinal walls darker than the rest of the cell, wall rugose, together 20–22 μm long (\( \overline x = {2}0.{2}\;\mu {\text{m}} \)) second cell from base 6–7 μm (\( \overline x = {6}.{5}\;\mu {\text{m}} \)); third cell 7–7.5 μm (\( \overline x = {7}.{1}\;\mu {\text{m}} \)); fourth cell 6–7.5 μm (\( \overline x = {6}.{8}\;\mu {\text{m}} \)); apical cell hyaline, conic to subcylindrical, 3–6 μm long (\( \overline x = {4}.{3}\;\mu {\text{m}} \)); with 1–3 tubular apical appendages (mostly 3), arising from the apex of the apical cell, 25–30 μm long (\( \overline x = {28}\;\mu {\text{m}} \)), unequal; basal appendage present 7–11 μm (\( \overline x = {8}.{7}\;\mu {\text{m}} \)).
Colonies on PDA reaching 7 cm diam. after 13 days at 25 °C, with edge crenate, whitish to pale yellow, with dense aerial mycelium on surface, fruiting bodies black, developing in concentric circles; reverse of culture yellow to pale orange.
Habitat/Distribution: Endophyte in leaves of Taxus sp., Kunming, Yunnan Province China.
Material examined: CHINA, Yunnan Province, Kunming, Kunming Botanical Garden, on living leaves of Taxus sp., 19 March 2002, Wenping Wu KBG13-9 (HMAS047218, holotype; MFLU12-0415, isotype; ex-type living culture NN047218 = MFLUCC 12-0273).
Notes: The conidial size of Pestalotiopsis chinensis overlaps with P. funerea (Desm.) Steyaert (21–29 × 7–9.5 μm) (Steyaert 1949), P. macrochaeta (Speg.) J. Xiang Zhang & T. Xu (22–31 × 8–10 μm) (Zhang et al. 2002), P. mayumbensis (Steyaert) Steyaert (22–28 × 6.5–8.5 μm) (Steyaert 1949) and P. osyridis (Thüm.) H.T. Sun & R.B. Cao (22–28 × 5–7 μm) (Guba 1961). However, P. chinensis can be distinguished from P. mayumbensis and P. osyridis by its relatively large conidial size and also by its long apical appendages (in P. mayumbensis 8–15 μm and in P. osyridis up to 14 μm). Pestalotiopsis chinensis (1–3 tubular apical appendages) can be differentiated by the number of apical appendages (3–6 apical appendages (mostly 4–5) in P. funerea and three apical appendages in P. macrochaeta). It was not possible to obtain PCR products of P. chinensis for β-tubulin and tef1 and thus in phylogenetic tree it clusters with P. verruculosa, which is morphologically distinct.
Pestalotiopsis chrysea Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800533
Figure 9 a–g.
Etymology: The specific refers to the golden yellow colour of the colony (Latin- chryseus) of this species.
Conidiophores indistinct. Conidiogenous cells discrete or integrated, lageniform, hyaline, smooth-walled. Conidia 20–24 × 5.5–7 μm (\( \overline x = {22}.{3} \times {6}.{1}\;\mu {\text{m}} \)), fusiform, straight to slightly curved, 4-septate; basal cell obconic to conic, hyaline, thin and smooth-walled, 3–5 μm long (\( \overline x = {4}.{3}\;\mu {\text{m}} \)); three median cells 14–16 μm long (\( \overline x = {14}.{8}\;\mu {\text{m}} \)), dark brown to olivaceous, septa and periclinal walls darker than the rest of the cell, versicoloured, verruculose, second cell from base pale brown, 4–5 μm (\( \overline x = {4}.{6}\;\mu {\text{m}} \)); third cell darker brown, 4–5 μm (\( \overline x = {4}.{6}\;\mu {\text{m}} \)); fourth cell darker, 4–5 μm (\( \overline x = {4}.{5}\;\mu {\text{m}} \)); apical cell 3.5–4.5 μm long (\( \overline x = {4}\;\mu {\text{m}} \)), hyaline, conic to obconic; apical appendages 22–30 μm long (\( \overline x = {26}.{8}\;\mu {\text{m}} \)), 3, tubular, arising from the apex; basal appendage, 3 –6 μm long (\( \overline x = {4}.{4}\;\mu {\text{m}} \)), filiform.
Colonies on PDA reaching 7 cm diam. after 10 days at 25 °C, edge irregular, yellowish to pale brown, aerial mycelium on surface, fruiting bodies black, gregarious; reverse of the colony orange to brown.
Habitat/Distribution: Saprobe on dead plant material, Guangxi and Hunan provinces, China.
Material examined: CHINA, Guangxi Province, Shangsi, Shiwandashan, Wangle, dead leaves of unidentified plant, 2 January 1997, Wenping Wu WUFH1303a (HMAS042855, holotype; MFLU12-0411, isotype; ex-type living culture NN042855 = MFLUCC 12-0261).
Additional culture examined: CHINA, Hunan Province, Yizhang County, Mangshanon dead plant material, 12 April 2002, Wenping Wu HN27-10 (NN047037 = MFLUCC 12-0262).
Notes: Pestalotiopsis chrysea is a morphologically distinct species in the genus with its yellowish colony; its conidiogenous cells and conidia are also slightly yellowish. It can clearly be differentiated from its phylogenetically related sibling species, P. umberspora (19–29 × 6–8 μm) in having relatively narrow conidia (20–24 × 5.5–7 μm) and also in tef1 and combined genes phylogenetic trees (Figs. 3 and 4).
Pestalotiopsis clavata Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800524
Figure 10 a–f.
Etymology: In Latin, clavatus refers to the clavate conidia.
Conidiophores most often indistinct. Conidiogenous cells discrete ampulliform to lageniform, smooth, thin-walled, hyaline, short. Conidia fusoid to ellipsoid, straight to slightly curved, 4-septate, 20–27 × 6.5–8 μm (\( \overline x = {22}.{6} \times {7}.{3}\;\mu {\text{m}} \)), basal cell conic to obconic with obtuse end, hyaline, thin-walled and verruculose, 4–5 μm long (\( \overline x = {4}.{6}\;\mu {\text{m}} \)), with three median cells, doliiform, concolorous, olivaceous to brown, septa and periclinal walls darker than the rest of the cell; wall rugose, together 15–16 μm long (\( \overline x = {15}.{2}\;\mu {\text{m}} \)) second cell from base 5–6 μm (\( \overline x = {5}.{2}\;\mu {\text{m}} \)); third cell 4–5 μm (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); fourth cell 5–5.5 μm (\( \overline x = {5}.{2}\;\mu {\text{m}} \)); apical cell hyaline, conic to cylindrical 3–5 μm long (\( \overline x = {3}.{75}\;\mu {\text{m}} \)), with 2– 3 tubular apical appendages (mostly 3) arising from the apex of the apical cell, 20–25 μm long (\( \overline x = {23}\;\mu {\text{m}} \)); basal appendage mostly present, 7–9 μm long (\( \overline x = {7}.{8}\;\mu {\text{m}} \)).
Colonies on PDA reaching 7 cm diam. after 8 days at 25 °C, with edge entire, whitish to pale brown, with dense, aerial mycelium on the surface, with black fruiting bodies; reverse of culture pale brown to brown.
Habitat/Distribution: Endophyte in living leaves of Buxus sp. and Euonymus sp., Hunan and Yunnan provinces, China.
Material examined: CHINA, Yunnan Province, Kunming, Kunming Botanical Garden, living leaf of Buxus sp., 19 March 2002, Wenping Wu KBG26-5 (HMAS047134, holotype; MFLU12-0412, isotype; ex-type living culture NN047134 = MFLUCC 12-0268).
Additional culture examined: CHINA, Hunan Province, Yizhang County, Mangshan, living leaf of Euonymus sp., 12 April 2002, Wenping Wu HN49-6 (NN047005 = MFLUCC 12-0269)
Notes: Pestalotiopsis clavata is a distinct species recognized based on its morphology and phylogeny. It has similar sized conidia to P. heterocornis (Guba) Y.X. Chen (18–26 × 6.5–8 μm) (Guba 1961). However, these species are distinct in the length and number of their apical appendages. P. clavata has conidia with 2–3 apical appendages (mostly 3) which are 20–25 μm long, while in P. heterocornis the apical appendages are unequal in length being 9–21 μm long (Guba 1961). Pestalotiopsis carveri (Guba) P.L. Zhu, Q.X. Ge & T. Xu (20–26 × 6–7 μm) (Guba 1961) also has a somewhat similar conidial morphology with P. clavata, but they differ in the length and number of their apical appendages. In P. carveri the two apical appendages are unequal in length being 12–26 μm long.
Pestalotiopsis clavispora (G.F. Atk.) Steyaert, Bull. Jard. bot. État Brux. 19: 335 (1949)
Basionym: Pestalotia clavispora G.F. Atk., Bulletin of Cornell University 3: 37 (1897)
MycoBank: MB289191
Description from holotype (Fig. 11 a–h)
Conidiomata 150–250 μm in diam., black, numerous, scattered, rupturing the epidermis and dehiscing irregularly. Conidia 18–26 × 6.5–8.5 μm (\( \overline x = {21} \times {7}.{5}\;\mu {\text{m}} \)), fusiform, 4-septate, straight or slightly curved and clavate-fusiform; basal cell long and conic, hyaline, thin and verruculose, 4–5 μm long (\( \overline x = {4}.{2}\;\mu {\text{m}} \)); with three median cells 13.7–15.3 μm long (\( \overline x = {14}.{7}\;\mu {\text{m}} \)), dark brown to olivaceous, septa and periclinal walls darker than the rest of the cell, versicolored, verruculose, second cell from base pale brown, 4.3–5.3 μm (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); third cell darker brown, 5.5–6.4 μm (\( \overline x = {5}.{8}\;\mu {\text{m}} \)); fourth cell darker, 4.5–5.8 μm (\( \overline x = {5}\;\mu {\text{m}} \)); apical cell 3.3–4.2 μm long (\( \overline x = {3}.{7}\;\mu {\text{m}} \)), short, broad conic, hyaline, subcylindric; with apical appendages 19–30 μm long (\( \overline x = {24}.{5}\;\mu {\text{m}} \)), tubular, 2–3 (rarely 2), arising from the apex of the apical cell; with basal appendage present, filiform.
