Abstract
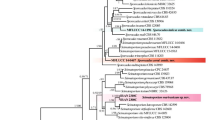
Septoria leaf spot is a severe disease of lettuce in tropical and subtropical regions, inducing yield/quality losses and increasing production costs. Globally, Septoria lactucae has been reported as the major pathogen associated with this disease. However, a complex of Septoria species has been also reported in association with lettuce crop around the world. Extensive surveys of Septoria species associated with this disease have not been conducted in the Neotropics. Therefore, the objective of the present study was to undergo a phenotypic and genetic characterization of a collection of 32 Brazilian Septoria isolates causing lettuce leaf spot. Morphometrical characterization of conidia and pycnidia was conducted with ten isolates obtained from three lettuce-producing regions in Brazil. In addition, partial sequences of four gene regions (β-tubulin, RPB2, actin, and calmodulin) were used in phylogenetic analyses of all 32 Septoria isolates. All tested isolates were pathogenic to two lettuce cultivars and morphometrically similar to each other. These isolates grouped together into a single monophyletic clade composed exclusively by genuine S. lactucae isolates. Therefore, S. lactucae was found to be the sole species associated with the Septoria leaf spot of lettuce in Brazil. The number of polymorphic sites (n = 27), mutations (n = 190), the level of nucleotide diversity (0.00262), and the average number of nucleotide differences (2.961) were relatively low. Notwithstanding, this genetic variability allowed the identification of 17 haplotypes amongst the 32 Brazilian isolates. This information will help to guide resistance breeding programs and establish more effective management strategies of this disease.




Similar content being viewed by others
References
Bedlan G (1999) Septoria birgitae Bedlan sp. n., a new pathogen causing leaf spots on Lactuca sativa. Mycotaxon 70:51–54
Bertus AL (1972) The eradication of seed-borne Septoria lactucae Pass, from lettuce with aerated steam. J Horticult Sci 47:259–261
Blancard D, Lot H, Maisonneuve BA (2006) Color atlas of diseases of lettuce and related salad crops – observation, biology and control. Academic Press, USA, p 375
Boiteux LS, Fonseca MEN, Simon PW (1999) Effects of plant tissue and DNA purification method on randomly amplified polymorphic DNA-based genetic fingerprinting analysis in carrot. J Am Soc Hortic Sci 124:32–38
Carbone I, Kohn LM (1999) A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 91:553–556
Castellani A (1939) Viability of some pathogenic fungi in distilled water. J Trop Med Hyg 24:270–276
Clement M, Snell Q, Walker P, Posada D, Crandall K (2002) TCS: estimating gene genealogies. Parallel and distributed processing symposium, International Proceedings 2, p 184
Crous PW, Quaedvlieg W, Sarpkaya K, Can C, Erkılıç A (2013) Septoria-like pathogens causing leaf and fruit spot of pistachio. IMA Fungus 4:187–199. https://doi.org/10.5598/imafungus.2013.04.02.04
Dingra OD, Sinclair JB (1995) Basic plant pathology methods, 2nd edn. Lewis Publishers, London
FAO (2019) FAOSTAT database. Rome (2017). http://www.fao.org/faostat/en/#data. Accessed 26 March 2019
Farr DF, Rossman AY (2019) Fungal Databases, U.S. National Fungus Collections, ARS, USDA. https://nt.ars-grin.gov/fungaldatabases/ Accessed 26 March 2019
Hasegawa M, Kishino H, Yano T (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755. https://doi.org/10.1093/bioinformatics/17.8.754
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Leigh JW, Bryant D (2015) POPART: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Liu Y, Whelen S, Hall B (1999) Phylogenetic relationships among ascomycetes: evidence from an RNA polymerse II subunit. Mol Biol Evol 16:1799–1808
