Abstract
Whilst Brazil is the fourth largest cotton producer globally, incidence of ramularia leaf spot (RLS) has decreased yield. In 2017–18 and 2018–19, ca. 300 fungal samples were collected throughout Brazil. Hyphal tip cultures were obtained for amplification of the RNA polymerase II (RPB2), 28S rRNA, the ribosomal DNA internal transcribed spacers (ITS), actin (ACT), elongation factor (EF1-α) and histone H3 (HIS3) genomic regions. Additionally, sequences of the glyceraldehyde-3-phosphate dehydrogenase (GAPDH) were obtained by nanopore sequencing and the EF1-α region was selected as a marker for rapid recognition of Ramulariopsis species. Clade assignments based on the concatenated-sequence tree were identical to those in tree generated by RPB2-sequences, as well as in an RPB2 haplotype network and an ISSR (TGTC)4 dendrogram, in identification with species-specific primers and based on morphological comparisons. Out of 267 examined isolates, 252 were identified as Ramulariopsis pseudoglycines, indicating this species as the most widespread causal agent of cotton RLS in the Brazilian growing regions. Species-specific primers developed in the study that target the EF1-α gene provide an opportunity for extensive RLS sampling worldwide to study the distribution of Ramulariopsis species. Such data will aid breeders and plant pathologists in cotton disease resistance development and fungicide resistance avoidance.
Similar content being viewed by others
Introduction
Cotton (Gossypium spp.) is the world´s most cultivated fibre crop, mostly for the supply of raw materials for the textile industry, as well as for oil and protein extraction1. Since the 1990s, Brazil has ranked globally in fourth place in terms of cotton production, following strong investments in the production technology and expansion of cultivated areas, mainly in the Brazilian Cerrado biome2,3,4,5 (a savannah-like region in the Brazilian Midwest).
Favorable environmental conditions during the growing season, combined with the cultivation of a limited number of cotton genotypes over very large areas, favour epidemics of ramularia leaf spot4,6. This disease was first reported in Paraguari, Paraguay in 18837, and soon after was followed by a report from Alabama, USA in 18908. Since then, the disease has been reported in more than 40 cotton-producing countries9,10. Typically, ramularia leaf spot (RLS) is observed at the end of the cotton cycle and has therefore generally been considered a disease of secondary importance.
The disease was first reported in Brazil by the Agricultural Inspection and Defense Service of São Paulo in 1919 and was considered of secondary importance until the 1990s9,10,11. Currently, RLS is the major cotton disease in Brazil, with up to eight fungicide spray applications required to reduce the negative effects on yield and cotton fibre quality4,5,12.
Historically, Ramularia areola was the specific epithet used to designate the causal agent of RLS13. In 1961, this species was recombined to Ramularia gossypii (Speg.) Cif. and, due to the morphological similarities with the related genus Ramulariopsis (conidiophores severely branched at the base with terminal and lateral conidiogenic cells), the taxon was recombined again in 1993 to Ramulariopsis gossypii (Speg.) Braun14. Later, a multigenic study including a representative number of isolates of Ramularia and allied genera revealed a new species, Ramulariopsis pseudoglycines, associated with RLS15.
Up to now, the etiology of the disease in Brazil has remained inaccurate, given that most identifications are based on only few isolates or limited to the examination of morphological data15,16,17. Furthermore, it has been shown that a molecular perspective, associated with morphological data, is required to resolve plant pathogen species complexes, with this combined approach effective in revealing previously uncharacterized species affecting different crops15,18,19. For example, molecular characterization of isolates previously identified as Ramularia eucalypti based exclusively on morphological comparisons revealed a total of seven species associated with the eucalyptus leaf spot20.
Although the sequencing of target regions is efficient for the accurate identification of Ramulariopsis species, recommended GAPDH sequences as secondary barcodes for identification of Ramularia and allied genera are currently absent for R. pseudoglycines in public databases15,20. Furthermore, sequencing-based approaches for molecular identification are time-consuming, costly and dependent upon trained personnel with expertise in phylogenetic analysis compared to simple visualization of amplicons obtained with species-specific primers21. As such, more accessible specific and rapid molecular tools are needed for routine identification of R. gossypii and R. pseudoglycines.
The diverse reactions of resistant and susceptible genotypes of cotton to different isolates of Ramulariopsis have been ascribed to the genetic variability of the pathogen22. An analysis of the genetic diversity of 16 isolates of Ramulariopsis previously revealed three genetic groups23. Such information indicates a greater variability of the causal agent of the RLS than has so far been acknowledged. Nevertheless, there are still relatively few studies addressing the genetic diversity of Ramulariopsis associated with cotton in Brazil4,22,23,24,25.
Accurate identification of the RLS pathogen in Brazil, as well as determination of the relative abundancy of each causal agent, and respective geographic distribution, are particularly important for disease management approaches, and are paramount for cotton breeding programs. The precise identification of the Ramulariopsis species in each growing area, combined with appropriate phytosanitary measures, can minimize damage caused by RLS.
Here, Ramulariopsis isolates collected in the main cotton-growing regions in Brazil were morphologically and molecularly characterized. In addition, a PCR-based method was developed to distinguish between isolates of R. gossypii and R. pseudoglycines, which will aid breeders and plant pathologists in cotton disease resistance development and fungicide resistance avoidance.