Description from epitype (Fig. 12 a–g)
Conidiophores indistinct. Conidiogenous cells hyaline, simple, short or relatively long, filiform, 4–10 um long. Conidia 20–24 × 6–8 μm (\( \overline x = {22} \times {7}.{2}\;\mu {\text{m}} \)), fusiform, straight to slightly curved, 4-septate, clavate-fusiform when mature; basal cell conical, hyaline, thin and verruculose, 3–5 μm long (\( \overline x = {3}.{8}\;\mu {\text{m}} \)); three median cells 13–15 μm long (\( \overline x = {13}.{9}\;\mu {\text{m}} \)), dark brown to olivaceous, verruculose-walled, septa and periclinal walls darker than the rest of the cell, versicoloured, second cell from base pale brown, 4–5 μm (\( \overline x = {4}.{5}\;\mu {\text{m}} \)); third cell darker brown, 4–5 μm (\( \overline x = {4}.{6}\;\mu {\text{m}} \)); fourth cell darker, 4–5 μm (\( \overline x = {4}.{5}\;\mu {\text{m}} \)); apical cell 3–5 μm long (\( \overline x = {4}.{3}\;\mu {\text{m}} \)), hyaline, subcylindric; with apical appendages 22–32 μm long (\( \overline x = {26}.{5}\;\mu {\text{m}} \)), tubular, 2–3 (rarely 2), arising from apex of the apical cell; with basal appendage, 3–5.5 μm (\( \overline x = {4}\;\mu {\text{m}} \)), filiform.
Colonies on PDA reaching 7 cm diam. after 7 days at 25 °C, edge undulate, whitish, aerial mycelium on surface, fruiting bodies black, concentric; reverse of culture pale luteous.
Habitat/Distribution: Known to inhabit in Quercus rubra in USA and Magnolia sp. in China.
Material examined: USA, Auburn, Alabama, on fallen leaves of Quercus rubra L., 10 March 1891, F. Atkinson (CUP-A-032389, holotype); CHINA, Guangxi Province, Shiwandashan, on dead leaves of Magnolia sp., 28 Dec 1997, Wenping Wu WUFH1486c (HMAS043133 = MFLU12-0418, epitype designated here; ex-type living culture NN043133 = MFLUCC 12-0281).
Additional culture examined: CHINA, Yunnan Guangxi Province, Shiwandashan, on dead leaves of Magnolia sp., 28 December 1997, Wenping Wu (NN043011 = MFLUCC 12-0280)
Notes: Pestalotiopsis clavispora is known as a plant pathogen but has been isolated as a common endophyte in recent studies (Keith et al. 2006; Espinoza et al. 2008; Liu et al. 2007; Wei et al. 2007). The holotype of P. clavispora was recorded from fallen leaves of Quercus rubra, in Auburn, Alabama, USA. In addition, P. clavispora has been recorded from leaves of black oak, Quercus minima and on fruit husks and leaves of Aleurites fordii grows in different parts of USA and on living leaves of Bruchellia bubalina in South Africa (Guba 1961). Thus, P. clavispora appears to have a wide host range and distribution. Since no ex-type culture is available for this species, an epitype with a living culture is designated from a sample collected in Guangxi Province, China. We would prefer to choose an epitype from USA and the original host however, in order to expedite the understanding of this poorly resolved genus, we designate an epitype which has conidial characters (length, width and length of apical appendages) fitting that of the holotype. The present material is a good match for P. clavispora.
Pestalotiopsis diversiseta Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800526
Figure 13 a–g.
Etymology: The specific epithet is based on the diverse arrangement, Latin = diversisetae of the apical appendages.
Conidia fusoid to ellipsoid, straight to slightly curved, 4–septate, 27–34 × 5.5–8 μm (\( \overline x = {29}.{7} \times {6}.{3}\;\mu {\text{m}} \)), with basal cell obconic and obtuse at the base, hyaline, thin-walled and verruculose, 3–6 μm long (\( \overline x = {4}.{5}\;\mu {\text{m}} \)), with three median cells, doliiform, concolorous, olivaceous, septa and periclinal walls darker than the rest of the cell, wall rugose, together 17–21 μm long (\( \overline x = {19}\;\mu {\text{m}} \)) second cell from base 5–7 μm (\( \overline x = {5}.{8}\;\mu {\text{m}} \)); third cell 6–8 μm (\( \overline x = {6}.{8}\;\mu {\text{m}} \)); fourth cell 6–7 μm (\( \overline x = {6}.{3}\;\mu {\text{m}} \)); apical cell hyaline, cylindrical 4–7 μm long (\( \overline x = {6}\;\mu {\text{m}} \)); with 3–5 tubular appendages (rarely 2); some appendages branched, slightly swollen at the tip, arising from the apex of the apical cell and sometimes arising from the different parts of the apical cell, 22–30 μm long (\( \overline x = {26}\;\mu {\text{m}} \)); with basal appendage 5–9 μm long, rarely absent.
Colonies on PDA reaching 7 cm diam. after 8 days at 25 °C, edge fimbriate, whitish, with dense, aerial mycelium on surface, with black fruiting bodies, gregarious; reverse of the culture white.
Habitat/Distribution: Endophyte on living leaf of Rhododendron sp., Yunnan Province, China.
Material examined: CHINA, Yunnan Province, Kunming, Kunming Botanical Garden, living leaves of Rhododendron sp., 19 March 2002, Wenping Wu HN26-5 (HMAS047261, holotype; MFLU12-0423, isotype; ex-type living culture NN047261 = MFLUCC 12-0287). Table 6
Notes:
Pestalotiopsis diversiseta is a morphologically distinct species, also shown in its DNA phylogeny. However, it has an overlapping conidial size with P. leucopogonis, P. perseae and P. theae (Guba 1961; Nag Rag 1993). Pestalotiopsis diversiseta can be differentiated from all these species by its morphological distinction. P. diversiseta has 3–5 apical appendages which differ from P. leucopogonis (7–11 apical appendages), P. perseae (2–4 apical appendages) and P. theae (7–11 apical appendages) (Guba 1961; Nag Rag 1993). Its apical appendages are also knobbed unlike those in P. leucopogonis (Nag Rag 1993). Although P. diversiseta and P. perseae have knobbed apical appendages with similar attachment to the apical cell, and length (22–30 vs 10–23 μm), the species can be distinguished by its concolorous median cells which in P. perseae are versicoloured with irregular longitudinal ridges (Guba 1961; Nag Rag 1993).
Pestalotiopsis ellipsospora Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800528
Figure 14 a–h.
Etymology: The specific epithet is based on the ellipsoid shape, Latin = ellipsospora, of the conidia.
Conidia 19–25 × 5–6.5 μm (\( \overline x = {21}.{7} \times {6}\;\mu {\text{m}} \)), fusiform, straight to slightly curved, 4-septate; with basal cell conical with obtuse end, hyaline, thin and smooth-walled, 4–5 μm long (\( \overline x = {4}.{3}\;\mu {\text{m}} \)); with three median cells 13–15 μm long (\( \overline x = {14}.{1}\;\mu {\text{m}} \)), dark brown, septa and periclinal walls darker than the rest of the cell, versicoloured, second cell from base pale brown, 4–5 μm (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); third cell darker brown, 4–5 μm (\( \overline x = {4}.{7}\;\mu {\text{m}} \)); fourth cell darker, 4–5 μm (\( \overline x = {4}.{5}\;\mu {\text{m}} \)); apical cell 3–4 μm long (\( \overline x = {3}.{8}\;\mu {\text{m}} \)), hyaline, conical; with apical appendages 5–12 μm long (\( \overline x = {8}\;\mu {\text{m}} \)), tubular, 1–3, arising from the apex of the apical cell; basal appendage small or absent, 3–4 μm long (\( \overline x = {3}.{4}\;\mu {\text{m}} \)), filiform.
Colonies on PDA reaching 7 cm diam. after 6 days at 25 °C, edge crenate, whitish, with aerial mycelium on the surface, with black, gregarious fruiting bodies; reverse of the culture white.
Habitat/Distribution: Saprobe on dead plant material in Yunnan Province, China and Chiang Rai Province Thailand.
Material examined: CHINA, Yunnan Province, on dead plant materials, Guo Liang-Dong Guo986 (MFLU12-0420; holotype; ex-type living culture MFLUCC 12-0283).
Additional culture examined: THAILAND, Chiang Rai, Tool Kwan, Huay Mesak waterfall, on dead plant material, 12 January 2010, S.S.N Maharachchikumbura (MFLUCC 12-0280)
Notes: Pestalotiopsis ellipsospora (conidia 19–25 × 5–6.5 μm) can be morphologically distinguished from its phylogenetically closely related species, P. samarangensis (conidia 18–21 × 6.5–7.5 μm) (Maharachchikumbura et al. 2012b). Pestalotiopsis samarangensis has three long apical appendages (12–18 μm long) whereas in P. ellipsospora the 1–3 appendages are shorter (5–12 μm).