Lohmeier U, Farahani-Kofoet RD, Kofoet A, Grosch R (2013) Factors affecting incidence and severity of leaf spot disease on lettuce caused by Septoria birgitae. Ann Appl Biol 162:221–230. https://doi.org/10.1111/aab.12016
Lourenço Jr. V, Moya A, González-Candelas F, Carbone I, Maffia LA, Mizubuti ESG (2009) Molecular diversity and evolutionary processes of Alternaria solani in Brazil inferred using genealogical and coalescent approaches. Phytopathology 99:765–774
Nao M (2008) Effects of inoculum density, leaf wetness duration and nitrate concentration on the occurrence of lettuce leaf spot. J Gen Plant Pathol 74:208–212. https://doi.org/10.1007/s10327-008-0086-4
Peixoto CC, Cabral CS, Fonseca MEN, Boiteux LS, Reis A (2021) Species diversity, novel interactions and absence of well-supported host-guided phylogenetic groupings of Neotropical Alternaria isolates causing foliar lesions in Solanaceae. J Appl Microbiol. https://doi.org/10.1111/jam.15115
Punithalingam E, Holliday P (1972) Septoria lactucae. CMI Descriptions of Pathogenic Fungi and Bacteria no. 335
Quaedvlieg W, Kema GHJ, Groenewald JZ, Verkley GJM, Seifbarghi S, Razavi M, Gohari AM, Mehrabi R, Crous PW (2011) Zymoseptoria gen. nov.: a new genus to accommodate Septoria-like species occurring on graminicolous hosts. Persoonia 26:57–69. https://doi.org/10.3767/003158511X571841
Quaedvlieg W, Groenewald JZ, de Jesús Yáñez-Morales M, Crous PW (2012) DNA barcoding of Mycosphaerella species of quarantine importance to Europe. Persoonia 29:101–115
Quaedvlieg W, Verkley GJM, Shin H-D, Barreto RW, Alfenas AC, Swart WJ, Groenewald JZ, Crous PW (2013) Sizing up Septoria. Stud Mycol 75:307–390. https://doi.org/10.3114/sim0017
Sousa CS, Kerr WE, Santos MR, Arruda AS, Spini VBMG, Juliatti FC, Takatsu A (2003) Mancha de Septoria da alface: isolamento, inoculação e avaliação de cultivares em condições de campo e casa de vegetação. Fitopatol Bras 28:555–558. https://doi.org/10.1590/S0100-41582003000500016
Stukenbrock EH, Quaedvlieg W, Javan-Nikhah M, Zala M, Crous PW, McDonald BA (2012) Zymoseptoria ardabilia and Z. pseudotritici, two progenitor species of the septoria tritici leaf blotch fungus Z. tritici (synonym: Mycosphaerella graminicola). Mycologia 104: 1397–1407
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Verkley GJM, Quaedvlieg W, Shin H-D, Crous PW (2013) A new approach to species delimitation in Septoria. Stud Mycol 75:213–305. https://doi.org/10.3114/sim0018
Zafari D, Razaghi P (2013) Septoria sonchi causes Sonchus oleraceus leaf spot in Iran. Aust Plant Dis Notes 8:55–57. https://doi.org/10.1007/s13314-013-0094-x
Acknowledgements
Maria Esther de Noronha Fonseca, Ailton Reis and Leonardo S. Boiteux were supported by fellowships from the Brazilian National Research Council (CNPq). Cléia Santos Cabral was supported by fellowships from CAPES and CNPq.
Funding
Not applicable.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors do not have any conflict of interest.
Data availability
Alignments from this study were submitted to TreeBase and the sequences to GenBank.
Informed consent
All authors have reviewed the manuscript and approved its submission to Journal of Plant Disease and Protection.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cabral, C.S., Fonseca, M.E.N., Boiteux, L.S. et al. Phenotypic and genetic variability of fungal isolates associated with the Septoria leaf spot disease of lettuce (Lactuca sativa) in Brazil. J Plant Dis Prot 129, 53–62 (2022). https://doi.org/10.1007/s41348-021-00537-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41348-021-00537-9