Results
Sampling and isolates
Symptomatic leaf samples were collected from 24 growing fields representing seven Brazilian states (Fig. 1). Naturally occurring symptoms included light green to yellow-green lesions delimited by the veinlets, giving them an angular or irregular shape, with white powdery sporulation on both sides of the leaves. Under favourable disease conditions, lesions coalesced, become chlorotic and then necrotic, often resulting in severe defoliation (Fig. 2). A total of 267 Ramulariopsis isolates (Supplementary Table S1) were obtained in the Brazilian states of Bahia (n = 44), Distrito Federal (n = 32), Goiás (n = 31), Maranhão (n = 30), Mato Grosso (n = 87), Mato Grosso do Sul (n = 39) and Paraíba (n = 4). The voucher specimens of R. gossypii and R. pseudoglycines were deposited at the Herbarium of the University of Brasilia (UB).
(a–h) Ramularia leaf spot symptoms in cotton. (a) Healthy leaves observed in the field. (b, c, d) Initial symptoms; Early sporulation of Ramulariopsis on adaxial sides of cotton leaf; (e, f) Late sporulation on necrotic lesions on the upper surface of cotton leaf; (g) Necrotic lesions covering the leaf. (h) Collapsed leaf with advanced necrotic lesions.
Phylogenetic analysis
RPB2 amplicons were obtained for all 267 isolates, generating sequences of approximately 930 bp, which were deposited in GenBank under accession nos. MZ039858 to MZ040124. The RPB2 matrix included 271 taxa (267 isolates from this study and 4 taxa from GenBank), composed of 847 sites (740 conserved) and 91 parsimony-informative characters. The BI tree was reconstructed using the GTR nucleotide substitution model. The RPB2 tree (Fig. 3) showed that the Ramulariopsis isolates were grouped into two distinct clades (the nucleotide matrices and phylogenetic tree are available in TreeBASE; study number S28159). Clade II gathered most (94.4%) of the isolates from the states of Bahia (44), Distrito Federal (21), Goiás (31), Maranhão (30), Mato Grosso (87) and Mato Grosso do Sul (39). The remaining 15 isolates (5.6%) were grouped in Clade I, with 11 isolates from the Distrito Federal and four from the state of Paraíba.
Bayesian phylogenetic tree based on RPB2 sequences of Ramulariopsis species. Bayesian posterior probabilities (BPP) and Maximum Likelihood bootstrap support values (MLBS) are indicated at the nodes (BPP/MLBS), and the scale bar represents the number of expected changes per site. Ex-type isolates are highlighted in bold. The ex-type CBS 141099 of Ramulariopsis gossypii was used as outgroup.
The GAPDH sequences obtained by nanopore sequencing were 571 and 636 bp in length for R. gossypii and R. pseudoglycines, respectively. The GAPDH sequence of R. pseudoglycines showed 135 single nucleotide polymorphisms compared to R. gossypii sequences. The GAPDH sequences of Ramulariopsis gossypii obtained by nanopore sequencing were identical to the sequences obtained by Sanger sequencing. To correctly delimit the Ramulariopsis isolates at the species level, a multilocus approach was adopted using the RPB2, LSU, EF1-α, ITS, ACT, and HIS3 sequences. A total of 21 taxa (Supplementary Table S2) were included in the BI and ML phylogenetic analyses. The RPB2, LSU, EF1-α, ITS, ACT, and HIS3 individually aligned data sets were 942, 873, 1112, 182, 156, and 346 bp in length, respectively (single gene trees are available in TreeBASE; study number S28159). The concatenate alignment comprised 3611 characters, with 3316 and 281 conserved and variable sites, respectively. Also, 279 sites were determined as phylogenetically informative. The Bayesian phylogenetic tree was reconstructed considering the best nucleotide substitution model for each partition in the concatenate data, GTR (RPB2), HKY (EF1-α, HIS3, ITS, LSU) and K80 (ACT). The Ramulariopsis isolates reported here were grouped into two distinct phylogenetic clades (Fig. 4), corresponding to R. gossypii (clade I) and R. pseudoglycines (clade II).
Bayesian phylogenetic tree based on concatenate sequences (RPB2, LSU, EF1-α, ITS, ACT, and HIS3) of Ramulariopsis species. Bayesian posterior probabilities (BPP) and Maximum Likelihood bootstrap support values (MLBS) are indicated at the nodes (BPP/MLBS), and the scale bar represents the number of expected changes per site. Ex-type isolates are highlighted in bold. The type CBS 141100 of Ramulariopsis pseudoglycines was used as outgroup.
Primer design and validation
Primer sequences (Table 1) were compared against obtained sequences in GenBank, with BLAST (Basic Local Alignments Search Tool) analysis showing 100% homology of primers with sequences of isolates belonging to the species for which primers were designed. The primers targeting EF1-α gene were able to specifically amplify only isolates of R. gossypii and R. pseudoglycines (Fig. 5).
Amplicons obtained using RG-TEF-F/RG-TEF-R and RP-TEF-F/RG-TEF-R primers and visualized on 1.5% agarose gels for isolates of R. pseudoglycines (UBADN 2229, UBADN 2551, UBADN 2713, UBADN 3343), R. gossypii (UBADN 2272, UBADN 2940, CCUB 3195, CCUB 3194), Fusarium sp. (CCUB 3293), Colletotrichum sp. (CCUB 460), Talaromyces sp. (CCUB 2934) and Baudoinia sp. (CCUB 2938), Cercospora sp. (CCUB 848), Aspergillus sp. (CCUB 2929), Lasiodiplodia sp. (CCUB 1436), Macrophomina sp. (CCUB 3905), Trichoderma sp. (CCUB 3421) and Phytophthora sp. (CCUB 1739). M = molecular marker 100 bp DNA Ladder Cellco Biotec. NC = negative control. A full-length gel is presented in Supplementary Figure S1.