Pestalotiopsis foedans (Sacc. & Ellis) Steyaert, Bull. Jard. bot. État Brux. 14: 329 (1949) Basionym: Pestalotia foedans Sacc. & Ellis, Michelia 2(no. 8): 575 (1882)
MycoBank: MB289196
Description from holotype (Fig. 15 a–h)
Conidiomata acervuli, with basal stroma and lateral wall 1–3 cells thick; the wall cells pale brown, textura angularis, 200– 400 × 150– 300 μm. Conidiophores reduced to conidiogenous cells arising in the concavity of acervuli. Conidiogenous cells discrete, simple, short, filiform. Conidia 19–24 × 5.7–6.9 μm (\( \overline x = {2}0.{7} \times {6}.{4}\;\mu {\text{m}} \)), fusiform to ellipsoid, straight to slightly curved, 4-septate; basal cell conic, hyaline, thin and smooth-walled, 3.2–4.5 μm long (\( \overline x = {4}\;\mu {\text{m}} \)); three median cells 12.5–14.6 μm long (\( \overline x = {14}\;\mu {\text{m}} \)), hyaline, versicoloured, verruculose; second cell from base pale brown to olivaceous, 4.3–5.7 μm (\( \overline x = {4}.{9}\;\mu {\text{m}} \)); third cell darker brown to olivaceous, 4.7–6 μm (\( \overline x = {5}\;\mu {\text{m}} \)); fourth cell darker, 4.5–5 μm (\( \overline x = {4}.{7}\;\mu {\text{m}} \)); apical cell 4–5 μm long (\( \overline x = {4}.{3}\;\mu {\text{m}} \)), hyaline, cylindric to subcylindric; apical appendages 6–18 μm long (\( \overline x = {13}.{3}\;\mu {\text{m}} \)), 2–3 (mostly 3), arising from the apex of the apical cell; basal appendage present (rarely absent), filiform 3–5 μm (\( \overline x = {4}\;\mu {\text{m}} \)).
Description from epitype (Fig. 16 a–h)
Conidiophores indistinct, arising in the concavity of acervuli. Conidiogenous cells discrete, simple, short, filiform, 2–4 um. Conidia 19.2–23.4 × 5.5–7 μm (\( \overline x = {2}0.{6} \times {6}.{7}\;\mu {\text{m}} \)), fusiform to ellipsoid, straight to slightly curved, 4-septate; basal cell conic, hyaline, thin and smooth-walled, 3.2–5 μm long (\( \overline x = {4}.{4}\;\mu {\text{m}} \)), three median cells hyaline, versicoloured, verruculose, 12.7–15.3 μm long (\( \overline x = {13}.{7}\;\mu {\text{m}} \)); second cell from base pale brown to olivaceous, 4.1–5.2 μm (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); third cell darker brown to olivaceous, 4.7–5.3 μm (\( \overline x = {5}\;\mu {\text{m}} \)); fourth cell darker, 4.9–5.7 μm (\( \overline x = {5}.{3}\;\mu {\text{m}} \)); apical cell 4–5 μm long (\( \overline x = {4}.{3}\;\mu {\text{m}} \)), hyaline, cylindric to subcylindric; apical cell hyaline, subcylindric to conic 3.3–4.4 μm (\( \overline x = {3}.{7}\;\mu {\text{m}} \)); apical appendages 8–15 μm long (\( \overline x = {12}.{6}\;\mu {\text{m}} \)), 2–3 (mostly 3), arising from the apex of the apical cell; basal appendage present (rarely absent), filiform, 3–6 μm long (\( \overline x = {4}.{3}\;\mu {\text{m}} \)).
Colonies on PDA reaching 7 cm diam. after 6 days at 25 °C, edge undulate, whitish, aerial mycelium on surface, with black fruiting bodies, gregarious; reverse of culture whitish (rarely pale luteous).
Habitat/Distribution: Saprobe on bark of Thuja occidentalis, USA and mangrove leaves, Calliandra haematocephala and Neodypsis decaryi, China.
Material examined: USA, Newfield, New Jersey, on decaying bark of white cedar, Thuja occidentalis L., October 1880, Ellis and Harkness (BPI 0405695, holotype); CHINA, Xinglong, Hainan, on mangrove leaves, April 2005, A.R. Liu L443 (MFLU 12-0424, epitype designated here; extype living culture-CGMCC 3.9123).
Additional culture examined: CHINA, Xinglong, Hainan, on leaves of Calliandra haematocephala, May 2004, A.R. Liu L101 (CGMCC 3.9202); CHINA, Xinglong, Hainan, on leaves of Neodypsis decaryi, May 2004, A.R. Liu L96 (CGMCC 3.9178).
Notes: The holotype of P. foedans was recorded from decaying bark of white cedar, in New Jersey, USA. In addition, P. foedans was recorded from Cupressus thyoides in New Jersey, USA; on Cryptomeria japonica in Philadelphia and Japan; leaves and twigs of C. japonica in Princeton and on needles of Pinus mugo in Pennington (Guba 1961). Thus, P. foedans appears to have a wide host range and distribution. Recently, P. foedans was discovered as a source of bioactive metabolites of high economic importance (Ding et al. 2008). Since no ex-type culture is available for this species, an epitype with a living culture is designated from a sample collected in Hainan Province, China. We would prefer to choose an epitype from USA and the original host however, in order to expedite the understanding of this poorly resolved genus we choose to be pragmatic and designated an epitype which has conidial characters (length, width and length of apical appendages) similar to that of the holotype and hence this is a good match for P. foedans.
Pestalotiopis inflexa Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800530
Figure 17 a–i.
Etymology: From the Latin, inflexus in reference to the curved nature of the conidia.
Conidiophores most often reduced to conidiogenous cells, simple, hyaline, smooth-walled. Conidiogenous cells discrete, ampulliform to lageniform, smooth, thin-walled, hyaline or pale olivaceous. Conidia fusoid to ellipsoid, straight to slightly curved, 4-septate, 24–31 × 6–9 μm (\( \overline x = {27} \times {7}.{6}\;\mu {\text{m}} \)), basal cell conic to obconic, hyaline or slightly olivaceous, thin-walled and verruculose, 5–7 μm long (\( \overline x = {5}.{7}\;\mu {\text{m}} \)), with 3 median cells, doliiform to cylindrical, with thick verruculose walls, constricted at the septa, concolorous, olivaceous, with septa and periclinal walls darker than the rest of the cell, wall rugose, together 15–19 μm long (\( \overline x = {17}.{1}\;\mu {\text{m}} \)) second cell from base 5–7 μm (\( \overline x = {5}.{7}\;\mu {\text{m}} \)); third cell 5–7 μm (\( \overline x = {5}.{8}\;\mu {\text{m}} \)); fourth cell 4.5–6 μm (\( \overline x = {5}.{3}\;\mu {\text{m}} \)); apical cell hyaline, subcylindrical to cylindrical 4–5 μm long (\( \overline x = {4}.{6}\;\mu {\text{m}} \)); 2–5 tubular apical appendages (mostly 3–4), often arising from the apex of the apical cell or rarely arising from just below the apex of apical cell, 20–30 μm long (\( \overline x = {24}\;\mu {\text{m}} \)), unequal, rarely branched; basal appendage present, relatively long 9–15 μm (\( \overline x = {12}\;\mu {\text{m}} \)).
Colonies on PDA reaching 7 cm diam. after 18 days at 25 °C, edge undulate, whitish, with dense, aerial mycelium on surface, with black, gregarious fruiting bodies; reverse of the culture yellowish.
Habitat/Distribution: Endophyte in living leaves of unidentified plant, Hunan Province, China.
Material examined: CHINA, Hunan Province, Yizhang County, Mangshan, living leaf of unidentified tree, 12 April 2002, Wenping Wu HN14-2 (HMAS047098, holotype; MFLU12-0413, isotype; ex-type living culture NN047098 = MFLUCC 12-0270).
Notes: Pestalotiopsis inflexa can be differentiated from its close relatives in the β-tubulin, tef1 and combined phylogram in Clade A. The characteristic morphology of P. inflexa is due to its divergent, 2 to 5 apical appendages, sometimes arising from the middle of apical cell and by a relatively long basal appendage (9–15 μm). Morphologically similar species to P. inflexa in conidial size is P. thujicola (J.L. Maas) Y. Suto & Tak. Kobay (25–31 × 6.5–10 μm) (Maas 1971). However, P. thujicola can be differentiated by its 3–6 apical appendages radiating from different parts of the apical cell. In P. inflexa, the appendages usually arise from the tip of the apical cell and rarely from the middle.
Pestalotiopsis intermedia Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800532
Figure 18 a–h.
Etymology: From Latin, intermediate pertaining to the intermediate size of the conidia.
Conidiophores indistinct. Conidiogenous cells discrete, simple, filiform, smooth, thin-walled, hyaline, and short. Conidia fusoid to ellipsoid, straight to slightly curved, 4-septate, 24–28 × 5.5–6.5 μm (\( \overline x = {25}.{7} \times {6}\;\mu {\text{m}} \)), basal cell conic to obconic with obtuse end, hyaline, thin- and verruculose, 4–5 μm long (\( \overline x = {4}.{8}\;\mu {\text{m}} \)), with three median cells, doliiform, concolorous, olivaceous to brown, septa and periclinal walls darker than the rest of the cell, wall rugose, together 15–19 μm long (\( \overline x = {17}\;\mu {\text{m}} \)) second cell from base 5–6 μm (\( \overline x = {5}.{7}\;\mu {\text{m}} \)); third cell 5–6 μm (\( \overline x = {5}.{7}\;\mu {\text{m}} \)); fourth cell 5–6.5 μm (\( \overline x = {5}.{2}\;\mu {\text{m}} \)); apical cell hyaline, conic to cylindrical 4–5 μm long (\( \overline x = {4}.{5}\;\mu {\text{m}} \)); with 2–3 tubular apical appendages (rarely 4), arising from the apex of the apical cell, 10–28 μm long (\( \overline x = {18}.{5}\;\mu {\text{m}} \)), unequal; basal appendage present 6–10 μm (\( \overline x = {7}.{5}\;\mu {\text{m}} \)), rarely absent.
Colonies on PDA reaching 7 cm diam. after 6 days at 25 °C, edge entire, whitish, with dense, aerial mycelium on surface, fruiting bodies black; reverse of culture whitish to pale yellow.
Habitat/Distribution: Saprobe/endophyte on unidentified trees, Hubei and Yunnan provinces, China.
Material examined: CHINA, Hubei Province, Shengnongjia, on dead leave of unidentified tree, 24 March 2003, Wenping Wu WUFH7033 (HMAS047642, holotype; MFLU12-0410, isotype; ex-type living culture NN047642 = MFLUCC 12-0259).
Additional culture examined: CHINA, Yunnan Hunan Province, Yizhang County, Mangshan, on living leaf of unidentified plant, 12 April 2002, Wenping Wu HN28-16 (NN047073 = MFLUCC 12-0260).
Notes: The morphologically similar species to P. intermedia (24–28 × 5.5–6.5 μm) in conidial size are P. lespedezae (Syd.) Bilgrami (20–25 × 7–9 μm) (Guba 1961), P. osyridis (Thüm.) H.T. Sun & R.B. Cao (22–28 × 5–7 μm) and P. cocculi (Guba) G.C. Zhao & N. Li (22–29 × 5.5–7 μm) (Guba 1961). Pestalotiopsis intermedia can be differentiated from P. lespedezae by its long and thin conidia; and from P. osyridis and P. cocculi by its long apical appendages (P. osyridis usually has 3 apical appendages (rarely 2) measuring up to 14 μm long and in P. cocculi there are three apical appendages (sometimes 2), up to 11–12 μm long).
Pestalotiopsis jesteri Strobel, J.Yi Li, E.J. Ford & W.M. Hess, in Strobel, Li, Ford, Worapong, Baird & Hess, Mycotaxon 76: 260 (2000)
MycoBank: MB466231
Figure 19 a–j.
Conidiophores indistinct. Conidiogenous cells lageniform to subcylindrical, colourless, smooth, proliferation once or twice. Conidia fusoid to ellipsoid, straight to slightly curved, 4-septate, 23–29 × 5.5–7 μm (\( \overline x = {25} \times {6}.{1}\;\mu {\text{m}} \)), basal cell obconic, colorless, thin- and verruculose, 5–6 μm long (\( \overline x = {5}.{3}\;\mu {\text{m}} \)), with three median cells, subcylindrical, with thick verruculose walls, constricted at the septa, concolorous, pale brown, together 14–16.5 μm long (\( \overline x = {15}.{2}\;\mu {\text{m}} \)) second cell from base 4.5–6.5 μm (\( \overline x = {5}.{8}\;\mu {\text{m}} \)); third cell 4–6 μm (\( \overline x = {5}.{2}\;\mu {\text{m}} \)); fourth cell 5–6 μm (\( \overline x = {5}.{2}\;\mu {\text{m}} \)); apical cell colorless, obconic, acute at the apex, 4–5 μm long (\( \overline x = {4}.{3}\;\mu {\text{m}} \)); with 4 tubular appendages, 14–20 μm long, one arising from the apex and rest arising from just above the septum separating upper median and apical cell; basal appendage present, 4–5 μm long (\( \overline x = {4}.{5}\;\mu {\text{m}} \)).
Colonies on PDA reaching 7 cm diam. after 7 days at 25 °C, edge lobate, whitish yellow, with dense, aerial mycelium on surface, with black, gregarious fruiting bodies; reverse of the colony whitish yellow.
Habitat/Distribution: Endophyte on Fagraea bodenii, Papua New Guinea and saprobic on dead plant material, China.
Culture examined: CHINA, Yunnan Province, on dead plant material, Wenping Wu (NN042849 = MFLUCC 12-0279).
Notes: Pestalotiopsis jesteri has a similar morphology to that of P. montellica (Sacc. & Voglino) Tak. Kobay (Guba 1961) and may be a synonym. There are no sequence data available for P. montellica in GenBank, while ITS sequence data is available for the type of P. jesteri (Strobel et al. 2000). We therefore refrain from taking any action concerning synonymy at this stage. Pestalotiopsis jesteri is, however, distinct from all other species in the genus both in morphology and phylogeny. The attachment and arrangement of apical appendages at the apical cell are noticeably distinct.
Pestalotiopis linearis Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800531
Figure 20 a–h.
Etymology: The specific epithet is based on the linear shape of the conidia and in Latin, linear is linearis.
Conidiophores often reduced to conidiogenous cells, sometimes sparsely septate at the base and unbranched or branched, hyaline, smooth. Conidiogenous cells discrete ampulliform to lageniform, smooth, thin-walled, hyaline, with 1–2 proliferations, sometimes remain vegetative. Conidia fusiform, straight to slightly curved, 4-septate, 24–33 × 4.7–6 μm (\( \overline x = {29} \times {5}.{5}\;\mu {\text{m}} \)), basal cell conic to obconic, hyaline or slightly olivaceous, thin- and verruculose, 3.5–5.5 μm long (\( \overline x = {4}.{4}\;\mu {\text{m}} \)), with three median cells, doliiform to cylindrical, with thick verruculose walls, constricted at the septa, concolorous, olivaceous, septa and periclinal walls darker than the rest of the cell, together 17–21 μm long (\( \overline x = {19}\;\mu {\text{m}} \)) second cell from base 5–6.2 μm (\( \overline x = {5}.{5}\;\mu {\text{m}} \)); third cell 6–7 μm (\( \overline x = {6}.{3}\;\mu {\text{m}} \)); fourth cell 6–8 μm (\( \overline x = {6}.{6}\;\mu {\text{m}} \)); apical cell hyaline, cylindrical to subcylindrical 4–5 μm long (\( \overline x = {4}.{2}\;\mu {\text{m}} \)); with 2–3 tubular apical appendages (rarely 1), arising from the apex of the apical cell, 10–20 μm long (\( \overline x = {15}\;\mu {\text{m}} \)), unequal in length; basal appendage present, rarely two, 4–7 μm long (\( \overline x = {5}.{7}\;\mu {\text{m}} \)).
Colonies on PDA reaching 7 cm diam. after 6 days at 25 °C, edge entire, whitish, with dense, aerial mycelium on surface, with black, gregarious fruiting bodies; reverse of culture white.
Habitat/Distribution: Endophytes on living leaves of Trachelospermum sp. and Tsuga sp., Yunnan Province, China.
Material examined: CHINA, Yunnan Province, Kunming, Kunming Botanical Garden, on living leaves of Trachelospermum sp., 19 March 2002, Wenping Wu KBG14-3 (HMAS047190 holotype; MFLU12-0414, isotype; ex-type living culture NN047190 = MFLUCC 12-0271).
Additional culture examined: CHINA, Yunnan Province, Kunming, Kunming Botanical Garden, on living leaves of Tsuga sp., 19 March 2002, Wenping Wu KBG16-7 (NN047141 = MFLUCC 12-0272).
Notes: Pestalotiopsis linearis is a distinct species both in conidial morphology and phylogeny. It can be easily differentiated from its phylogenetically related species P. intermedia both in tef1 and combined trees (Figs. 3 and 4) and morphologically related species in the concolorous groups such as P. macrochaeta (Speg.) J. Xiang Zhang & T. Xu (22–31 × 8–10 μm) and P. caudata (Syd.) B. Sutton (22–31 × 8–10 μm) (Saccardo 1902). In P. linearis (24–33 × 4.7–6 μm) conidia are much thinner than these two species.
Pestalotiopsis rosea Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800521
Figure 21 a–k.
Etymology: The specific epithet is based on the Latin roseus in reference to the rose-colored, colony of this species
Conidiophores septate, unbranched, up to 20 μm long, often reduced to conidiogenous cells, smooth walled; Conidiogenous cells discrete, ampulliform to lageniform, smooth, thin-walled, slightly red, rarely hyaline, with 2–3 proliferations. Conidia fusoid to ellipsoid, straight to slightly curved, 4-septate, 17.5–21.8 × 5.7–7 μm (\( \overline x = {19}.{2} \times {6}.{2}\;\mu {\text{m}} \)), basal cell obconic, hyaline, thin- and verruculose, 3.1–4 μm long (\( \overline x = {3}.{6}\;\mu {\text{m}} \)), with three median cells, doliiform to subcylindrical, with thick verruculose walls, constricted at the septa, concolorous, olivaceous with slightly red, septa and periclinal walls darker than the rest of the cell, wall rugose, together 11.8–13.8 μm long (\( \overline x = {12}.{9}\;\mu {\text{m}} \)) second cell from base 4–5.3 μm (\( \overline x = {4}.{5}\;\mu {\text{m}} \)); third cell 3.3–5.1 μm (\( \overline x = {4}.{3}\;\mu {\text{m}} \)); fourth cell 4.2–5.4 μm (\( \overline x = {4}.{7}\;\mu {\text{m}} \)); apical cell hyaline, conic to cylindrical 2.6–4.2 μm long (\( \overline x = {3}.{3}\;\mu {\text{m}} \)); with 1–3 tubular apical appendages, some appendages branched, arising from the apex of the apical cell, 14–22 μm long (\( \overline x = {16}.{5}\;\mu {\text{m}} \)); basal appendage present 2–5.7 μm (\( \overline x = {4}.{1}\;\mu {\text{m}} \)), rarely absent.
Colonies on PDA reaching 7 cm diam. after 27 days at 25 °C, edge undulate, whitish or pale red, with dense, aerial mycelium on surface, with black to reddish brown fruiting bodies, gregarious; reverse of culture white or slightly red to red.
Habitat/Distribution: Endophyte on living leaves of Pinus sp., Yunnan Province, China.
Material examined: CHINA, Yunnan Province, Kunming, Kunming Botanical Garden, on living leaves of Pinus sp., 19 March 2002, Wenping Wu KBG25-3 (HMAS047135, holotype; MFLU12-0409, isotype; ex-type living culture NN047135 = MFLUCC 12-0258).
Notes: Pestalotiopsis rosea is a distinct species in the genus. The reddish colony is unique to the species and this can be found even in conidiogenous cells and in some conidia. This species was quite similar to the type species of the genus Pestalotiopsis; P. guepinii (Desm.) Steyaert (Guba 1961; Nag Rag 1993) isolated from Camellia japonica. In P. guepini, conidia are 14–21 × 5.5–6.6 μm, with 1–3 apical appendages that are sometimes knobbed at their apices. However, in P. rosea apical appendages are not knobbed and according to Guba (1961) P. guepinii is restricted to Camellia species.