The primers sets RG-TEF-F/RG-TEF-R and RP-TEF-F/RP-TEF-R, specifically designed for the recognition of R. gossypii and R. pseudoglycines, respectively, successfully amplified a fragment of 900 bp from each R. gossypii isolate (Fig. 5) and an amplicon of 750 bp from each isolate of R. pseudoglycines (Fig. 5), respectively. The other fungal species (samples 10–20) did not have any fragment amplified.
Resultant amplicons were sequenced to confirm primer specificity. Comparison of their sequences with the target regions selected for primer design showed 100% homology, confirming the species-specificity of the primers. No cross-reactions were observed with the other species or genera tested.
Morphological characterization
The morphological characteristics of the isolates belonging to R. gossypii and R. pseudoglycines in the concatenated tree matched well with the description of each species (Table 2). The long conidiophores of R. pseudoglycines are readily distinguished from the short conidiophores of R. gossypii by visualization under stereomicroscope or light microscopy. These morphological differences were recorded by SEM and are illustrated here for the first time (Fig. 6).
(a–h) Ramulariopsis spp. on leaves of Gossypium hirsutum. (a) Lesion with signs of the fungus on the leaf abaxial face. (b) Hyaline conidiophores of R. gossypii on the abaxial side of the leaves. (c) Hyaline conidiophores of R. pseudoglycines on the abaxial side of the leaves. (d) Hyaline conidiophores of R. gossypii (left) and R. pseudoglycines (right) viewed under light microscopy. (e) Fascicle of R. gossypii formed by conidiophores with presence of hyaline conidia. (f) Conidiophores of R. pseudoglycines. (g) Conidiophores of R. gossypii visualized in SEM. (h) Conidiophores of R. pseudoglycines visualized in SEM.
Genetic characterization
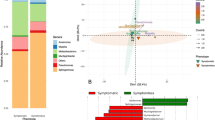
A high interspecific polymorphism and a low intraspecific polymorphism were observed among Ramulariopsis isolates for all 14 markers. The dendrograms based on the binary matrix produced from the band patterns generated with all markers separately were used to analyze the interspecific diversity of Ramulariopsis (data not shown). The ISSR (TGTC)4 molecular marker was selected to estimate the interspecific diversity due to its´ simplicity for species discrimination. The resultant dendrogram of 267 isolates of Ramulariopsis revealed two distinct clades (Fig. 7) corresponding to the clades observed previously in the phylogenetic analysis.
Genealogical network based on the RPB2 gene
Analysis of the genealogical network based on RPB2 gene sequences revealed four distinct haplotypes among the 267 Ramulariopsis isolates (Supplementary table S1). The RPB2 haplotype network (Fig. 8) distinguished two clear-cut clades corresponding to the clades observed in both the phylogenetic tree and the dendrogram. The first clade is restricted only to R. gossypii isolates from the Distrito Federal and Paraíba. The second clade is represented by isolates of R. pseudoglycines from multiple geographic regions, including all sampled locations. Two out of three RPB2 haplotypes were represented by only one isolate of R. pseudoglycines. All isolates of R. gossypii (n = 15) grouped into a single haplotype (H3) while 250 out of the 252 isolates belong to the most frequent R. pseudoglycines haplotype (H1), indicating a strong prevalence of clonal populations.
Haplotype network generated for RPB2 sequences representing seven Brazilian states using Network. Each circle represents a distinct haplotype, proportional in size to its´ frequency in the sample. Hatch marks along the network branches indicate hypothetical mutational steps not detected in the dataset. Geographic origin of isolates from each haplotype by state is represented by colour.
Discussion
Large-scale studies that investigate RLS etiology and the genetic variation of the causal agent are scarce in the literature. The centre of origin of cotton has not been determined, but the main centres of diversity are distributed among regions of Central America, Africa, Arabia and Australia43. In most of the countries within these regions, RLS is considered as a disease of secondary importance, in contrast to the present situation in Brazil.
Global cotton yield has been affected by R. gossypii since 18837. Historically, this species is most widespread and economically important in Brazil, although it has been mostly identified based upon morphological data4,15. Although conidiophore length is useful for separating Ramulariopsis species, taxonomic expertise is required. Interestingly, several isolates previously putatively identified as R. gossypii were molecularly identified as either R. gossypii or R. pseudoglycines15. Comparison of ITS sequences of the Ramulariopsis isolates deposited in GenBank (nucleotide matrices and phylogenetic tree available in TreeBASE; study number S28159) revealed that R. gossypii and R. pseudoglycines were described in previous studies17,44. Isolates collected between 2017 and 2020 and molecularly characterized with sequences of the ITS region were identified as R. pseudoglycines in Brazil25.
Here, the molecular identification of the Ramulariopsis isolates causing RLS on cotton using a polyphasic approach confirmed the presence of R. gossypii and R. pseudoglycines in Brazil. In total, 252 out of 267 isolates were identified as R. pseudoglycines (94%), indicating that this species is the most widespread causal agent of cotton RLS in the Brazilian growing regions today. Additionally, isolates of R. gossypii were restricted to small and isolated farms located in the Distrito Federal and the state of Paraíba, while isolates of R. pseudoglycines were obtained from all sampled locations, and all extensive farms in the Brazilian Cerrado, which is the main cotton growing area in Brazil.
The clade assignments based on the concatenated-sequence tree (RPB2, LSU, EF1-α, ITS, ACT, and HIS3) were identical to those generated by RPB2-sequences trees, the RPB2 haplotype network, the ISSR (TGTC)4 dendrogram, and the morphological comparisons. The most widely employed genomic regions for Ramulariopsis DNA-based identifications have, to date, been based upon ITS sequences, given both their high copy number and easy amplification, and the availability of universal primers. However, for various fungi, the RPB2 molecular marker has been proposed in place of the ITS sequences, due to the lack of resolution in the latter and the potential presence of non-homologous ITS copies in individual fungal genomes21.
On a molecular level, RPB2 sequences are recommended for accurate molecular-based identification of Ramulariopsis, given their universal application, speed, and the presumption that this molecular marker safely approximates taxonomic expertise. However, this technique is laborious, expensive, and requires time and knowledge of phylogenetic analysis for identifying species45. The desire for rapid, automated approaches, such as those obtained here using the ISSR (TGTC)4 primer and EF-α species-specific primers, indicates that the RPB2 region can also potentially be applied for future simple and inexpensive diagnosis and detection assays.
This study showed that EF-α species-specific primers can be used for accurate molecular identification of Ramulariopsis isolates in Brazil, facilitating large-scale surveys of the distribution of species and monitoring of epidemics. Nevertheless, the primers developed here need to be validated for isolates collected in other countries. Prior to this study, the diversity among isolates of Ramulariopsis was verified through ERIC- and REP-PCR profiles for Brazilian isolates23 and RAPD profiles for Indian isolates46, although in both studies, only few isolates were examined, and accurate species identification was not achieved.
Considering the wide distribution of haplotype H1 of R. pseudoglycines, there is evidence for a predominant clonal lineage occurring in Brazil, indicating the existence of a highly efficient mechanism of dispersion over long distances. Although RLS caused by R. gossypii has been recognized for a long time, R. pseudoglycines seems to be firmly prevalent amongst the cotton-producing regions today. When comparing the morphology of Ramulariopsis specimens from earlier studies4,16, the morphological characteristics matched well with the description of R. gossypii.
Given the higher susceptibility of today’s main cotton genotypes and the frequency of fungicide applications, as well as the presence of the G143A substitution in the CYTB25 gene of R. pseudoglycines, which reduces sensitivity to strobilurins, we can hypothesize that extensive cultivation of a limited number of cotton genotypes over many successive growing seasons in the Cerrado has resulted in the increase in population size of R. pseudoglycines. An alternative hypothesis may be that R. pseudoglycines inhabited the native Cerrado vegetation and then spread to cotton plants. Clearly, this species has thus become the most important pathogen negatively impacting cotton production in Brazil today.
This is the first large-scale study that investigates the diversity of Ramulariopsis isolates associated with cotton. Validation of the EF-α species-specific primers as a tool to study the abundance and distribution of Ramulariopsis species will make it possible to carry out extensive RLS sampling studies worldwide. Finally, the correct identification of the RLS causal agent and its geographical distribution is essential for predicting resistance breakdown, guiding pesticide regimes and the development of disease-resistant genotypes.
Methods
Sampling and isolation
Cotton leaves showing typical symptoms of RLS were collected in the 2017–18 and 2018–19 growing seasons from 24 commercial fields in the Brazilian states of Bahia, Distrito Federal, Goiás, Maranhão, Mato Grosso, Mato Grosso do Sul and Paraíba (Fig. 1).
Fungal isolation into pure culture was carried out by the direct method47 in Petri dishes containing water-agar (WA) medium (20 g/L of agar). After 14 days of growth in WA, pure cultures were established by transferring a fragment of a hyphal tip to a new Petri dish containing malt extract (ME) medium (20 g/L malt extract and 20 g/L agar).
Isolates (Supplementary Table S1) were deposited in the Coleção de Culturas da Universidade de Brasília (CCUB; Brasília, Brazil) and stored at 18 ± 1 °C in sterile water48, 10% (v / v) sterile glycerol, and half potato-dextrose-agar (500 mL/L potato broth, 20 g/L agar and 20 g/L dextrose) slopes covered with sterile mineral oil.
DNA extraction
Four mycelial discs (5 mm in diameter) were removed from the margin of 20-day-old pure cultures on ME and transferred to 250 mL conical flasks containing 50 mL of potato dextrose broth with the addition of streptomycin (500 mL/L potato broth, 20 g/L dextrose, 100 µg/mL streptomycin) and incubated at 25 °C, with a 12 h photoperiod.
After seven days growth, the developed mycelium was recovered on filter paper and transferred to 1.5 mL microtubes containing 30 µL of Tris–EDTA (TE) buffer, four metal beads (2.8 mm), and 600 mL of Nuclei Lysis Solution (Promega®). Total DNA extraction was performed using the Wizard Genomic DNA Purification Kit (Promega®) according to the manufacturer’s instructions. Total DNA preparations were analyzed via 1% agarose gel electrophoresis, stained with GelRed (Biotium R), and visualized under UV light. The DNA samples were stored at − 20 °C.