Pestalotiopsis saprophyta Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800525
Figure 22. a–h.
Etymology: From the Latin saprophyta.
Conidiophores 0–1-septate, unbranched or irregularly branched, colorless, smooth-walled. Conidiogenous cells discrete or integrated, lageniform, subcylindric to cylindric, hyaline. Conidia 22–30 × 5–6 μm (\( \overline x = {24}.{9} \times {5}.{7}\;\mu {\text{m}} \)), fusiform, straight to slightly curved, 4-septate; basal cell conical to obtuse, hyaline, thin and smooth-walled, 4–7 μm long (\( \overline x = {5}\;\mu {\text{m}} \)); three median cells 14–20 μm long (\( \overline x = {15}.{5}\;\mu {\text{m}} \)), dark brown to olivaceous, septa and periclinal walls darker than the rest of the cell, versicoloured, verruculose, second cell from base pale brown to olivaceous, 4.5–7 μm (\( \overline x = {5}.{3}\;\mu {\text{m}} \)); third cell darker brown to dark olivaceous, 4–5 μm (\( \overline x = {4}.{7}\;\mu {\text{m}} \)); fourth cell darker, 4–6 μm (\( \overline x = {5}\;\mu {\text{m}} \)); apical cell 4–5 μm long (\( \overline x = {4}.{3}\;\mu {\text{m}} \)), hyaline, cylindric to subcylindric; apical appendages 23–35 μm long (\( \overline x = {27}.{3}\;\mu {\text{m}} \)), tubular, 2–4 (often 3), arising from the apex of the apical cell; basal appendage, 4–7 μm (\( \overline x = {6}\;\mu {\text{m}} \)), filiform.
Colonies on PDA reaching 7 cm diam. after 7 days at 25 °C, edge crenate, off white, aerial mycelium on surface, fruiting bodies black, gregarious; reverse of culture off white.
Habitat/Distribution: Saprobes on leaves of Magnolia sp., Yunnan Province, China.
Material examined: CHINA, Yunnan Province, Kunming, Kunming Botanical Garden, on leaves of Magnolia sp., 19 March 2002, Wenping Wu KBG29-2 (HMAS047136, holotype; MFLU12-0419, isotype; ex-type living culture NN047136 = MFLUCC 12-0282).
Notes: Pestalotiopsis saprophyta is a distinct species in the versicolour group (Clade B) with a higher conidial length to width ratio compared with other species. In β-tubulin and tef1 phylograms, it separates well with other species in the Clade B. P. saprophyta separates from its phylogenetic relative, P. foedans (19–25 × 5.5–7 μm) by having larger conidia (22–30 × 5–6 μm) and longer apical appendages (23–35 μm in P. saprophyta and 6–18 um in P. foedans). Other morphologically related species are P. batatas (Ellis & Everh.) G.C. Zhao & N. Li (23–28 × 7–8 μm) (Zhao and Li 1995), P. matildae (Richatt) S.J. Lee & Crous (22–32 × 6–8 μm) (Lee et al. 2006), and P. paeoniae (Servazzi) Steyaert (20–28 × 6–8 μm) (Guba 1961). However, in P. saprophyta conidia are thinner and apical appendages are longer.
Pestalotiopsis trachicarpicola Y. M. Zhang & K.D. Hyde in Zhang, Maharachchikumbura, McKenzie and Hyde, Cryptogamie mycol (in press) (2012a)
MycoBank: MB 564879
Figure 23 a–f.
Conidiophores indistinct. Conidiogenous cells discrete, simple, filiform, smooth, thin-walled, hyaline. Conidia fusoid to ellipsoid, broad-clavate, straight to slightly curved, 4-septate, 20–25 × 5.5–7.2 μm (\( \overline x = {23}.{5} \times {6}.{5}\;\mu {\text{m}} \)), basal cell conic to acute, hyaline, thin-walled and verruculose, 4–6 μm long (\( \overline x = {4}.{9}\;\mu {\text{m}} \)), with three median cells, doliiform to cylindrical, concolorous, olivaceous, verruculose, septa and periclinal walls darker than the rest of the cell, together 13–17 μm long (\( \overline x = {14}\;\mu {\text{m}} \)) second cell from base 4–6 μm (\( \overline x = {4}.{7}\;\mu {\text{m}} \)); third cell 4–6 μm (\( \overline x = {4}.{5}\;\mu {\text{m}} \)); fourth cell 4–6 μm (\( \overline x = {4}.{4}\;\mu {\text{m}} \)); apical cell hyaline, conic to subcylindrical 4–6 μm long (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); with 2–3 tubular apical appendages, arising from the apex of the apical cell, 9–18 μm long (\( \overline x = {13}.{6}\;\mu {\text{m}} \)); with a basal appendage, rarely two, 4–8 μm (\( \overline x = {6}.{3}\;\mu {\text{m}} \)) long.
Colonies fast growing on PDA, reaching 7 cm diam. after 6 days at 25 °C, edge fimbriate, yellowish white, dense, aerial mycelium on surface, fruiting bodies black; reverse of the culture yellowish white.
Habitat/Distribution Pathogen on Trachycarpus fortunei and saprobes on dead plant materials in China.
Material examined: CHINA, Yunnan Province, Kunming, Kunming Botanical Gardens, on leaf spots on living leaves of Trachycarpus fortunei, March 2011, K.D. Hyde OP068 (IFRD 9026,holotype; ex-type living culture IFRDCC 2440).
Additional culture examined: CHINA, Hunan Province, Yizhang County, Mangshan, on living leaf of unidentified tree, 12 April 2002, Wenping Wu HN53-2 (NN047072 = MFLUCC 12-0263); CHINA, Yunnan Province, Kunming, Kunming Botanical Garden, on living leaf of Chrysophullum sp., 19 March 2002, Wenping Wu KBG19-3 (NN047196 = MFLUCC 12-0264); CHINA, Hunan Province, Yizhang County, Mangshan, on living leaf of Schima sp., 12 April 2002, Wenping Wu HN40-8 (NN046983 = MFLUCC 12-0265); CHINA, Hunan Province, Yizhang County, Mangshan, on living leaf of Sympolocos sp., 12 April 2002, Wenping Wu HN38-6 (NN046978 = MFLUCC 12-0266); CHINA, Hunan Province, Yizhang County, Mangshan, on living leaf of unidentified tree, 12 April 2002, Wenping Wu HN14-4 (NN047099 = MFLUCC 12-0267).
Notes: Pestalotiopsis trachicarpicola is a recently described species from Kunming, China, and it causes serious leaf blotch and defoliation in Trachycarpus fortunei (Zhang et al. 2012a). The isolates we obtained as endophytes are morphologically and phylogenetically consistent to the type of P. trachicarpicola.
Pestalotiopsis umberspora Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800534
Figure 24 a–g.
Etymology: The specific epithet is based on the Latin = umber, in reference to the umber earth brown colour of the median cells of the conidia.
Conidiophores reduced to conidiogenous cells. Conidiogenous cells discrete or integrated, lageniform, hyaline, smooth walled, sometimes septate. Conidia 19–25 × 6–8 μm (\( \overline x = {21}.{3} \times {6}.{5}\;\mu {\text{m}} \)), fusiform, straight to slightly curved, 4-septate; basal cell obconic to conic, hyaline or pale brown, thin and verruculose, 3–4.5 μm long (\( \overline x = {3}.{8}\;\mu {\text{m}} \)); three median cells 12–14 μm long (\( \overline x = {13}.{1}\;\mu {\text{m}} \)), umber brown to olivaceous, septa and periclinal walls darker than the rest of the cell, versicoloured, verruculose, second cell from base pale brown, 3–4.5 μm (\( \overline x = {3}.{9}\;\mu {\text{m}} \)); third cell darker brown, 3.5–5 μm (\( \overline x = {4}.{3}\;\mu {\text{m}} \)); fourth cell darker, 3.5–4.5 μm (\( \overline x = {4}.{2}\;\mu {\text{m}} \)); apical cell 3–4.5 μm long (\( \overline x = {3}.{9}\;\mu {\text{m}} \)), hyaline, conic to obconic; with apical appendages 22–35 μm long (\( \overline x = {27}.{7}\;\mu {\text{m}} \)), tubular, 1–3 (mainly 3), arising from the upper portion of the apical cell; basal appendage, 5–7 μm (\( \overline x = {5}.{9}\;\mu {\text{m}} \)), filiform.
Colonies on PDA reaching 7 cm diam. after 6 days at 25 °C, edge entire, whitish, aerial mycelium on surface, fruiting bodies black, gregarious; reverse of culture pale yellow.
Habitat/Distribution: Saprobe on dead plant material, Guangxi Province, China.
Material examined: CHINA, Guangxi Province, Shiwandashan, on dead leaves of unidentified plant, 30 December 1997, Wenping Wu WU1554j (HMAS042986, holotype; MFLU12-0421, isotype; ex-type living culture NN042986 = MFLUCC 12-0285).
Notes: Pestalotiopsis umberspora is a phylogenetically distinct species in the genus and separates well in tef1 and combined multi-locus tree with its phylogenetically related species P. crysea. Its umber coloured and relatively wider mature conidia are characteristic to the species.
Pestalotiopsis unicolor Maharachchikumbura & K.D. Hyde, sp. nov.
MycoBank: MB 800523
Figure 25 a–g.
Etymology: Specific epithet in reference to concolorous median cells.