Amplification and sequencing
Partial sequences of the gene encoding the second largest RNA polymerase II subunit (RPB2) were amplified using the specific PCR primers shown in Table 1. This genomic region was employed as the primary barcode for identification of Ramulariopsis species, given the high PCR success rate and easy alignment of the nucleotide sequences. To assign definite species demarcations for the Ramulariopsis isolates, partial nucleotide sequences of six nuclear genes, namely: 28S rRNA (LSU), the internal transcribed spacers of the ribosomal DNA (ITS), actin (ACT), elongation factor (EF1-α), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), and histone H3 (HIS3) were obtained from representative isolates of different clades and locations preliminarily identified based on RPB2 sequence data (Fig. 3). The amplification of the GAPDH gene of R. pseudoglycines isolates resulted in double bands from which the band with the correct estimated size was subsequently purified from agarose gels and sequenced by nanopore sequencing49,50. All primers employed are listed in Table 1, with respective annealing and extension parameters. The PCR mixtures consisted of 6.25 µL of MyTaq PCR Master Mix (2 ×), 0.3 µL of each primer (Table 1), 1 µL of genomic DNA (25 ng/µL) and 4.65 µL of ultrapure water. The cycling conditions were: Initial denaturation at 95 °C for 1.5 min, followed by 35 cycles at 95 °C for 20 s; annealing and extension according to Table 1 and a final extension at 72 °C for 5 min. PCR products were purified and bidirectionally Sanger-sequenced.
Phylogenetic analyses
To determine to which Ramulariopsis species each isolate shared the highest nucleotide identity, the partial nucleotide sequences and the BLASTn algorithm were used to search the NCBI-GenBank nonredundant nucleotide database. A Bayesian phylogenetic tree was initially reconstructed using the RPB2 sequences from the 267 isolates characterized here, and four representative isolates of Ramulariopsis. The ex-epitype CBS 141099 of R. gossypii was used as an outgroup. Also, phylogenetic trees were individually inferred from each genomic region analyzed here. Multiple sequence alignments were obtained with MAFFT v751. Finally, Bayesian Inference (BI) and Maximum Likelihood (ML) phylogenetic trees were reconstructed using the concatenate data (RPB2, LSU, EF1-α, ITS, ACT, and HIS3). For BI, the best nucleotide substitution models were determined, for each partition, with MrModeltest. The CIPRES web portal52 was used to run MrBayes v3.2.153. The Markov Chain Monte Carlo (MCMC) analysis was run with a total of 10 million generations, sampling every 1,000 generations. The convergence of the log likelihoods was confirmed using TRACER v1.7.154. The first 25% of the sampled trees were discarded as burn-in, with the posterior probability (PP) values calculated with the remaining trees. The ML tree was reconstructed using RAxML v.855, accessed through the CIPRES web portal52, assuming a general time reversible (GTR) nucleotide substitution model with a gamma (G) rate of heterogeneity, and 1,000 bootstrap replicates. Phylogenetic trees were visualized and edited in FigTree v1.456 and Inkscape.
Primer design and validation
The EF1-α sequences of R. gossypii and R. pseudoglycines were selected and aligned to enable searching for species-specific primers using Primer3 Plus and Primer-BLAST57,58. Additionally, divergent regions within the EF1-α sequences were selected for manual primers development. The specificities of the primer sequences were in-silico-tested prior to synthesis by searching similar DNA sequences on the NCBI database. Each specific primer was checked for the following parameters: primer length, primer melting temperature, GC content, GC clamp, primer secondary structures (hairpins, self-dimer, and cross dimer), repeats, runs and 3′ end stability45.
Seven species-specific primers were designed and screened against eight isolates from R. gossypii (n = 4) and R. pseudoglycines (n = 4). The screening also included ten fungal genera (Aspergillus sp., Baudoinia sp., Cercospora sp., Colletotrichum sp., Fusarium sp., Lasiodiplodia sp., Phytophthora sp., Macrophomina sp., Talaromyces sp., and Trichoderma sp.) that may occur on cotton plants or that can be found as contaminants. Each amplification was repeated at least twice in separate assays. The PCR parameters were the same as those mentioned above. Amplification products were visualized on 1.5% agarose gels stained with EtBr. After the initial screening, the validated primers were tested on all isolates.
Light microscopy and SEM morphological characterization
For morphological characterization, specimens were initially observed with a Leica 205C stereomicroscope (Leica Biosystems, Nussloch GmbH, Nussloch, Germany). The microscopical characteristics were analyzed by mounting asexual structures in clear lactoglycerol, and 50 measurements for each morphological parameter were carried out at a magnification of × 1,000 using a Leica DM2500 light microscope equipped with a Leica DFC 490 digital camera, coupled to a computer containing the Leica Qwin-Plus software. The morphological characteristics of the isolates were compared with the description of R. gossypii and R. pseudoglycines14,15.
For examination on a scanning electron microscope (JOEL JSM-700 1F model), fragments of symptomatic dry leaves were fixed in 10 mm diameter copper stubs with double-sided carbon tape and coated with 25 mA gold, 1.10–2 mbar, for 2.5 min.
Genetic characterization
Seventeen isolates of R. gossypii (n = 3) and R. pseudoglycines (n = 14) were subjected to characterization with different molecular markers (CIIRAP1-4, CIIRAP2-4, REP, ERIC, BOX, M13, N21, CAG5, GA8, GACAC3, TGTC4, GATA4, GTG5 and GACA4) which are typically highly polymorphic and useful in analysis of genetic variability of fungi23,32,36.
PCR amplifications were performed in a final volume of 12.5 µL: 6.25 µL of MyTaq PCR Master Mix (2 ×), 2.5 µL of primer, 1 µL of genomic DNA (25 ng/µL) and 2.75 µL of ultrapure water. Different volumes of primer were used for REP and ERIC (0.5 μl), and BOX (1 μl) molecular markers, with a final reaction volume again adjusted to 12.5 μL. The PCR conditions for each molecular marker are shown in the references listed in Table 1. Each amplification was repeated at least twice in separate assays.
The amplified products were evaluated as presence (1) or absence (0) of bands and recorded in a binary matrix. This matrix was added to the PAST3 software59, where the Jaccard similarity index was calculated for each combination of two samples. From the similarity index, dendrograms were constructed according to the unweighted pair group method with arithmetic mean (UPGMA).
Genealogical network based on the RPB2 gene
To characterize genetic diversity of R. pseudoglycines and R. gossypii, an analysis of haplotypes was performed using the RPB2 sequences of the 267 isolates. Haplotype identification was performed using the program DnaSP ver. 5.10.160. A haplotype network to visualize the relationships among haplotypes representing seven Brazilian states was reconstructed using NETWORK 4.5.0.2 (Fluxus Technology Ltd.), with gaps and missing data excluded61.
Ethics statement
The study complies with relevant institutional, national, and international guidelines and legislation. The activity of access to Genetic Heritage was registered with SisGen, compliance with law no. 13,123/2015 and its regulations, under permit number A724B5B dated 04/30/2018.
Data availability
The datasets generated in this study can be found in Genbank: MZ039858-MZ040124, MZ066658-MZ066720, and OM419332- OM419338; Treebase: S28159. The results obtained in this study are included in the contents of this report.
References
Santos, R. F. & Barros, M. A. L. Economia do Algodão. In Algodão: o Produtor Pergunta, A Embrapa Responde (eds Beltrão, N. E. M. & Araújo, A. E.) (Embrapa Algodão, 2004).
CONAB. In Companhia Nacional de Abastecimento. Acompanhamento da safra brasileira de grãos. https://www.conab.gov.br/info-agro/safras/graos (2020).
Rodrigues, J. C. J. Algodão no Brasil: Mudança, associativismo e crescimento. In Algodão no Cerrado do Brasil (ed. Freire, E. C.) 21–38 (Positiva, 2015).
Silva, J. C., Bettiol, W. & Suassuna, N. D. Ramularia leaf spot: an emergent disease of cotton in Brazil. Trop. Plant Pathol. 44, 473–482. https://doi.org/10.1007/s40858-019-00308-w (2019).
Xavier, T. W. et al. Identification of ramularia leaf blight cotton disease infection levels by multispectral, multiscale UAV imagery. Drones 3(2), 33. https://doi.org/10.3390/drones3020033 (2019).
Suassuna, N. D. & Coutinho, W. M. Manejo das principais doenças do algodoeiro no Cerrado brasileiro. In Algodão no Cerrado do Brasil (ed. Freire, E. C.) 365–408 (Positiva, 2015).
Spegazzini, C. Fungi guaranitici. Pugillus I. Anales de la Sociedad Científica Argentina 22(4), 209 (1886).
Atkinson, G. F. A new Ramularia on cotton. In Botanical Gazette (eds Arthur, J. C. et al.) 166–168 (Carlon & Hollenbeck, 1890).
Chitarra, L. G. Identificação e Controle das Principais Doenças do Algodoeiro 3rd edn, 1–82 (Embrapa Algodão, 2014).
Farr, D. F. & Rossman, A. Y. Fungal Databases, U.S. National Fungus Collections, ARS, USDA. https://nt.ars-grin.gov/fungaldatabases/ (2021).
Ehrlic, E. & Wolf, F. A. Areolate mildew of cotton. Phytopathology 22, 229–240 (1932).
Tormen, N. R. & Blum, L. E. B. Ramularia leaf spot effect on yield and fiber quality of cotton submitted to fungicide application. Revista Caatinga 32, 634–646. https://doi.org/10.1590/1983-21252019v32n308rc (2019).
Suassuna, N. D., Chitarra, L. G., Asmus, G. L. & Inomoto, M. M. Manejo de doenças do algodoeiro. Circular Técnica 97. https://www.infoteca.cnptia.embrapa.br/bitstream/doc/274818/1/CIRTEC97.pdf (2006).
Braun, U. Studies on Ramularia and allied genera (VI). Nova Hedwigia 56, 432–433 (1993).
Videira, S. I. R., Groenewald, J. Z., Braun, U., Shin, H. D. & Crous, P. W. All that glitters is not Ramularia. Stud. Mycol. 83, 49–163. https://doi.org/10.1016/j.simyco.2016.06.001 (2016).
Curvelo, C. R. S., Rodrigues, F. A., Berger, P. G. & Rezende, D. C. Microscopia eletrônica de varredura do processo infeccioso de Ramularia areola em folhas de algodoeiro. Trop. Plant Pathol. 35(2), 108–113. https://doi.org/10.1590/S1982-56762010000200006 (2010).