Conidiophores indistinct. Conidiogenous cells discrete ampulliform to lageniform, smooth, thin-walled, hyaline, with 1–2 proliferations. Conidia fusoid to ellipsoid, straight to slightly curved, 4-septate, 20–24.5 × 4–6 μm (\( \overline x = {22} \times {5}.{1}\;\mu {\text{m}} \)), basal cell conic to obconic, hyaline or slightly olivaceous, thin- and verruculose, 4–5.5 μm long (\( \overline x = {4}.{9}\;\mu {\text{m}} \)), with three median cells, doliiform to cylindrical, with thick verruculose walls, constricted at the septa, concolorous, olivaceous, septa and periclinal walls darker than the rest of the cell, wall rugose, together 13–16 μm long (\( \overline x = {14}.{7}\;\mu {\text{m}} \)) second cell from base 4–5 μm (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); third cell 4–5 μm (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); fourth cell 4–6 μm (\( \overline x = {5}\;\mu {\text{m}} \)); apical cell hyaline, conic to subcylindrical 3–5 μm long (\( \overline x = {4}.{2}\;\mu {\text{m}} \)); with 2–3 tubular apical appendages, arising from the apex of the apical cell, 11–20 μm long (\( \overline x = {17}.{5}\;\mu {\text{m}} \)), of unequal length; basal appendages present, rarely two, 4–10 μm (\( \overline x = {6}.{9}\;\mu {\text{m}} \)) long.
Habitat/Distribution: Endophyte on Rhododendron sp. and unidentified plant, Hunan Province, China.
Material examined: CHINA, Hunan Province, Yizhang County, Mangshan, on living leaf of Rhododendron sp., 12 April 2002, Wenping Wu HN42-1 (HMAS046974, holotype; MFLU12-0417, isotype; ex-type living culture NN046974 = MFLUCC 12-0276).
Additional culture examined: CHINA, Hunan Province, Yizhang County, Mangshan, on living leaf of unidentified tree, 12 April 2002, Wenping Wu HN51-1 (NN047308 = MFLUCC 12-0275).
Notes: Pestalotiopsis unicolor is a distinct species in the genus from molecular and morphological characters. The morphologically similar species in conidial size are P. kawakamii Sawada (20–24 × 5–7 μm) (Guba 1961) and P. algeriensis (Sacc. & Berl.) W.P. Wu. (17–23 × 5–7 μm) (Guba 1961). However, 2–3 tubular apical appendages (11–20 μm long) of P. unicolor are longer than in Pestalotia kawakamii (3 apical appendages; 5–10 μm long). The conidial width of P. algeriensis is similar to P. unicolor but the length of conidia and apical appendages are smaller in P. algeriensis and length (up to 16 μm) is shorter than in P. unicolor.
Pestalotiopsis verruculosa Maharachchikumbura, & K.D. Hyde, sp. nov.
MycoBank: MB 800527
Figure 26 a–h.
Etymology: The specific epithet is based on the Latin verruculose in reference to the verrucose pattern in walls of three median cells.
Conidia ellipsoid, straight to slightly curved, 4-septate, 28–35 × 9–11 μm (\( \overline x = {3}0.{6} \times {1}0.{3}\;\mu {\text{m}} \)), basal cell conic with obtuse end, hyaline, thin-walled and verruculose, 5–7 μm long (\( \overline x = {5}.{7}\;\mu {\text{m}} \)), with three median cells, doliiform to cylindrical, with thick verruculose walls, constricted at the septa, concolorous, olivaceous, septa and periclinal walls darker than the rest of the cell, wall rugose, together 18–26 μm long (\( \overline x = {21}.{6}\;\mu {\text{m}} \)) second cell from base 6–9 μm (\( \overline x = {6}.{8}\;\mu {\text{m}} \)); third cell 6–9 μm (\( \overline x = {7}.{5}\;\mu {\text{m}} \)); fourth cell 6–9 μm (\( \overline x = {7}.{3}\;\mu {\text{m}} \)); apical cell hyaline, conic to subcylindrical 4–6 μm long (\( \overline x = {4}.{8}\;\mu {\text{m}} \)); with 2–6 tubular apical appendages (mostly 3–4), arising from the apex of the apical cell (rarely 1 appendage arising from just above the septum separating upper median and apical cell), 25–40 μm long (\( \overline x = {34}\;\mu {\text{m}} \)); basal appendage present 8–12 μm (\( \overline x = {9}\;\mu {\text{m}} \)).
Colonies on PDA reaching 7 cm diam. after 15 days at 25 °C, edge undulate, whitish to pale yellow, with dense, aerial mycelium on surface, with black, gregarious fruiting bodies; reverse of culture yellow to pale orange.
Habitat/Distribution: Endophytes on living leaf of Rhododendron sp., Yunnan Province, China.
Material examined: CHINA, Yunnan Province, Kunming, Kunming Botanical Garden, on living leaf of Rhododendron sp., 19 March 2002, Wenping Wu KBG25-8 (HMAS047309, holotype; MFLU12-0416, isotype; ex-type culture NN047309 = MFLUCC 12-0274). Table 7
Notes: Pestalotiopsis verruculosa is a distinct species in terms of morphology, and its molecular phylogeny. It has a relatively large conidial size (28–35 × 9–11 μm) when compared with morphologically similar species (P. funerea, 21–29 × 7–9.5 μm); (P. multiseta, 22–26 × 7.5–9.5 μm). It also has a long apical appendage (25–40 μm) when compared to P. funerea (5–20 μm), P. multiseta (9–16 μm) and P. macrospora (15–22 μm) (Nag Rag 1993).
Discussion
In this study, we include 40 sequenced isolates of Pestalotiopsis to provide a backbone tree for the genus Pestalotiopsis. Based on morphological and molecular data, we determined that the 40 sequenced isolates comprise 23 species of Pestalotiopsis, of which 14 are new. Three species are epitypified, and six are recognized as existing species, namely; P. camelliae (Zhang et al. 2012b), P. furcata (Maharachchikumbura et al. 2012a), P. jesteri (Strobel et al. 2000), P. samarangensis (Maharachchikumbura et al. 2012b), P. theae (Maharachchikumbura et al. 2012a) and P. trachicarpicola (Zhang et al. 2012a). There is a need to deposit more sequences of epitypified species of Pestalotiopsis in GenBank and this communication provides data on three genes for 14 new species, three epitypified species and six species with type/epitype sequences from GenBank.
In our studies, we analyzed sequence data from three gene regions, individually and in combination, to evaluate their ability to resolve species limits of Pestalotiopsis. In all cases, the genes resolved the species into three strongly supported Clades (A, B and C). These clades corresponded to three conidial types: Clade A with pale brown or olivaceous concolorous median conidial cells, Clade B with versicolorous median conidial cells and Clade C with dark-coloured concolorous median conidial cells and with knobbed apical appendages. These Clades were also evident in the studies of Jeewon et al. (2003), Liu et al. (2010) and Maharachchikumbura et al. (2011). Steyaert (1949) and Guba (1961) had previously grouped species with versicolorous conidia into two groups based on the intensity of colour of the median cells (umber olivaceous and fuliginous olivaceous). Our findings, based on combined genes analysis, did not support this arrangement. In the combined phylogenetic tree (Fig. 4) P. clavispora, P. ellipsospora, P. foedans, P. samarangensis and P. saprophyta, Clade B diverges from a common ancestor. Within this Sub-clade P. ellipsospora and P. foedans have pale brown, versicolorous median conidial cells, while all other species belonging in this sub-clade have dark brown versicolorous median conidial cells. Thus, we agree with Jeewon et al. (2003), Liu et al. (2010) and Maharachchikumbura et al. (2011) in concluding that the division of the “versicolor group” based on colour intensities of the median conidial cell is not a taxonomically good character. In addition to differences in median, conidial cells (Clades A, B and C); the size variation in conidia, and presence or absence of basal appendages, can be used as additional taxonomic characters for defining Pestalotiopsis species. Apical appendage characters, such as branching pattern, number, position of attachment to apical cell, and knobbed or knotted nature are also useful as a species, but not as a generic character (Crous et al. 2012). Differentiation of species based on these characters is also supported by molecular data and has been previously noted by Jeewon et al. (2003), Hu et al. (2007), Liu et al. (2010) and Maharachchikumbura et al. (2011).
In our study, we tested 10 gene regions for suitability in resolving species in Pestalotiopsis, but narrowed this down to three most applicable regions, which were tested individually and in combination, to evaluate the differences between species. The ITS is the universal barcode for fungi (Schoch et al. 2012). It has a high sequence and PCR success rate with higher resolving power within some fungal lineages (Bridge et al. 2005; Schoch et al. 2012). The species of Pestalotiopsis sequenced with ITS in this study had a high PCR and sequence success rate. However, ITS could not fulfill the role as candidate gene for species discrimination, as the data did not have high variation between species. Thus, possible cryptic taxa could not be discriminated from one another.
A gene-by-gene assessment of phylogenetic resolution yielded higher levels with protein genes as compared to ribosomal regions (Schoch et al. 2009). Cryptic taxa can be better resolved using slow evolving protein coding genes (Liu et al. 1999; Liu and Hall 2004). Hu et al. (2007) and Liu et al. (2010) used a β-tubulin fragment to study species relationships within Pestalotiopsis. This region has also been shown to resolve species in other genera in groups such as Discosia (Tanaka et al. 2011), Seimatosporium (Tanaka et al. 2011) and Seiridium (Barnes et al. 2001). Tef1 is a widely used taxonomic marker and this has been successfully utilized to investigate the kingdom-level phylogeny of eukaryotes (Roger et al. 1999; Baldauf et al. 2000) and species in fungal genera such as Diaporthe (Santos et al. 2010; Udayanga et al. 2012), Fusarium (Geiser et al. 2004; O’Donnell et al. 2010) and Trichoderma and Hypocrea (Druzhinina et al. 2005). In the present study, β-tubulin and tef1 gene regions proved to be favorable taxonomic markers for Pestalotiopsis since they resolved the taxonomic relationships of most species studied. Further, tef1 had better PCR amplification success rates (95 %) and was found to be superior to β-tubulin (90 %). Tef1 is therefore a powerful tool to resolve lineages within Pestalotiopsis. Because of the better PCR and sequencing success rate and fewer difficulties with alignment, editing and better resolution, the tef1 gene appears to be a very good molecular marker for phylogenetic investigation of Pestalotiopsis.
Combined sequence analysis of ITS, β-tubulin and tef1 genes successfully resolved most of the Pestalotiopsis species used in this study with high bootstrap support. Hu et al. (2007) and Liu et al. (2010) have previously shown that a combination of β-tubulin and ITS genes gave better species resolution than a single gene and they suggested that at least two genes should be used to resolve species in Pestalotiopsis. Similar results have been shown in Fusarium (Summerell et al. 2010); Calonectria (Lombard et al. 2010), Phyllosticta (Wikee et al. 2011), and Colletotrichum (Phoulivong et al. 2010), however the genes best suited for each genus differed. In addition to the above three genes, we tested LSU, SSU, ACT and GPDH (low resolution), GS and RPB1 (cannot be synthesized using available primers or multiple copies) and Calmodulin (species resolution is high, PCR success rate low) and these were less successful in PCR amplification and/or resolving species.
Three epitypes based on live cultures derived from fresh collections from China and Fiji were chosen. When choosing epitypes it is desirable (but not mandatory) to use fresh collections with living isolates from the original host, with the same symptoms and as near to the original location of the holotype as possible (Zhang and Hyde 2008). However, this is not straightforward for Pestalotiopsis species which may be endophytes, weak pathogens or saprobes on a wide range of hosts. There are also numerous cryptic species, very few distinct species, species with wide host ranges, those with cosmopolitan distribution and some species being opportunistic pathogens. We have, therefore, been pragmatic in choosing epitypes so that we can advance Pestalotiopsis understanding. In this way future workers can pin names to isolated endophytic Pestalotiopsis species (Ko Ko et al. 2011) which may be important for chemical bioprospecting and other research in the genus (Xu et al. 2010; Maharachchikumbura et al. 2011).
In the present study multi-locus phylogeny for Pestalotiopsis species is presented and strives towards providing biological species concepts based on taxonomically important morphological characters. This backbone tree needs expanding by re-examining type materials of some of the important species described in Pestalotiopsis and using multi-locus analysis to establish epitypes. We have not epitypified some of the more commonly known Pestalotiopsis names (e.g. P. disseminata, P. fici, P. longiseta, P. microspora, P. neglecta, P. pauciseta, P. photiniae and P. uvicola) due of lack of fresh collections and living material. Our new species, however, differ from putatively named examples of the common species in GenBank (e.g. P. microspora, P. neglecta) and thus we are confident that the species newly introduced in this paper are distinct.
References
Aly AH, Debbab A, Kjer J, Proksch P (2010) Fungal endophytes from higher plants: a prolific source of phytochemicals and other bioactive natural products. Fungal Divers 41:1–16
Baldauf SL, Roger AJ, Wenk-Siefert I, Doolittle WF (2000) A kingdom-level phylogeny of eukaryotes based on combined protein data. Science 290:972–977
Barnes I, Roux J, Coetzee MPA, Wingfield MJ (2001) Characterization of Seiridium spp. associated with cypress canker based on ȕ-tubulin and histone sequences. Plant Dis 85:317–321
Barr ME (1975) Pestalosphaeria, a new genus in the Amphisphaeriaceae. Mycologia 67:187–194
Barr ME (1990) Prodromus to nonlichenized, pyrenomycetous members of class Hymenoascomycetes. Mycotaxon 39:43–184
Bate-Smith EC, Metcalfe CR (1957) Leucanthocyanins.3. The nature and systematic distribution of tannin in dicotyledonous plants. J Linn Soc (Bot) 55:669–705
Bridge PD, Spooner BM, Roberts PJ (2005) The impact of molecular data in fungal systematics. Adv Bot Res 42:33–67
Cai L, Giraud T, Zhang N, Begerow D, Cai G, Shivas RG (2011) The evolution of species concepts and species recognition criteria in plant pathogenic fungi. Fungal Divers 50:121–133
Carbone I, Kohn LM (1999) A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 91:553–556
Chomnunti P, Schoch CL, Aguirre-Hudson B, Ko Ko TW, Hongsanan S, Jones EBG, Kodsueb R, Phookamsak R, Chukeatirote E, Bahkali AH, Hyde KD (2011) Capnodiaceae. Fungal Divers 51:103–134
Crous PW, Verkley GJM, Christensen M, Castañeda-Ruiz RF, Groenewald JZ (2012) How important are conidial appendages? Persoonia 28:126–137
Das R, Chutia M, Das K, Jha DK (2010) Factors affecting sporulation of Pestalotiopsis disseminata causing grey blight disease of Persea bombycina Kost., the primary food plant of muga silkworm. Crop Prot 29:963–968
Dennis RWG (1995) Fungi of the South East England. Royal Botanic Gardens, Kew
Ding G, Liu S, Guo L, Zhou Y, Che Y (2008) Antifungal metabolites from the plant endophytic fungus Pestalotiopsis foedan. J Nat Prod 71:615–618
Douanla-Meli C, Langer E (2009) Pestalotiopsis theae (Ascomycota, Amphisphaeriaceae) on seeds of Diospyros crassiflora (Ebenaceae). Mycotaxon 107:441–448
Druzhinina IS, Kopchinskiy AG, Komoń M, Bissett J, Szakacs G, Kubicek CP (2005) An oligonucleotide barcode for species identification in Trichoderma and Hypocrea. Fungal Genet Biol 42:813–828
Ellis MB, Ellis JP (1997) Microfungi on land plants: an identification handbook, 2nd edn (New Enlarged). Richmond Publishing
Espinoza JG, Briceno EX, Keith LM, Latorre BA (2008) Canker and Twig Dieback of blueberry caused by Pestalotiopsis spp. and a Truncatella sp. in Chile. Plant Dis 92:1407–1414
Felsenstein J (1985) Confidence limits on phylogenies: An approach using the bootstrap. Evolution 39:783–791
Geiser DM, Jiménez-Gasco M, Kang S, Makalowska I, Veeraraghavan N, Ward TJ, Zhang N, Kuldau GA, O’Donnell K (2004) FUSARIUM-ID v. 1.0: A DNA sequence database for identifying Fusarium. Eur J Plant Pathol 110:473–479
Glass NL, Donaldson GC (1995) Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 61:1323–1330
Guba EF (1961) Monograph of Pestalotia and Monochaetia. Harvard University Press, Cambridge
Guerber JC, Liu B, Correll JC, Johnston PR (2003) Characterization of diversity in Colletotrichum acutatum sensu lato by sequence analysis of two gene introns, mtDNA and intron RFLPs, and mating compatibility. Mycologia 95:872–895
Guo LD, Hyde KD, Liew ECY (2000) Identification of endophytic fungi from Livistona chinensis (Palmae) using morphological and molecular techniques. New Phytol 147:617–630
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hu HL, Jeewon R, Zhou DQ, Zhou TX, Hyde KD (2007) Phylogenetic diversity of endophytic Pestalotiopsis species in Pinus armandii and Ribes spp.: evidence from rDNA and β- tubulin gene phylogenies. Fungal Divers 24:1–22
Hyde KD, Bussaban B, Paulus B, Crous PW, Lee S, McKenzie EHC, Photita W, Lumyong S (2007) Diversity of saprobic microfungi. Biodivers Conserv 16:7–35
Jeewon R, Liew ECY, Simpson JA, Hodgkiss IJ, Hyde KD (2003) Phylogenetic significance of morphological characters in the taxonomy of Pestalotiopsis species. Mol Phylogenet Evol 27:372–383
Kang JC, Hyde KD, Kong RYC (1999) Studies on the Amphisphaeriales. The Amphisphaeriaceae (sensu stricto). Mycol Res 103:53–64
Kang JC, Kong RYC, Hyde KD (1998) Studies on the Amphisphaeriales I. Amphisphaeriaceae (sensu stricto) and its phylogenetic relationships inferred from 5.8 S rDNA and ITS2 sequences. Fungal Divers 1:147–157
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9:276–285
Keith LM, Velasquez ME, Zee FT (2006) Identification and characterization of Pestalotiopsis spp. causing scab disease of guava, Psidium guajava in Hawaii. Plant Dis 90:16–23
Kishino H, Hasegawa M (1989) Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data. J Mol Evol 29:170–179
Kohlmeyer J, Kohlmeyer VB (2001) Fungi on Juncus roemerianus 16. More new coelomycetes, including Tetranacriella gen. nov. Bot Mar 44:147–156
Ko Ko T, Stephenson SL, Bahkali AH, Hyde KD (2011) From morphology to molecular biology: can we use sequence data to identify fungal endophytes? Fungal Divers 50:113–120
Lee S, Crous PW, Wingfield MJ (2006) Pestalotioid fungi from Restionaceae in the Cape Floral Kingdom. Stud Mycol 55:175–187
Li E, Jiang L, Guo L, Zhang H, Che Y (2008) Pestalachlorides A–C, antifungal metabolites from the plant endophytic fungus Pestalotiopsis adusta. Bioorg Med Chem 16:7894–7899
Liu AR, Chen SC, Wu SY, Xu T, Guo LD, Jeewon R, Wei JG (2010) Cultural studies coupled with DNA based sequence analyses and its implication on pigmentation as a phylogenetic marker in Pestalotiopsis taxonomy. Mol Phylogenet Evol 57:528–535
Liu YJ, Hall BD (2004) Body plan evolution of ascomycetes, as inferred from an RNA polymerase II phylogeny. Proc Natl Acad Sci 101:4507–4512