Mehta, Y. R. et al. Mycosphaerella areola - the teleomorph of Ramularia areola of cotton in Brazil, and its epidemiological significance. Am. J. Plant Sci. 7, 1415–1422. https://doi.org/10.4236/ajps.2016.710135 (2016).
Crous, P. W., Groenewald, J. Z., Risède, J., Simoneau, P. & Hywel-Jones, N. L. Calonectria species and their Cylindrocladium anamorphs: species with sphaeropedunculate vesicles. Stud. Mycol. 50, 415–430. https://doi.org/10.3114/sim.55.1.213 (2004).
Decloquement, J. et al. Phytophthora theobromicola sp. Nov.: a new species causing black pod disease on Cacao in Brazil. Front. Microbiol. 15, 1–15. https://doi.org/10.3389/fmicb.2021.537399 (2021).
Videira, S. I. R. et al. Elucidating the Ramularia eucalypti species complex. Persoonia 34, 50–64. https://doi.org/10.3767/003158515X685670 (2015).
Lücking, R. et al. Unambiguous identification of fungi: Where do we stand and how accurate and precise is fungal DNA barcoding?. IMA Fungus 11(14), 1–32. https://doi.org/10.1186/s43008-020-00033-z (2020).
Pezenti, L. F. et al. Phenotypic variability among isolates of Ramularia areola from Brazilian cotton. Trop. Plant Pathol. 38(4), 329–331. https://doi.org/10.1590/S1982-56762013005000023 (2013).
Girotto, L. et al. Identification of phenotypic and genotypic variability among the isolates of Ramularia areola of Brazilian cotton. Am. J. Plant Sci. 4(9), 1893–1898. https://doi.org/10.4236/ajps.2013.49232 (2013).
Zandona, C. et al. Mechanisms of resistance and presence of different resistance genes to Ramularia areola in two cotton genotypes. Trop. Plant Pathol. 37(3), 175–178. https://doi.org/10.1590/S1982-56762012000300002 (2012).
Mathioni, S. M. et al. Species determination and CYTB-G143A monitoring of Ramulariopsis spp. isolated from cotton in Brazil. Plant Health Progr. 23, 4–6. https://doi.org/10.1094/PHP-05-21-0081-SC (2022).
Carbone, I. & Kohn, L. M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 91(3), 553–556. https://doi.org/10.1080/00275514.1999.12061051 (1999).
Martin, B. et al. A highly conserved repeated DNA element located in the chromosome of Steptococcus pneumoniae. Nucleic Acid Res. 20(13), 3479–3483. https://doi.org/10.1093/nar/20.13.3479 (1992).
Jacobs, K. et al. Leptographium wingfieldii introduced into North America and found associated with exotic Tomicus piniperda and native bark beetles. Mycol. Res. 108(4), 411–418. https://doi.org/10.1017/S0953756204009748 (2004).
Hulton, C. S., Higgins, C. F. & Sharp, P. M. ERIC sequences: a novel family of repetitive elements in the genomes of Escherichia coli, Salmonela typhimurium and other enterobacteria. Mol. Microbiol. 5(4), 825–834. https://doi.org/10.1111/j.1365-2958.1991.tb00755.x (1991).
Berbee, M. L., Pirseyedi, M. & Hubbard, S. Cochliobolus phylogenetics and the origin of known, highly virulent pathogens, inferred from ITS and glyceraldehyde-3-phosphate dehydrogenase gene sequences. Mycologia 91, 964–977 (1999).
Santos, L. V. et al. Development of new molecular markers for the Colletotrichum genus using RetroCl1 sequences. World J. Microbiol. Biotechnol. 28(3), 1087–1095. https://doi.org/10.1007/s11274-011-0909-x (2011).
Santana, M. F., Batista, A. D., Ribeiro, L. E., Araújo, E. F. & Queiroz, M. V. Terminal repeat retrotransposons as DNA markers in fungi. J. Basic Microbiol. 53(10), 823–827. https://doi.org/10.1002/jobm.201200453 (2012).
Rodrigues, R. J. & Yoder, O. C. A family of conserved repetitive DNA elements from the fungal plant pathogen Glomerella cingulata (Colletotrichum lindemathianum). Exp. Micol. 15, 232–242. https://doi.org/10.1016/0147-5975(91)90025-9 (1991).
Andrea, G. & Xitlali, A. Inter simple sequence repeats (ISSRs). In Ecología Molecular (eds Eguiarte, L. E. et al.) 567–571 (INE, 2007).
Weising, K. et al. Polymorphic simple GATA/GACA repeats in plant genomes. Nucleic Acids Res. 17(23), 10128. https://doi.org/10.1093/nar/17.23.10128 (1989).
Gente, S., Sohier, D., Coton, E., Duhamel, C. & Guéguen, M. Identification of Geotrichum candidum at the species and strain level: proposal for a standardized protocol. J. Ind. Microbiol. Biotechnol. 33, 1019–1031. https://doi.org/10.1007/s10295-006-0130-3 (2006).
Liu, Y. J., Whelen, S. & Hall, B. D. Phylogenetic relationships among ascomycetes: evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 16(12), 1799–1808. https://doi.org/10.1093/oxfordjournals.molbev.a026092 (1999).