Liu YJ, Whelen S, Hall BD (1999) Phylogenetic relationships among ascomycetes: evidence from an RNA polymerase II subunit. Mol Biol Evol 16:1799–808
Liu AR, Xu T, Guo LD (2007) Molecular and morphological description of Pestalotiopsis hainanensis sp. nov., a new endophyte from a tropical region of China. Fungal Divers 24:23–36
Lombard L, Crous PW, Wingfield BD, Wingfield MJ (2010) Phylogeny and systematics of the genus Calonectria. Stud Mycol 66:31–69
Maharachchikumbura SSN, Guo LD, Chukeatirote E, Bahkali AH, Hyde KD (2011) Pestalotiopsis–morphology, phylogeny, biochemistry and diversity. Fungal Divers 50:167–187
Maharachchikumbura SSN, Chukeatirote E, Guo LD, Crous PW, McKenzie EHC, Hyde KD (2012a) Pestalotiopsis species associated with Camellia sinensis (tea). Mycotaxon (in press)
Maharachchikumbura SSN, Guo LD, Chukeatirote E, McKenzie EHC, Hyde KD (2012b) A destructive new disease of Syzygium samarangense in Thailand caused by the new species Pestalotiopsis samarangensis. Trop plant pathol (in press)
Maas JL (1971) Hyalotia pistacina a new species and a notes on Pestalotiopsis thujae. Mycologia 63:663–668
Moriya S, Inoue S, Ohkuma M, Yaovapa T, Kohjima T, Suwanarit P, Sangwanti U, Vongkaluang C, Noparatnaraporn N, Kudo T (2005) Fungal community analysis of fungal gardens in termite nests. Microbes Environ 20:243–252
Myllys L, Stenroos S, Thell A (2002) New genes for phylogenetic studies of lichenized fungi: glyceraldehyde-3-phosphate dehydrogenase and beta-tubulin genes. Lichenologist 34:237–246
Nag Rag TR (1993) Coelomycetous anamorphs with appendage bearing conidia. Mycologue, Waterloo
O’Donnell K, Cigelnik E (1997) Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol Phylogenet Evol 7:103–116
O’Donnell K, Nirenberg H, Aoki T, Cigelnik E (2000) A multigene phylogeny of the Gibberella fujikuroi species complex: detection of additional phylogenetically distinct species. Mycoscience 41:61–78
O’Donnell K et al (2010) Internet-accessible DNA sequence database for identifying fusaria from human and animal infections. J Clin Microbiol 48:3708–3718
Page RDM (1996) TREEVIEW: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Phoulivong S, Cai L, Chen H, McKenzie EHC, Abdelsalam K, Chukeatirote E, Hyde KD (2010) Colletotrichum gloeosporioides is not a common pathogen on tropical fruits. Fungal Divers 44:33–43
Rehner SA, Samuels GJ (1994) Taxonomy and phylogeny of Gliocladium analysed from nuclear large subunit ribosomal DNA sequences. Mycol Res 98:625–634
Rehner SA (2001) Primers for Elongation Factor 1-alpha (EF1-alpha). http://ocid.nacse.org/research/deephyphae/EF1primer.pdf
Roger AJ, Sandblom O, Doolittle WF, Philippe H (1999) An evaluation of elongation factor 1 alpha as a phylogenetic marker for eukaryotes. Mol Biol Evol 16:218–233
Saccardo PA (1902) Deuteromycetes, melanconiaceae, phragmasporae, monochaetia. Sylloge Fungorum 16:1017
Saccardo PA (1914) Fungi italici. Ann Mycol 12:310
Saccardo PA (1931) Deuteromycetes, melanconiaceae, monochaetia. Sylloge Fungorum 25:610
Santos JM, Correia VG, Phillips AJL (2010) Primers for mating-type diagnosis in Diaporthe and Phomopsis: their use in teleomorph induction in vitro and biological species definition. Fungal Biology 114:255–270
Schoch CL et al (2009) The ascomycota tree of life: a phylum-wide phylogeny clarifies the origin and evolution of fundamental reproductive and ecological traits. Syst Biol 58:224–239
Schoch CL et al (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for fungi. Proc Natl Acad Sci USA. doi:10.1073/pnas.1117018109, Available at: www.pnas.org/cgi/doi/10.1073/pnas.1117018109
Stephenson SA, Green JR, Manners JM, Maclean DJ (1997) Cloning and characterisation of glutamine synthetase from Colletotrichum gloeosporioides and demonstration of elevated expression during pathogenesis on Stylosanthes guianensis. Curr Genet 31:447–454
Steyaert RL (1949) Contributions à l’étude monographique de Pestalotia de Not. et Monochaetia Sacc. (Truncatella gen. nov. et Pestalotiopsis gen. nov.). Bull Jard Bot Brux 19:285–354
Strobel G, Li JY, Ford E, Worapong J, Gary IB, Hess WM (2000) Pestalotiopsis jesteri, sp. nov. an endophyte from Fragraea bodenii Wernh, a common plant in the southern highlands of Papua New Guinea. Mycotaxon 76:257–266
Strobel G, Ford E, Worapong J, Harper JK, Arif AM, Grant DM, Fung PC, Chau MW (2002) Isopestacin, an isobenzofuranone from Pestalotiopsis microspora, possessing antifungal and antioxidant activities. Phytochemistry 60:179–183
Strobel G, Yang XS, Sears J, Kramer R, Sidhu RS, Hess WM (1996) Taxol from Pestalotiopsis microspora of Taxus wallachiana. Microbiology 142:435–440
Summerell BA, Laurence MH, Liew ECY, Leslie JF (2010) Biogeography and phylogeography of Fusarium: a review. Fungal Divers 44:3–13
Swofford DL (2002) PAUP* 4.0: phylogenetic analysis using parsimony (* and other methods). Sinauer Associates, Sunderland
Tanaka K, Endo M, Hirayama K, Okane I, Hosoya T, Sato T (2011) Phylogeny of Discosia and Seimatosporium, and introduction of Adisciso and Immersidiscosia genera nova. Persoonia 26:85–98
Tejesvi MV, Kini KR, Prakash HS, Subbiah V, Shetty HS (2007a) Genetic diversity and antifungal activity of species of Pestalotiopsis isolated as endophytes from medicinal plants. Fungal Divers 24:37–54
Tejesvi MV, Nalini MS, Mahesh B, Prakash HS, Kini KR, Shetty HS, Subbiah V (2007b) New hopes from endophytic fungal secondary metabolites. Bol Soc Quím de Méx 1(1):19–26
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Thongkantha S, Lumyong S, McKenzie EHC, Hyde KD (2008) Fungal saprobes and pathogens occurring on tissues of Dracaena lourieri and Pandanus spp. in Thailand. Fungal Divers 30:149–169
Udayanga D, Liu XZ, Crous PW, McKenzie EHC, Chukeatirote E, Hyde KD (2012) A multi-locus phylogenetic evaluation of Diaporthe (Phomopsis). Fungal Divers (in press)
Venkatasubbaiah P, Grand LF, Dyke CGV (1991) A new species of Pestalotiopsis on Oenothera. Mycologia 83:511–513
Wei JG, Xu T (2004) Pestalotiopsis kunmingensis, sp. nov., an endophyte from Podocarpus macrophyllus. Fungal Divers 15:247–254
Wei JG, Xu T, Guo LD, Liu AR, Zhang Y, Pan XH (2007) Endophytic Pestalotiopsis species associated with plants of Podocarpaceae, Theaceae and Taxaceae in southern China. Fungal Divers 24:55–74
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic, New York, pp 315–322
Wikee S, Cai L, Pairin N, McKenzie EHC, Su YY, Chukeatirote E, Thi HN, Bahkali AH, Moslem MA, Abdelsalam K (2011) Colletotrichum species from jasmine (Jasminum sambac). Fungal Divers 46:171–182
Wu CG, Tseng HY, Chen ZC (1982) Fungi inhabiting on Schoenoplectus triqueter (L.) Palla (I). Taiwania 27:35–38
Xu J, Ebada SS, Proksch P (2010) Pestalotiopsis a highly creative genus: chemistry and bioactivity of secondary metabolites. Fungal Divers 44:15–31
Yasuda F, Kobayashi T, Watanabe H, Izawa H (2003) Addition of Pestalotiopsis spp. to leaf spot pathogens of Japanese persimmon. J Gen Plant Pathol 69:29–32
Zhao GC, Li N (1995) Thirty-four species of pestalotiopsis in Yunnan. J Northeast For Univ 23:21–33
Zhang J, Xu T, Ge Q (2002) Notes on Pestalotiopsis from southern China. Mycotaxon 85:91–99
Zhang Y, Hyde KD (2008) Epitypification: should we epitypify? J Zhejiang Univ Sci B 9:842–846
Zhang YM, Maharachchikumbura SSN, McKenzie EHC, Hyde KD (2012a) A novel species of Pestalotiopsis causing leaf spots of Trachycarpus fortunei. Cryptogamie mycol (in press)
Zhang YM, Maharachchikumbura SSN, McKenzie EHC, Hyde KD (2012b) Pestalotiopsis camelliae sp. nov. associated with grey blight of Camellia japonica in China (unpublished)
Zhou D, Hyde KD (2001) Host-specificity, host-exclusivity and host-recurrence in saprobic fungi. Mycol Res 105:1449–1457
Acknowledgments
We thank Thailand Research Fund (grant: BRG5280002); The National Research Council of Thailand (grant for Pestalotiopsis No: 55201020008); Mae Fah Luang University (grant for Pestalotiopsis No: 55101020004); The National Natural Science Foundation of China (grant: 30930005); the Knowledge Innovation Program of the Chinese Academy of Sciences (grant: KSCX2-YW-Z-0935); Mushroom Research Foundation, Chiang Mai, Thailand; State Key Laboratory of Mycology, Institute of Microbiology, Chinese Academy of Sciences, Beijing, China and the King Saud University for supporting this research. We kindly acknowledge Novozymes, CGMCC and ICMP for providing cultures and BPI, CUP and NY herbaria for providing type materials for this study.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Maharachchikumbura, S.S.N., Guo, LD., Cai, L. et al. A multi-locus backbone tree for Pestalotiopsis, with a polyphasic characterization of 14 new species. Fungal Diversity 56, 95–129 (2012). https://doi.org/10.1007/s13225-012-0198-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-012-0198-1