Stern, M. J., Ames, G. F. L., Smith, N. H., Robinson, E. C. & Higgins, C. F. Repetitive extragenic palindromic sequences: a major component of the bacterial genome. Cell 37(3), 1015–1026. https://doi.org/10.1016/0092-8674(84)90436-7 (1984).
Bulat, S. A., Mironenko, N. V. & Zholkevich, Y. G. Genetic structure of soil population of fungus Fusarium oxysporum Schlechtend.: Fr.: Molecular reidentification of the species and genetic differentiation of isolates using polymerase chain reaction technique with universal primers (UP PCR). Russ. J. Genet. 31(3), 271–278 (1995).
Vassart, G. et al. A sequence in M13 phage detects hypervariable minisatellites in human and animal DNA. Science 235(4789), 683–684. https://doi.org/10.1126/science.2880398 (1987).
De Hoog, G. S. & Ende, A. H. G. Molecular diagnostics of clinical strains of filamentous Basidiomycetes. Mycoses 41, 183–189. https://doi.org/10.1111/j.1439-0507.1998.tb00321.x (1998).
Vilgalys, R. & Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 172, 4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990 (1990).
Beltrão, N. E. M. Origem e evolução do algodoeiro. In Algodão: O Produtor Pergunta, a Embrapa Responde (eds Beltrão, N. E. M. & Araujo, A. E.) 15–20 (Embrapa Algodão, 2004).
Ganiger, M., Ashtaputre, S. A. & Kulkarni, V. R. Molecular variability of Ramularia areola isolates causing grey mildew of cotton. J. Farm Sci. 30(2), 216–219 (2017).
Santos, K. M. et al. Novel specific primers for rapid identification of Macrophomina species. Eur. J. Plant Pathol. 156, 1213–1218. https://doi.org/10.1007/s10658-020-01952-8 (2020).
Ramanagouda, G. & Ashtaputre, S. A. Collection and characterization of grey mildew (Ramularia areola Atk.) pathogen of cotton. Indian Phytopathol. 72, 301–307. https://doi.org/10.1007/s42360-019-00123-y (2019).
Alfenas, A. C., Ferreira, F. A., Mafia, R. G. & Gonçalves, R. C. Isolamento de fungos fitopatogênicos. In Métodos em Fitopatologia (eds Alfenas, A. C. & Mafia, R. G.) 55–91 (UFV, 2016).
Castellani, A. Viability of some pathogenic fungi in distilled water. Am. J. Trop. Med. Hyg. 24, 270–276 (1939).
Loman, N. J., Quick, J. & Simpson, J. T. A complete bacterial genome assembled de novo using only nanopore sequencing data. Nat. Methods 12, 733–735. https://doi.org/10.1038/nmeth.3444 (2015).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Katoh, K. & Standley, D. M. MAFFT Multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evolut. 30, 772–780. https://doi.org/10.1093/molbev/mst010 (2013).
Miller, M. A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. in Proceedings of the Gateway Computing Environments Workshop (GCE) 1–8 (New Orleans, LA) https://doi.org/10.1109/GCE.2010.5676129 (2010).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19(12), 1572–1574. https://doi.org/10.1093/bioinformatics/btg180 (2003).
Rambaut, A. & Drummond, A. in Tracer MCMC Trace Analysis Tool. http://tree.bio.ed.ac.uk/software/tracer/ (2018).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313. https://doi.org/10.1093/bioinformatics/btu033 (2014).
Rambaut, A. FigTree 1.4.0. http://tree.bio.ed.ac.uk/software/figtree/ (2012).
Hung, J. H. & Weng, Z. Designing polymerase chain reaction primers using Primer3Plus. Cold Spring Harb. Protoc. 2016, 821–826. https://doi.org/10.1101/pdb.prot093096 (2016).
Ye, J. et al. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformat. 13, 134. https://doi.org/10.1186/1471-2105-13-134 (2012).
Hammer, O., Harper, D. A. T. & Ryan, P. D. Paleontological statistics software: package for education and data analysis. Palaeontol. Electron. 4(1), 1–9 (2001).
Librado, P. J. R. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25(11), 1451–1452. https://doi.org/10.1093/bioinformatics/btp187 (2009).
Bandelt, H. J., Forster, P. & Roehl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48. https://doi.org/10.1093/oxfordjournals.molbev.a026036 (1999).
Acknowledgements
The authors thank Rede de Pesquisa Ramularia for sampling of Ramulariopsis isolates in the main cotton producing regions in Brazil.
Funding
This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, Brazil (CAPES), Finance Code 001 and the Universidade de Brasília (UnB). We also acknowledge the financial support of the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Fundação de Apoio a Pesquisa do Distrito Federal (FAP-DF) through Grants 432974/2016-4 and 0193.001491/2017, respectively. A. C. Café-Filho, D. B. Pinho and R.N.G. Miller acknowledge CNPq for research productivity fellowships.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by A.S.S., M.H.L.R., A.C.R.Q. and D.B.P. Conceptualization, review and editing were performed by A.C.C.-F., A.E.A. and R.N.G.M. The first draft of the manuscript was written by A.S.S. and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
da Silva, A.S., Rennó, M.H.L., Quitania, A.C.R. et al. Ramularia leaf spot: PCR-based methods reveal widespread distribution of Ramulariopsis pseudoglycines and limited presence of R. gossypii in Brazil. Sci Rep 13, 9826 (2023). https://doi.org/10.1038/s41598-023-33530-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-33530-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.